+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: EMDB / ID: EMD-9915 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Cryo-EM structure of Holo-bacterioferritin form-II from Streptomyces coelicolor | |||||||||
 マップデータ マップデータ | Cryo_EM map of bacterioferritin from Streptomyces coelicolor form II | |||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | Ferritin / Nanocage / Bacterioferritin / METAL TRANSPORT | |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報ferroxidase / ferroxidase activity / ferric iron binding / iron ion transport / intracellular iron ion homeostasis / iron ion binding / heme binding / cytosol 類似検索 - 分子機能 | |||||||||
| 生物種 |  Streptomyces coelicolor (バクテリア) Streptomyces coelicolor (バクテリア) | |||||||||
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 4.5 Å | |||||||||
 データ登録者 データ登録者 | Jobichen C / Sivaraman J | |||||||||
 引用 引用 |  ジャーナル: To Be Published ジャーナル: To Be Publishedタイトル: Cryo-EM structure of Bacterioferritin from Streptomyces coelicolor 著者: Jobichen C / Sivaraman J | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_9915.map.gz emd_9915.map.gz | 48.8 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-9915-v30.xml emd-9915-v30.xml emd-9915.xml emd-9915.xml | 11 KB 11 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| 画像 |  emd_9915.png emd_9915.png | 81.1 KB | ||
| Filedesc metadata |  emd-9915.cif.gz emd-9915.cif.gz | 5.1 KB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9915 http://ftp.pdbj.org/pub/emdb/structures/EMD-9915 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9915 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9915 | HTTPS FTP |
-関連構造データ
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_9915.map.gz / 形式: CCP4 / 大きさ: 52.7 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_9915.map.gz / 形式: CCP4 / 大きさ: 52.7 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Cryo_EM map of bacterioferritin from Streptomyces coelicolor form II | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.11 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
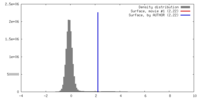
| 密度 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
CCP4マップ ヘッダ情報:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-添付データ
- 試料の構成要素
試料の構成要素
-全体 : Bacterioferritin in complex with HEME and IRON
| 全体 | 名称: Bacterioferritin in complex with HEME and IRON |
|---|---|
| 要素 |
|
-超分子 #1: Bacterioferritin in complex with HEME and IRON
| 超分子 | 名称: Bacterioferritin in complex with HEME and IRON / タイプ: organelle_or_cellular_component / ID: 1 / 親要素: 0 / 含まれる分子: #1 |
|---|---|
| 由来(天然) | 生物種:  Streptomyces coelicolor (バクテリア) Streptomyces coelicolor (バクテリア) |
| 分子量 | 理論値: 500 KDa |
-分子 #1: Bacterioferritin
| 分子 | 名称: Bacterioferritin / タイプ: protein_or_peptide / ID: 1 / コピー数: 24 / 光学異性体: LEVO / EC番号: ferroxidase |
|---|---|
| 由来(天然) | 生物種:  Streptomyces coelicolor (バクテリア) Streptomyces coelicolor (バクテリア) |
| 分子量 | 理論値: 19.242629 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: MQGDPEVIEF LNEQLTAELT AINQYFLHAK LQDHKGWTKL AKYTRAESFD EMRHAEVLTD RILLLDGLPN YQRLFHVRVG QSVTEMFQA DREVELEAID RLRRGIEVMR AKHDITSANV FEAILADEEH HIDYLETQLD LIEKLGESLY LSTVIEQTQP D PSGPGSL UniProtKB: Bacterioferritin |
-分子 #2: FE (II) ION
| 分子 | 名称: FE (II) ION / タイプ: ligand / ID: 2 / コピー数: 24 / 式: FE2 |
|---|---|
| 分子量 | 理論値: 55.845 Da |
-分子 #3: PROTOPORPHYRIN IX CONTAINING FE
| 分子 | 名称: PROTOPORPHYRIN IX CONTAINING FE / タイプ: ligand / ID: 3 / コピー数: 12 / 式: HEM |
|---|---|
| 分子量 | 理論値: 616.487 Da |
| Chemical component information |  ChemComp-HEM: |
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 濃度 | 1.8 mg/mL |
|---|---|
| 緩衝液 | pH: 7 |
| 凍結 | 凍結剤: ETHANE |
| 詳細 | This sample was monodisperse with 24mer nanocage assemblies. |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: FEI FALCON II (4k x 4k) 平均電子線量: 1.6 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | C2レンズ絞り径: 2.7 µm / 照射モード: OTHER / 撮影モード: BRIGHT FIELD |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
- 画像解析
画像解析
| 初期モデル | モデルのタイプ: PDB ENTRY PDBモデル - PDB ID: |
|---|---|
| 最終 再構成 | 想定した対称性 - 点群: O (正8面体型対称) / 解像度のタイプ: BY AUTHOR / 解像度: 4.5 Å / 解像度の算出法: FSC 0.143 CUT-OFF / ソフトウェア - 名称: cisTEM / 使用した粒子像数: 13793 |
| 初期 角度割当 | タイプ: RANDOM ASSIGNMENT / ソフトウェア - 名称: cisTEM |
| 最終 角度割当 | タイプ: ANGULAR RECONSTITUTION / ソフトウェア - 名称: cisTEM |
 ムービー
ムービー コントローラー
コントローラー
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)