[English] 日本語
 Yorodumi
Yorodumi- EMDB-21113: Cryo EM Structure of VPS13 protein, 1-1390 from C. thermophilum -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-21113 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo EM Structure of VPS13 protein, 1-1390 from C. thermophilum | |||||||||
 Map data Map data | VPS13 protein | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Chaetomium thermophilum (fungus) Chaetomium thermophilum (fungus) | |||||||||
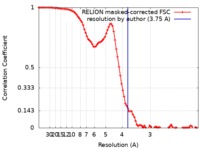
| Method | single particle reconstruction / cryo EM / Resolution: 3.75 Å | |||||||||
 Authors Authors | Li P / Lees JA / Reinisch KM | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: J Cell Biol / Year: 2020 Journal: J Cell Biol / Year: 2020Title: Cryo-EM reconstruction of a VPS13 fragment reveals a long groove to channel lipids between membranes. Authors: PeiQi Li / Joshua Aaron Lees / C Patrick Lusk / Karin M Reinisch /  Abstract: A single particle cryo-EM reconstruction of an ∼160-kD N-terminal fragment of the lipid transport protein VPS13 reveals an ∼160-Å long channel lined with hydrophobic residues suitable for ...A single particle cryo-EM reconstruction of an ∼160-kD N-terminal fragment of the lipid transport protein VPS13 reveals an ∼160-Å long channel lined with hydrophobic residues suitable for solubilizing multiple lipid fatty acid moieties. The structure suggests that VPS13 and related proteins, like the autophagy protein ATG2, can act as bridges between organelle membranes to allow bulk lipid flow between organelles. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_21113.map.gz emd_21113.map.gz | 55.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-21113-v30.xml emd-21113-v30.xml emd-21113.xml emd-21113.xml | 9.9 KB 9.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_21113_fsc.xml emd_21113_fsc.xml | 8.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_21113.png emd_21113.png | 94.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-21113 http://ftp.pdbj.org/pub/emdb/structures/EMD-21113 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21113 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21113 | HTTPS FTP |
-Validation report
| Summary document |  emd_21113_validation.pdf.gz emd_21113_validation.pdf.gz | 78 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_21113_full_validation.pdf.gz emd_21113_full_validation.pdf.gz | 77.1 KB | Display | |
| Data in XML |  emd_21113_validation.xml.gz emd_21113_validation.xml.gz | 492 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21113 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21113 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21113 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21113 | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_21113.map.gz / Format: CCP4 / Size: 59.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_21113.map.gz / Format: CCP4 / Size: 59.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | VPS13 protein | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.05 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
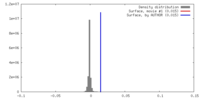
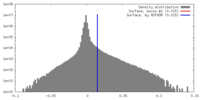
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Chaetomium thermophilum VPS13 1-1390
| Entire | Name: Chaetomium thermophilum VPS13 1-1390 |
|---|---|
| Components |
|
-Supramolecule #1: Chaetomium thermophilum VPS13 1-1390
| Supramolecule | Name: Chaetomium thermophilum VPS13 1-1390 / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Chaetomium thermophilum (fungus) Chaetomium thermophilum (fungus) |
| Molecular weight | Theoretical: 161.84 kDa/nm |
| Recombinant expression | Organism:  Homo sapiens (human) / Recombinant cell: Expi293F Homo sapiens (human) / Recombinant cell: Expi293F |
-Macromolecule #1: Vacuolar protein sorting-associate protein 13
| Macromolecule | Name: Vacuolar protein sorting-associate protein 13 / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Chaetomium thermophilum (fungus) Chaetomium thermophilum (fungus) |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MDYKDHDGDY KDHDIDYKDD DDKLAAA LE GLVAGLLNRF LGMYVKNFDP KQLKWEVWNG KVRLDNLELQ REALDQLKLP INVIKGHLGH LVLHIPW KT LASEQVKINI EDVFLLASPK EEAEYDEDEE ARRRHRLKME KLDSAELLKE RSQEGLSEEE QKRTQTFA Q ...String: MDYKDHDGDY KDHDIDYKDD DDKLAAA LE GLVAGLLNRF LGMYVKNFDP KQLKWEVWNG KVRLDNLELQ REALDQLKLP INVIKGHLGH LVLHIPW KT LASEQVKINI EDVFLLASPK EEAEYDEDEE ARRRHRLKME KLDSAELLKE RSQEGLSEEE QKRTQTFA Q ALVTKIVDNL QITIRNIHIR YEDAISAPGH PFALGITLEE FSAVSTDSDW TPAFITSIQS AHKLATLES LAIYWDTDAK LIGPGREPHE HSDQIPHDEM LKFFREMIAK GEADLSSEHQ FILKPVSGQA KIEIDKTGSH TVPRYKANL LFDEIGVVLD DQQYRDALMM VDLFHYFIRH QEYKKFQPKG VTPKEDPRAW FRFAGNAVLS K IHERNRRW SWDYFRERRD DRRRYIELFK KTKQNIQLTP EEREDLDKLE WKLSYEDLRF WRSLARNQLK KE NAEALKN KPPPQPQQQQ GWLSWVWGSK PVQPQQEEQQ GDENTRITEA QRKELYEVIQ WDEKAALAAE IDV PRDSVR LLIETSLSTG SFTLRQNPHG DARDLISLHF DLFRAKGLTR PDSFLIDISL GGFRVNDNTT PDSL YKEIV RVKDAPNTEG QKRYSIADLE LTVDEEAFFE LQVEQHPLDG QGDVAVTMKL KPLEIIWNPN VVVGI ADFF RPPERHMESI NALLETANAT VEGLRAQTRA GLQFALEEHK TVNAKLDLQA PLIILPESIT TPNSTC LIV DAGHISVNSE LVDKETMKQV QSTQDRPCTE EDLRRLEELM YDRFLVKLTS TQVLIGPSVE ITKQQLV QR DEKRQLHIVD QINLDFVVAM SILPKAPNLT KLKISGHLPV LQVNASDSKY KHLMRIIEVA IPKLYDVE P VLAPSGSHPT IRPRLASDAS TRSRRASFRK ASTQFLQFVS QQQEIVLDES DSDDDSEKFE DAKDTSVDE QLRIQQRIFD FKFTVDQLRG SLYRSDPEGK HPDQLLVELV AENFGVEYHL RPYDMSAMVS LGSVTMDDFV ENPPAEFKS IVSSGDIEDR KQARDLVRVK FVRVKKESPE FMSVYDGIET NVDVAISTIN LVVTRKTLLT L LDFILVTF SNPQPAAPVA TRMAVTDQES ETNIIVQPPP IESGPIRVKV DLKSIRMILN NDGIRLATLS FN HADVGVY ILGRYMRVSA KLGDLSLVDD VNLGVSEDSS LRQLVTIQGN ELADFRYEYF DPDKPEKNPG YDS SIYLRA GSVKVNFIEE PFRKIVDFLV KFGKMQAIYN AARMAAANQA QQLQQSQSRI KFDIVVKTPI VVFP RVVMS PKPKRDVITA YLGEIYAQNA FVPLDDSEKA DMAMKLTTGI RNIRLTSHFH YSEGRDEVLE LIDHV DLGF TIIYAEHKEG IKRPDLEIEG SMSDFNLRIT PYQLSALLAI SQSVPTVFAA DVEQHT |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)