+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-21816 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM of Form 2 like peptide filament, 29-20-2 | |||||||||
 Map data Map data | Cryo-EM of Form 2 like peptide filament, 29-20-2 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | filament / self-assembly peptide filament / Cryo-EM / PROTEIN FIBRIL | |||||||||
| Biological species | synthetic construct (others) | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 3.8 Å | |||||||||
 Authors Authors | Wang F / Gnewou OM | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2021 Journal: Nat Commun / Year: 2021Title: Structural analysis of cross α-helical nanotubes provides insight into the designability of filamentous peptide nanomaterials. Authors: Fengbin Wang / Ordy Gnewou / Charles Modlin / Leticia C Beltran / Chunfu Xu / Zhangli Su / Puneet Juneja / Gevorg Grigoryan / Edward H Egelman / Vincent P Conticello /  Abstract: The exquisite structure-function correlations observed in filamentous protein assemblies provide a paradigm for the design of synthetic peptide-based nanomaterials. However, the plasticity of ...The exquisite structure-function correlations observed in filamentous protein assemblies provide a paradigm for the design of synthetic peptide-based nanomaterials. However, the plasticity of quaternary structure in sequence-space and the lability of helical symmetry present significant challenges to the de novo design and structural analysis of such filaments. Here, we describe a rational approach to design self-assembling peptide nanotubes based on controlling lateral interactions between protofilaments having an unusual cross-α supramolecular architecture. Near-atomic resolution cryo-EM structural analysis of seven designed nanotubes provides insight into the designability of interfaces within these synthetic peptide assemblies and identifies a non-native structural interaction based on a pair of arginine residues. This arginine clasp motif can robustly mediate cohesive interactions between protofilaments within the cross-α nanotubes. The structure of the resultant assemblies can be controlled through the sequence and length of the peptide subunits, which generates synthetic peptide filaments of similar dimensions to flagella and pili. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_21816.map.gz emd_21816.map.gz | 16.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-21816-v30.xml emd-21816-v30.xml emd-21816.xml emd-21816.xml | 13.3 KB 13.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_21816.png emd_21816.png | 129.6 KB | ||
| Filedesc metadata |  emd-21816.cif.gz emd-21816.cif.gz | 4.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-21816 http://ftp.pdbj.org/pub/emdb/structures/EMD-21816 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21816 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21816 | HTTPS FTP |
-Related structure data
| Related structure data |  6wl7MC  6wkxC  6wkyC  6wl0C  6wl1C  6wl8C  6wl9C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_21816.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_21816.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
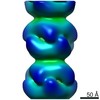
| Annotation | Cryo-EM of Form 2 like peptide filament, 29-20-2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.04 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : self-assembly peptide filament, 29-20-2
| Entire | Name: self-assembly peptide filament, 29-20-2 |
|---|---|
| Components |
|
-Supramolecule #1: self-assembly peptide filament, 29-20-2
| Supramolecule | Name: self-assembly peptide filament, 29-20-2 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all / Details: synthetic peptide |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
-Macromolecule #1: peptide 29-20-2
| Macromolecule | Name: peptide 29-20-2 / type: protein_or_peptide / ID: 1 / Number of copies: 150 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 3.253769 KDa |
| Sequence | String: QAEILEADAR ILRAYAEILK AHAEILKAQ |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Grid | Details: unspecified |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 55.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Helical parameters - Δz: 5.63 Å Applied symmetry - Helical parameters - Δ&Phi: 12.90 ° Applied symmetry - Helical parameters - Axial symmetry: C3 (3 fold cyclic) Resolution.type: BY AUTHOR / Resolution: 3.8 Å / Resolution method: OTHER / Details: Model:Map FSC 0.38 cut off and d99 / Number images used: 67941 |
|---|---|
| CTF correction | Type: PHASE FLIPPING AND AMPLITUDE CORRECTION |
| Startup model | Type of model: OTHER / Details: featureless cylinder |
| Final angle assignment | Type: NOT APPLICABLE |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















