[English] 日本語
 Yorodumi
Yorodumi- EMDB-7972: cryo-EM reconstruction of the mitochondrial calcium uniporter (MC... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-7972 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | cryo-EM reconstruction of the mitochondrial calcium uniporter (MCU) from Zebrafish | |||||||||
 Map data Map data | Cryo-EM structure of the mitochondrial calcium uniporter | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 8.5 Å | |||||||||
 Authors Authors | Long SB / Baradaran R / Wang C | |||||||||
 Citation Citation |  Journal: Nature / Year: 2018 Journal: Nature / Year: 2018Title: Cryo-EM structures of fungal and metazoan mitochondrial calcium uniporters. Authors: Rozbeh Baradaran / Chongyuan Wang / Andrew Francis Siliciano / Stephen Barstow Long /  Abstract: The mitochondrial calcium uniporter (MCU) is a highly selective calcium channel and a major route of calcium entry into mitochondria. How the channel catalyses ion permeation and achieves ...The mitochondrial calcium uniporter (MCU) is a highly selective calcium channel and a major route of calcium entry into mitochondria. How the channel catalyses ion permeation and achieves ion selectivity are not well understood, partly because MCU is thought to have a distinct architecture in comparison to other cellular channels. Here we report cryo-electron microscopy reconstructions of MCU channels from zebrafish and Cyphellophora europaea at 8.5 Å and 3.2 Å resolutions, respectively. In contrast to a previous report of pentameric stoichiometry for MCU, both channels are tetramers. The atomic model of C. europaea MCU shows that a conserved WDXXEP signature sequence forms the selectivity filter, in which calcium ions are arranged in single file. Coiled-coil legs connect the pore to N-terminal domains in the mitochondrial matrix. In C. europaea MCU, the N-terminal domains assemble as a dimer of dimers; in zebrafish MCU, they form an asymmetric crescent. The structures define principles that underlie ion permeation and calcium selectivity in this unusual channel. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_7972.map.gz emd_7972.map.gz | 58.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-7972-v30.xml emd-7972-v30.xml emd-7972.xml emd-7972.xml | 15.2 KB 15.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_7972.png emd_7972.png | 32.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-7972 http://ftp.pdbj.org/pub/emdb/structures/EMD-7972 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7972 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7972 | HTTPS FTP |
-Validation report
| Summary document |  emd_7972_validation.pdf.gz emd_7972_validation.pdf.gz | 78.6 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_7972_full_validation.pdf.gz emd_7972_full_validation.pdf.gz | 77.7 KB | Display | |
| Data in XML |  emd_7972_validation.xml.gz emd_7972_validation.xml.gz | 493 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-7972 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-7972 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-7972 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-7972 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_7972.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_7972.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM structure of the mitochondrial calcium uniporter | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.384 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
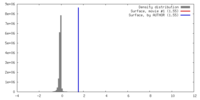
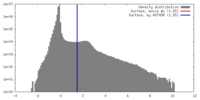
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Mitochondrial Calcium Uniporter (MCU) from Zebrafish
| Entire | Name: Mitochondrial Calcium Uniporter (MCU) from Zebrafish |
|---|---|
| Components |
|
-Supramolecule #1: Mitochondrial Calcium Uniporter (MCU) from Zebrafish
| Supramolecule | Name: Mitochondrial Calcium Uniporter (MCU) from Zebrafish / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 133 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) / Recombinant cell: HEK293S GnTI - Homo sapiens (human) / Recombinant cell: HEK293S GnTI - |
-Macromolecule #1: Mitochondrial Calcium Uniporter (MCU) from Zebrafish
| Macromolecule | Name: Mitochondrial Calcium Uniporter (MCU) from Zebrafish / type: protein_or_peptide / ID: 1 Details: Zebrafish (Danio rerio) MCU expressed in mammalian cells and purified in digitonin detergent. Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: LLCSPAAEDV SVVYQNGLPV ISVRLPSRRE R CQFTLKPL SDTVGVFLQQ LQAEDRGIDR VTIYSADGAR IASSTGIDIL LMDNFKLVIN DT SYLVQPP RRDLLPHEDG ERLNDVKILV QQLYTTLRIE EHQLNKEREL IGRLEDLNSQ LQP LEKVKE ELSKKAERRT ...String: LLCSPAAEDV SVVYQNGLPV ISVRLPSRRE R CQFTLKPL SDTVGVFLQQ LQAEDRGIDR VTIYSADGAR IASSTGIDIL LMDNFKLVIN DT SYLVQPP RRDLLPHEDG ERLNDVKILV QQLYTTLRIE EHQLNKEREL IGRLEDLNSQ LQP LEKVKE ELSKKAERRT TWVLWGGMAY MATQFGILAR LTWWEYSWDI MEPVTYFITY GTAM AMYAY FVLTRQEYLY PDARDRQYLL FFHRGAKRTR FDIEKYNKLK DAIAEAELDL KRLRD PLQL NLPIQQIDTS KD |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 5 mg/mL | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8.5 Component:
| |||||||||||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: AIR | |||||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 298 K / Instrument: FEI VITROBOT MARK IV / Details: 2 second blot, blot force of 0. | |||||||||||||||||||||
| Details | Monodisperse sample |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Digitization - Frames/image: 1-46 / Number grids imaged: 3 / Number real images: 3500 / Average exposure time: 0.13 sec. / Average electron dose: 2.2 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal magnification: 37000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)