+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-9361 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of chicken Otop3 in nanodiscs | |||||||||
 Map data Map data | Structure of chOtop3 in lipidic nanodiscs | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Proton channel / Otopetrin / MEMBRANE PROTEIN | |||||||||
| Biological species |   | |||||||||
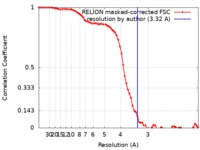
| Method | single particle reconstruction / cryo EM / Resolution: 3.32 Å | |||||||||
 Authors Authors | Saotome K / Lee WH | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2019 Journal: Nat Struct Mol Biol / Year: 2019Title: Structures of the otopetrin proton channels Otop1 and Otop3. Authors: Kei Saotome / Bochuan Teng / Che Chun Alex Tsui / Wen-Hsin Lee / Yu-Hsiang Tu / Joshua P Kaplan / Mark S P Sansom / Emily R Liman / Andrew B Ward /   Abstract: Otopetrins (Otop1-Otop3) comprise one of two known eukaryotic proton-selective channel families. Otop1 is required for otoconia formation and a candidate mammalian sour taste receptor. Here we report ...Otopetrins (Otop1-Otop3) comprise one of two known eukaryotic proton-selective channel families. Otop1 is required for otoconia formation and a candidate mammalian sour taste receptor. Here we report cryo-EM structures of zebrafish Otop1 and chicken Otop3 in lipid nanodiscs. The structures reveal a dimeric architecture, with each subunit forming 12 transmembrane helices divided into structurally similar amino (N) and carboxy (C) domains. Cholesterol-like molecules occupy various sites in Otop1 and Otop3 and occlude a central tunnel. In molecular dynamics simulations, hydrophilic vestibules formed by the N and C domains and in the intrasubunit interface between N and C domains form conduits for water entry into the membrane core, suggesting three potential proton conduction pathways. By mutagenesis, we tested the roles of charged residues in each putative permeation pathway. Our results provide a structural basis for understanding selective proton permeation and gating of this conserved family of proton channels. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_9361.map.gz emd_9361.map.gz | 28.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-9361-v30.xml emd-9361-v30.xml emd-9361.xml emd-9361.xml | 11.5 KB 11.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_9361_fsc.xml emd_9361_fsc.xml | 7.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_9361.png emd_9361.png | 171.7 KB | ||
| Filedesc metadata |  emd-9361.cif.gz emd-9361.cif.gz | 5.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9361 http://ftp.pdbj.org/pub/emdb/structures/EMD-9361 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9361 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9361 | HTTPS FTP |
-Related structure data
| Related structure data |  6nf6MC  9360C  6nf4C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_9361.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_9361.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Structure of chOtop3 in lipidic nanodiscs | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.03 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Zebrafish Otopetrin1 in lipidic nanodiscs
| Entire | Name: Zebrafish Otopetrin1 in lipidic nanodiscs |
|---|---|
| Components |
|
-Supramolecule #1: Zebrafish Otopetrin1 in lipidic nanodiscs
| Supramolecule | Name: Zebrafish Otopetrin1 in lipidic nanodiscs / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 65853 kDa/nm |
-Macromolecule #1: Otopetrin3
| Macromolecule | Name: Otopetrin3 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 63.744496 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: GPAGNKASKQ KFCHHCNSRS ASAPPGTSTI HYEKSWLYRH CSLQQRDRQA QKSGQLFSGL LALNVVFLGS AFISSMIFNH VAITLADVW ILLSILKVLC LCWIIYYLLG TSRQPHAVLY RDTHAAPVWI RGSLLLFGTF SILLNVFQIG YSVIQINCKS K VEIVFPSI ...String: GPAGNKASKQ KFCHHCNSRS ASAPPGTSTI HYEKSWLYRH CSLQQRDRQA QKSGQLFSGL LALNVVFLGS AFISSMIFNH VAITLADVW ILLSILKVLC LCWIIYYLLG TSRQPHAVLY RDTHAAPVWI RGSLLLFGTF SILLNVFQIG YSVIQINCKS K VEIVFPSI EILFVATQAF FLWHHSKDCI QVQHNLTRCG LMLTIATNLL LWLLAVTNDS IHMEIESQLR EVEQRLAGNE TD SCACPNT TTCKVFQKGY ILLYPFNTEY CLICCSVLYV MWKNVGRRIS HHHIAHIKPK FKLQGVVFGP LLGAAAVIIG ICV FMMYQI QATGSAPNYQ VFVLYYSYYI VLLPLMCVVA IIGTIIHTLE KRELDTLKNP TRSLDVVLLM GAALGQIAMS YFSI VAIVA TNPRDMLNSL ILSYSVLLIF QYITQNIFII DGLQRQPFAK EEEVSEEHNR EAPDQRRVSV LELGQEFRRV SLSYI HTYS HLGWKRKALK EISFFLVLCN IILWIMPTFG AHPVFENGLQ KSFYGYSTWF AIVNFGLPLS VFYRMHSVGG LLEVYV SA |
-Macromolecule #2: CHOLESTEROL HEMISUCCINATE
| Macromolecule | Name: CHOLESTEROL HEMISUCCINATE / type: ligand / ID: 2 / Number of copies: 2 / Formula: Y01 |
|---|---|
| Molecular weight | Theoretical: 486.726 Da |
| Chemical component information |  ChemComp-Y01: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2.1 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 283 K |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-6nf6: |
 Movie
Movie Controller
Controller














 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)