+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: EMDB / ID: EMD-4134 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Structure of the mammalian rescue complex with Pelota and Hbs1l assembled on a truncated mRNA. | ||||||||||||
 マップデータ マップデータ | Postprocessed, sharpened map. | ||||||||||||
 試料 試料 |
| ||||||||||||
 キーワード キーワード | Translation / Elongation / Ribosome | ||||||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報stalled ribosome sensor activity / Dom34-Hbs1 complex / RNA surveillance / nuclear-transcribed mRNA catabolic process, no-go decay / mRNA decay by 3' to 5' exoribonuclease / nuclear-transcribed mRNA catabolic process, non-stop decay / mesenchymal to epithelial transition / endoderm development / positive regulation of BMP signaling pathway / ribosome disassembly ...stalled ribosome sensor activity / Dom34-Hbs1 complex / RNA surveillance / nuclear-transcribed mRNA catabolic process, no-go decay / mRNA decay by 3' to 5' exoribonuclease / nuclear-transcribed mRNA catabolic process, non-stop decay / mesenchymal to epithelial transition / endoderm development / positive regulation of BMP signaling pathway / ribosome disassembly / inner cell mass cell proliferation / nonfunctional rRNA decay / stem cell population maintenance / chromosome organization / 90S preribosome / ubiquitin ligase inhibitor activity / positive regulation of signal transduction by p53 class mediator / translation elongation factor activity / phagocytic cup / translation regulator activity / rough endoplasmic reticulum / ribosomal small subunit export from nucleus / gastrulation / MDM2/MDM4 family protein binding / rescue of stalled cytosolic ribosome / cytosolic ribosome / class I DNA-(apurinic or apyrimidinic site) endonuclease activity / DNA-(apurinic or apyrimidinic site) lyase / ribosomal large subunit biogenesis / maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / positive regulation of apoptotic signaling pathway / maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / maturation of SSU-rRNA / small-subunit processome / spindle / rRNA processing / regulation of translation / rhythmic process / positive regulation of canonical Wnt signaling pathway / antimicrobial humoral immune response mediated by antimicrobial peptide / large ribosomal subunit / ribosomal small subunit assembly / ribosome binding / ribosomal small subunit biogenesis / 5S rRNA binding / small ribosomal subunit / ribosomal large subunit assembly / small ribosomal subunit rRNA binding / cytosolic small ribosomal subunit / large ribosomal subunit rRNA binding / killing of cells of another organism / cytosolic large ribosomal subunit / defense response to Gram-negative bacterium / 加水分解酵素; 酸無水物に作用; GTPに作用・細胞または細胞小器官の運動に関与 / perikaryon / cell differentiation / cytoplasmic translation / tRNA binding / mitochondrial inner membrane / postsynaptic density / rRNA binding / structural constituent of ribosome / ribosome / translation / ribonucleoprotein complex / cell division / DNA repair / mRNA binding / GTPase activity / apoptotic process / centrosome / synapse / dendrite / GTP binding / nucleolus / perinuclear region of cytoplasm / endoplasmic reticulum / Golgi apparatus / signal transduction / DNA binding / RNA binding / extracellular exosome / zinc ion binding / membrane / metal ion binding / nucleus / cytoplasm / cytosol 類似検索 - 分子機能 | ||||||||||||
| 生物種 |   Homo sapiens (ヒト) Homo sapiens (ヒト) | ||||||||||||
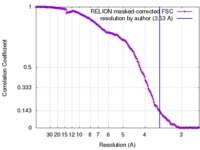
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 3.53 Å | ||||||||||||
 データ登録者 データ登録者 | Shao S / Murray J | ||||||||||||
| 資金援助 |  英国, 3件 英国, 3件
| ||||||||||||
 引用 引用 |  ジャーナル: Cell / 年: 2016 ジャーナル: Cell / 年: 2016タイトル: Decoding Mammalian Ribosome-mRNA States by Translational GTPase Complexes. 著者: Sichen Shao / Jason Murray / Alan Brown / Jack Taunton / V Ramakrishnan / Ramanujan S Hegde /   要旨: In eukaryotes, accurate protein synthesis relies on a family of translational GTPases that pair with specific decoding factors to decipher the mRNA code on ribosomes. We present structures of the ...In eukaryotes, accurate protein synthesis relies on a family of translational GTPases that pair with specific decoding factors to decipher the mRNA code on ribosomes. We present structures of the mammalian ribosome engaged with decoding factor⋅GTPase complexes representing intermediates of translation elongation (aminoacyl-tRNA⋅eEF1A), termination (eRF1⋅eRF3), and ribosome rescue (Pelota⋅Hbs1l). Comparative analyses reveal that each decoding factor exploits the plasticity of the ribosomal decoding center to differentially remodel ribosomal proteins and rRNA. This leads to varying degrees of large-scale ribosome movements and implies distinct mechanisms for communicating information from the decoding center to each GTPase. Additional structural snapshots of the translation termination pathway reveal the conformational changes that choreograph the accommodation of decoding factors into the peptidyl transferase center. Our results provide a structural framework for how different states of the mammalian ribosome are selectively recognized by the appropriate decoding factor⋅GTPase complex to ensure translational fidelity. | ||||||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_4134.map.gz emd_4134.map.gz | 13.9 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-4134-v30.xml emd-4134-v30.xml emd-4134.xml emd-4134.xml | 115.2 KB 115.2 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_4134_fsc.xml emd_4134_fsc.xml | 14.7 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_4134.png emd_4134.png | 189 KB | ||
| Filedesc metadata |  emd-4134.cif.gz emd-4134.cif.gz | 22.8 KB | ||
| その他 |  emd_4134_half_map_1.map.gz emd_4134_half_map_1.map.gz emd_4134_half_map_2.map.gz emd_4134_half_map_2.map.gz | 247 MB 245.9 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4134 http://ftp.pdbj.org/pub/emdb/structures/EMD-4134 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4134 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4134 | HTTPS FTP |
-関連構造データ
| 関連構造データ |  5lzwMC  4129C  4130C  4131C  4132C  4133C  4135C  4136C  4137C  5lzsC  5lztC  5lzuC  5lzvC  5lzxC  5lzyC  5lzzC M: このマップから作成された原子モデル C: 同じ文献を引用 ( |
|---|---|
| 類似構造データ |
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_4134.map.gz / 形式: CCP4 / 大きさ: 282.6 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_4134.map.gz / 形式: CCP4 / 大きさ: 282.6 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Postprocessed, sharpened map. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.34 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
CCP4マップ ヘッダ情報:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-添付データ
-ハーフマップ: Half map 1.
| ファイル | emd_4134_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Half map 1. | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-ハーフマップ: Half map 2.
| ファイル | emd_4134_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Half map 2. | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
+全体 : Affinity-purified 80S ribosome-nascent chain complex reconstitute...
+超分子 #1: Affinity-purified 80S ribosome-nascent chain complex reconstitute...
+分子 #1: uL2
+分子 #2: uL3
+分子 #3: uL4
+分子 #4: uL18
+分子 #5: eL6
+分子 #6: uL30
+分子 #7: eL8
+分子 #8: uL6
+分子 #9: uL16
+分子 #10: uL5
+分子 #11: eL13
+分子 #12: eL14
+分子 #13: eL15
+分子 #14: uL13
+分子 #15: uL22
+分子 #16: eL18
+分子 #17: eL19
+分子 #18: eL20
+分子 #19: eL21
+分子 #20: eL22
+分子 #21: uL14
+分子 #22: eL24
+分子 #23: uL23
+分子 #24: uL24
+分子 #25: eL27
+分子 #26: uL15
+分子 #27: eL29
+分子 #28: eL30
+分子 #29: eL31
+分子 #30: eL32
+分子 #31: eL33
+分子 #32: eL34
+分子 #33: uL29
+分子 #34: eL36
+分子 #35: eL37
+分子 #36: eL38
+分子 #37: eL39
+分子 #38: eL40
+分子 #39: eL41
+分子 #40: eL42
+分子 #41: eL43
+分子 #42: eL28
+分子 #43: uL10
+分子 #44: uL11
+分子 #45: Nascent chain
+分子 #52: uS2
+分子 #53: eS1
+分子 #54: uS5
+分子 #55: uS3
+分子 #56: eS4
+分子 #57: uS7
+分子 #58: eS6
+分子 #59: eS7
+分子 #60: eS8
+分子 #61: uS4
+分子 #62: eS10
+分子 #63: uS17
+分子 #64: eS12
+分子 #65: uS15
+分子 #66: uS11
+分子 #67: uS19
+分子 #68: uS9
+分子 #69: eS17
+分子 #70: uS13
+分子 #71: eS19
+分子 #72: uS10
+分子 #73: eS21
+分子 #74: uS8
+分子 #75: uS12
+分子 #76: eS24
+分子 #77: eS25
+分子 #78: eS26
+分子 #79: eS27
+分子 #80: eS28
+分子 #81: uS14
+分子 #82: eS30
+分子 #83: eS31
+分子 #84: RACK1
+分子 #86: Pelota
+分子 #87: Hbs1l
+分子 #46: P-site tRNA
+分子 #47: E-site tRNA
+分子 #48: 28S ribosomal RNA
+分子 #49: 5S ribosomal RNA
+分子 #50: 5.8S ribosomal RNA
+分子 #51: 18S ribosomal RNA
+分子 #85: mRNA (truncated)
+分子 #88: MAGNESIUM ION
+分子 #89: ZINC ION
+分子 #90: PHOSPHOMETHYLPHOSPHONIC ACID GUANYLATE ESTER
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 濃度 | 0.2 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 緩衝液 | pH: 7.4 構成要素:
| |||||||||||||||
| グリッド | モデル: Quantifoil R2/2 / 材質: COPPER / メッシュ: 400 / 支持フィルム - 材質: CARBON / 支持フィルム - トポロジー: CONTINUOUS / 支持フィルム - Film thickness: 5 / 前処理 - タイプ: GLOW DISCHARGE | |||||||||||||||
| 凍結 | 凍結剤: ETHANE / チャンバー内湿度: 100 % / チャンバー内温度: 277 K / 装置: FEI VITROBOT MARK III 詳細: 3 ul aliquots were applied to the grid and incubated for 30 s, before blotting for 3s to remove excess solution.. |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: FEI FALCON II (4k x 4k) 検出モード: INTEGRATING / 実像数: 1815 / 平均露光時間: 1.0 sec. / 平均電子線量: 30.0 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | C2レンズ絞り径: 70.0 µm / 倍率(補正後): 104478 / 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD / Cs: 2.7 mm / 倍率(公称値): 59000 |
| 試料ステージ | 試料ホルダーモデル: FEI TITAN KRIOS AUTOGRID HOLDER ホルダー冷却材: NITROGEN |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
+ 画像解析
画像解析
-原子モデル構築 1
| 精密化 | 空間: RECIPROCAL / プロトコル: OTHER / 温度因子: 84.1 / 当てはまり具合の基準: FSCaverage |
|---|---|
| 得られたモデル |  PDB-5lzw: |
 ムービー
ムービー コントローラー
コントローラー

























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)