+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4086 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | RNA polymerase I-Rrn3 complex | |||||||||
 Map data Map data | Pol I-Rrn3 complex | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 7.7 Å | |||||||||
 Authors Authors | Torreira E / Louro JA / Gil-Carton D / Gallego O / Calvo O / Fernandez-Tornero C | |||||||||
 Citation Citation |  Journal: Elife / Year: 2017 Journal: Elife / Year: 2017Title: The dynamic assembly of distinct RNA polymerase I complexes modulates rDNA transcription. Authors: Eva Torreira / Jaime Alegrio Louro / Irene Pazos / Noelia González-Polo / David Gil-Carton / Ana Garcia Duran / Sébastien Tosi / Oriol Gallego / Olga Calvo / Carlos Fernández-Tornero /  Abstract: Cell growth requires synthesis of ribosomal RNA by RNA polymerase I (Pol I). Binding of initiation factor Rrn3 activates Pol I, fostering recruitment to ribosomal DNA promoters. This fundamental ...Cell growth requires synthesis of ribosomal RNA by RNA polymerase I (Pol I). Binding of initiation factor Rrn3 activates Pol I, fostering recruitment to ribosomal DNA promoters. This fundamental process must be precisely regulated to satisfy cell needs at any time. We present in vivo evidence that, when growth is arrested by nutrient deprivation, cells induce rapid clearance of Pol I-Rrn3 complexes, followed by the assembly of inactive Pol I homodimers. This dual repressive mechanism reverts upon nutrient addition, thus restoring cell growth. Moreover, Pol I dimers also form after inhibition of either ribosome biogenesis or protein synthesis. Our mutational analysis, based on the electron cryomicroscopy structures of monomeric Pol I alone and in complex with Rrn3, underscores the central role of subunits A43 and A14 in the regulation of differential Pol I complexes assembly and subsequent promoter association. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4086.map.gz emd_4086.map.gz | 2.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4086-v30.xml emd-4086-v30.xml emd-4086.xml emd-4086.xml | 13.7 KB 13.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_4086_fsc.xml emd_4086_fsc.xml | 5.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_4086.png emd_4086.png | 120.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4086 http://ftp.pdbj.org/pub/emdb/structures/EMD-4086 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4086 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4086 | HTTPS FTP |
-Validation report
| Summary document |  emd_4086_validation.pdf.gz emd_4086_validation.pdf.gz | 246.4 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_4086_full_validation.pdf.gz emd_4086_full_validation.pdf.gz | 245.5 KB | Display | |
| Data in XML |  emd_4086_validation.xml.gz emd_4086_validation.xml.gz | 8.9 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-4086 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-4086 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-4086 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-4086 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_4086.map.gz / Format: CCP4 / Size: 16.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4086.map.gz / Format: CCP4 / Size: 16.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Pol I-Rrn3 complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.77 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
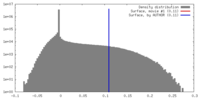
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : S. cerevisiae RNA polymerase I in complex with activating factor Rrn3
| Entire | Name: S. cerevisiae RNA polymerase I in complex with activating factor Rrn3 |
|---|---|
| Components |
|
-Supramolecule #1: S. cerevisiae RNA polymerase I in complex with activating factor Rrn3
| Supramolecule | Name: S. cerevisiae RNA polymerase I in complex with activating factor Rrn3 type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#14 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 594 KDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.06 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.8 / Component:
| |||||||||
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK III |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Detector mode: COUNTING / Digitization - Frames/image: 1-68 / Number grids imaged: 1 / Number real images: 1288 / Average exposure time: 4.0 sec. / Average electron dose: 68.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 79096 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 4.2 µm / Nominal defocus min: 1.9 µm / Nominal magnification: 47000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Overall B value: 537 |
|---|
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)