[English] 日本語
 Yorodumi
Yorodumi- EMDB-9896: CryoEM structure of S.typhimurium R-type flagellar filament made ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-9896 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of S.typhimurium R-type flagellar filament made of FljB (A461V) without domain D3 by masking out | |||||||||
 Map data Map data | CryoEM structure of S.typhimurium R-type straight flagellar filament made of FljB (A461V) | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | FljB / Helical reconstruction / Salmonella / Flagellar motor / PROTEIN FIBRIL | |||||||||
| Function / homology |  Function and homology information Function and homology informationbacterial-type flagellum / structural molecule activity / extracellular region Similarity search - Function | |||||||||
| Biological species |  Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 3.56 Å | |||||||||
 Authors Authors | Yamaguchi T / Toma S | |||||||||
| Funding support |  Japan, 1 items Japan, 1 items
| |||||||||
 Citation Citation |  Journal: Biomolecules / Year: 2020 Journal: Biomolecules / Year: 2020Title: Structural and Functional Comparison of Flagellar Filaments Composed of FljB and FliC. Authors: Tomoko Yamaguchi / Shoko Toma / Naoya Terahara / Tomoko Miyata / Masamichi Ashihara / Tohru Minamino / Keiichi Namba / Takayuki Kato /  Abstract: The bacterial flagellum is a motility organelle consisting of a long helical filament as a propeller and a rotary motor that drives rapid filament rotation to produce thrust. serovar Typhimurium has ...The bacterial flagellum is a motility organelle consisting of a long helical filament as a propeller and a rotary motor that drives rapid filament rotation to produce thrust. serovar Typhimurium has two genes of flagellin, and , for flagellar filament formation and autonomously switches their expression at a frequency of 10-10 per cell per generation. We report here differences in their structures and motility functions under high-viscosity conditions. A strain expressing FljB showed a higher motility than one expressing FliC under high viscosity. To examine the reasons for this motility difference, we carried out structural analyses of the FljB filament by electron cryomicroscopy and found that the structure was nearly identical to that of the FliC filament except for the position and orientation of the outermost domain D3 of flagellin. The density of domain D3 was much lower in FljB than FliC, suggesting that domain D3 of FljB is more flexible and mobile than that of FliC. These differences suggest that domain D3 plays an important role not only in changing antigenicity of the filament but also in optimizing motility function of the filament as a propeller under different conditions. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_9896.map.gz emd_9896.map.gz | 228.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-9896-v30.xml emd-9896-v30.xml emd-9896.xml emd-9896.xml | 21.6 KB 21.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_9896_fsc.xml emd_9896_fsc.xml | 14.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_9896.png emd_9896.png | 2 MB | ||
| Masks |  emd_9896_msk_1.map emd_9896_msk_1.map | 244.1 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-9896.cif.gz emd-9896.cif.gz | 6.7 KB | ||
| Others |  emd_9896_half_map_1.map.gz emd_9896_half_map_1.map.gz emd_9896_half_map_2.map.gz emd_9896_half_map_2.map.gz | 194.5 MB 194.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9896 http://ftp.pdbj.org/pub/emdb/structures/EMD-9896 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9896 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9896 | HTTPS FTP |
-Related structure data
| Related structure data |  6jy0MC  0980C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_9896.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_9896.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CryoEM structure of S.typhimurium R-type straight flagellar filament made of FljB (A461V) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_9896_msk_1.map emd_9896_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
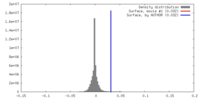
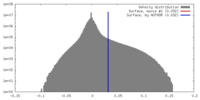
| Density Histograms |
-Half map: EM half map-1 of 3D Refinement CryoEM structure...
| File | emd_9896_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | EM half map-1 of 3D Refinement CryoEM structure of S.typhimurium R-type straight flagellar filament made of FljB (A461V) | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: EM half map-2 of 3D Refinement CryoEM structure...
| File | emd_9896_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | EM half map-2 of 3D Refinement CryoEM structure of S.typhimurium R-type straight flagellar filament made of FljB (A461V) | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : FljB
| Entire | Name: FljB |
|---|---|
| Components |
|
-Supramolecule #1: FljB
| Supramolecule | Name: FljB / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all / Details: R-type straight flagellar filament (A461V) |
|---|---|
| Source (natural) | Organism:  Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) Salmonella enterica subsp. enterica serovar Typhimurium (bacteria)Strain: SJW590 |
-Macromolecule #1: Flagellin
| Macromolecule | Name: Flagellin / type: protein_or_peptide / ID: 1 / Number of copies: 22 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) |
| Molecular weight | Theoretical: 52.608449 KDa |
| Recombinant expression | Organism:  Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) |
| Sequence | String: MAQVINTNSL SLLTQNNLNK SQSALGTAIE RLSSGLRINS AKDDAAGQAI ANRFTANIKG LTQASRNAND GISIAQTTEG ALNEINNNL QRVRELAVQS ANSTNSQSDL DSIQAEITQR LNEIDRVSGQ TQFNGVKVLA QDNTLTIQVG ANDGETIDID L KQINSQTL ...String: MAQVINTNSL SLLTQNNLNK SQSALGTAIE RLSSGLRINS AKDDAAGQAI ANRFTANIKG LTQASRNAND GISIAQTTEG ALNEINNNL QRVRELAVQS ANSTNSQSDL DSIQAEITQR LNEIDRVSGQ TQFNGVKVLA QDNTLTIQVG ANDGETIDID L KQINSQTL GLDSLNVQKA YDVKDTAVTT KAYANNGTTL DVSGLDDAAI KAATGGTNGT ASVTGGAVKF DADNNKYFVT IG GFTGADA AKNGDYEVNV ATDGTVTLAA GATKTTMPAG ATTKTEVQEL KDTPAVVSAD AKNALIAGGV DATDANGAEL VKM SYTDKN GKTIEGGYAL KAGDKYYAAD YDEATGAIKA KTTSYTAADG TTKTAANQLG GVDGKTEVVT IDGKTYNASK AAGH DFKAQ PELAEAAAKT TENPLQKIDA ALAQVDALRS DLGAVQNRFN SAITNLGNTV NNLSEVRSRI EDSDYATEVS NMSRA QILQ QAGTSVLAQA NQVPQNVLSL LR UniProtKB: Flagellin |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 7.8 Component:
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sugar embedding | Material: buffer | ||||||||||||
| Grid | Model: Quantifoil, UltrAuFoil, R1.2/1.3 / Material: MOLYBDENUM / Support film - topology: CONTINUOUS / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 24 sec. / Pretreatment - Atmosphere: AIR | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Detector mode: COUNTING / Digitization - Frames/image: 1-6 / Number grids imaged: 3 / Number real images: 2319 / Average electron dose: 10.3 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Calibrated defocus max: 1.6 µm / Calibrated defocus min: 0.21 µm / Calibrated magnification: 72273 / Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 75000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller


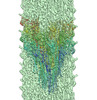





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)