+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-3726 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
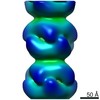
| Title | BtubABC mini microtubule | |||||||||
 Map data Map data | BtubABC mini microtubule | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | bacterial cytoskeleton / microtubules / structural protein | |||||||||
| Function / homology |  Function and homology information Function and homology informationchloroplast organization / microtubule-based process / structural constituent of cytoskeleton / microtubule / hydrolase activity / protein serine/threonine kinase activity / GTPase activity / GTP binding Similarity search - Function | |||||||||
| Biological species |  Prosthecobacter dejongeii (bacteria) / Prosthecobacter dejongeii (bacteria) /  Prosthecobacter vanneervenii (bacteria) Prosthecobacter vanneervenii (bacteria) | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 3.6 Å | |||||||||
 Authors Authors | Deng X / Bharat TAM | |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2017 Journal: Proc Natl Acad Sci U S A / Year: 2017Title: Four-stranded mini microtubules formed by BtubAB show dynamic instability. Authors: Xian Deng / Gero Fink / Tanmay A M Bharat / Shaoda He / Danguole Kureisaite-Ciziene / Jan Löwe /  Abstract: Microtubules, the dynamic, yet stiff hollow tubes built from αβ-tubulin protein heterodimers, are thought to be present only in eukaryotic cells. Here, we report a 3.6-Å helical reconstruction ...Microtubules, the dynamic, yet stiff hollow tubes built from αβ-tubulin protein heterodimers, are thought to be present only in eukaryotic cells. Here, we report a 3.6-Å helical reconstruction electron cryomicroscopy structure of four-stranded mini microtubules formed by bacterial tubulin-like BtubAB proteins. Despite their much smaller diameter, mini microtubules share many key structural features with eukaryotic microtubules, such as an M-loop, alternating subunits, and a seam that breaks overall helical symmetry. Using in vitro total internal reflection fluorescence microscopy, we show that bacterial mini microtubules treadmill and display dynamic instability, another hallmark of eukaryotic microtubules. The third protein in the gene cluster, BtubC, previously known as "bacterial kinesin light chain," binds along protofilaments every 8 nm, inhibits BtubAB mini microtubule catastrophe, and increases rescue. Our work reveals that some bacteria contain regulated and dynamic cytomotive microtubule systems that were once thought to be only useful in much larger and sophisticated eukaryotic cells. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_3726.map.gz emd_3726.map.gz | 78.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-3726-v30.xml emd-3726-v30.xml emd-3726.xml emd-3726.xml | 13.9 KB 13.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_3726_fsc.xml emd_3726_fsc.xml | 9.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_3726.png emd_3726.png | 182.7 KB | ||
| Filedesc metadata |  emd-3726.cif.gz emd-3726.cif.gz | 5.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-3726 http://ftp.pdbj.org/pub/emdb/structures/EMD-3726 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3726 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3726 | HTTPS FTP |
-Validation report
| Summary document |  emd_3726_validation.pdf.gz emd_3726_validation.pdf.gz | 328.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_3726_full_validation.pdf.gz emd_3726_full_validation.pdf.gz | 327.3 KB | Display | |
| Data in XML |  emd_3726_validation.xml.gz emd_3726_validation.xml.gz | 11.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3726 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3726 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3726 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3726 | HTTPS FTP |
-Related structure data
| Related structure data |  5o09MC  5o01C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_3726.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_3726.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | BtubABC mini microtubule | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.34 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : BtubABC mini microtubule
| Entire | Name: BtubABC mini microtubule |
|---|---|
| Components |
|
-Supramolecule #1: BtubABC mini microtubule
| Supramolecule | Name: BtubABC mini microtubule / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|
-Supramolecule #2: BtubABC mini microtubule
| Supramolecule | Name: BtubABC mini microtubule / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Prosthecobacter dejongeii (bacteria) Prosthecobacter dejongeii (bacteria) |
-Supramolecule #3: BtubABC mini microtubule
| Supramolecule | Name: BtubABC mini microtubule / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #3 |
|---|---|
| Source (natural) | Organism:  Prosthecobacter vanneervenii (bacteria) Prosthecobacter vanneervenii (bacteria) |
-Macromolecule #1: Tubulin
| Macromolecule | Name: Tubulin / type: protein_or_peptide / ID: 1 / Number of copies: 8 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Prosthecobacter dejongeii (bacteria) Prosthecobacter dejongeii (bacteria) |
| Molecular weight | Theoretical: 47.071582 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: VNNTIVVSIG QAGNQIAASF WKTVCLEHGI DPLTGQTAPG VAPRGNWSSF FSKLGESSSG SYVPRAIMVD LEPSVIDNVK ATSGSLFNP ANLISRTEGA GGNFAVGYLG AGREVLPEVM SRLDYEIDKC DNVGGIIVLH AIGGGTGSGF GALLIESLKE K YGEIPVLS ...String: VNNTIVVSIG QAGNQIAASF WKTVCLEHGI DPLTGQTAPG VAPRGNWSSF FSKLGESSSG SYVPRAIMVD LEPSVIDNVK ATSGSLFNP ANLISRTEGA GGNFAVGYLG AGREVLPEVM SRLDYEIDKC DNVGGIIVLH AIGGGTGSGF GALLIESLKE K YGEIPVLS CAVLPSPQVS SVVTEPYNTV FALNTLRRSA DACLIFDNEA LFDLAHRKWN IESPTVDDLN LLITEALAGI TA SMRFSGF LTVEITLREL LTNLVPQPSL HFLMCAFAPL TPPDRSKFEE LGIEEMIKSL FDNGSVFAAC SPMEGRFLST AVL YRGIME DKPLADAALA AMREKLPLTY WIPTAFKIGY VEQPGISHRK SMVLLANNTE IARVLDRICH NFDKLWQRKA FANW YLNEG MSEEQINVLR ASAQELVQSY QVAEESGA UniProtKB: Tubulin |
-Macromolecule #2: Tubulin BtubB
| Macromolecule | Name: Tubulin BtubB / type: protein_or_peptide / ID: 2 / Number of copies: 8 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Prosthecobacter dejongeii (bacteria) Prosthecobacter dejongeii (bacteria) |
| Molecular weight | Theoretical: 46.465508 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: VREILSIHVG QCGNQIADSF WRLALREHGL TEAGTLKEGS NAAANSNMEV FFHKVRDGKY VPRAVLVDLE PGVIARIEGG DMSQLFDES SIVRKIPGAA NNWARGYNVE GEKVIDQIMN VIDSAVEKTK GLQGFLMTHS IGGGSGSGLG SLILERLRQA Y PKKRIFTF ...String: VREILSIHVG QCGNQIADSF WRLALREHGL TEAGTLKEGS NAAANSNMEV FFHKVRDGKY VPRAVLVDLE PGVIARIEGG DMSQLFDES SIVRKIPGAA NNWARGYNVE GEKVIDQIMN VIDSAVEKTK GLQGFLMTHS IGGGSGSGLG SLILERLRQA Y PKKRIFTF SVVPSPLISD SAVEPYNAIL TLQRILDNAD GAVLLDNEAL FRIAKAKLNR SPNYMDLNNI IALIVSSVTA SL RFPGKLN TDLSEFVTNL VPFPGNHFLT ASFAPMRGAG QEGQVRTNFP DLARETFAQD NFTAAIDWQQ GVYLAASALF RGD VKAKDV DENMATIRKS LNYASYMPAS GGLKLGYAET APEGFASSGL ALVNHTGIAA VFERLIAQFD IMFDNHAYTH WYEN AGVSR DMMAKARNQI ATLAQSYRDA S UniProtKB: Tubulin BtubB |
-Macromolecule #3: Bacterial kinesin light chain
| Macromolecule | Name: Bacterial kinesin light chain / type: protein_or_peptide / ID: 3 / Number of copies: 8 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Prosthecobacter vanneervenii (bacteria) Prosthecobacter vanneervenii (bacteria) |
| Molecular weight | Theoretical: 27.135703 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: DTALERQIAS ASRSVEEARR LAYHDPIRVG ALVEQISVLA DLRQKEGDFR KAESLYREAL FRAQELRKQD PDLLTGIYSL LAHLYDRWG RMDKAAEFYE LALKISAENG LEESDKVATI KNNLAMIFKQ LRKFERAEGY YCEALETFQR LDGEQSARVA S VYNNLGVL ...String: DTALERQIAS ASRSVEEARR LAYHDPIRVG ALVEQISVLA DLRQKEGDFR KAESLYREAL FRAQELRKQD PDLLTGIYSL LAHLYDRWG RMDKAAEFYE LALKISAENG LEESDKVATI KNNLAMIFKQ LRKFERAEGY YCEALETFQR LDGEQSARVA S VYNNLGVL YYSHMDVDRA QVMHERALAI RQNLHEGQMD PADLSQTFIN LGAVYKAAGD FQKAEACVDR AKRIRAAMNG UniProtKB: Bacterial kinesin light chain |
-Macromolecule #4: GUANOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: GUANOSINE-5'-DIPHOSPHATE / type: ligand / ID: 4 / Number of copies: 16 / Formula: GDP |
|---|---|
| Molecular weight | Theoretical: 443.201 Da |
| Chemical component information |  ChemComp-GDP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | helical array |
- Sample preparation
Sample preparation
| Buffer | pH: 7.7 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI POLARA 300 |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Detector mode: INTEGRATING / Number real images: 6105 / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Tecnai Polara / Image courtesy: FEI Company |
- Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: RECIPROCAL / Protocol: OTHER / Target criteria: R-factor |
|---|---|
| Output model |  PDB-5o09: |
 Movie
Movie Controller
Controller