[English] 日本語
 Yorodumi
Yorodumi- EMDB-20832: Cryo-EM structure of the PrgHK periplasmic ring from the Salmonel... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-20832 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the PrgHK periplasmic ring from the Salmonella SPI-1 type III secretion needle complex solved at 3.3 angstrom resolution | |||||||||
 Map data Map data | primary map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | BACTERIAL NANOMACHINE / TYPE III SECRETION SYSTEM / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology information | |||||||||
| Biological species |  Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) | |||||||||
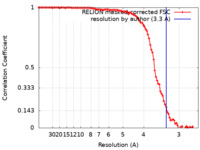
| Method | single particle reconstruction / cryo EM / Resolution: 3.3 Å | |||||||||
 Authors Authors | Butan C / Galan J | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2019 Journal: Proc Natl Acad Sci U S A / Year: 2019Title: High-resolution view of the type III secretion export apparatus in situ reveals membrane remodeling and a secretion pathway. Authors: Carmen Butan / Maria Lara-Tejero / Wenwei Li / Jun Liu / Jorge E Galán /  Abstract: Type III protein secretion systems are essential virulence factors for many important pathogenic bacteria. The entire protein secretion machine is composed of several substructures that organize into ...Type III protein secretion systems are essential virulence factors for many important pathogenic bacteria. The entire protein secretion machine is composed of several substructures that organize into a holostructure or injectisome. The core component of the injectisome is the needle complex, which houses the export apparatus that serves as a gate for the passage of the secreted proteins through the bacterial inner membrane. Here, we describe a high-resolution structure of the export apparatus of the type III secretion system in association with the needle complex and the underlying bacterial membrane, both in isolation and in situ. We show the precise location of the core export apparatus components within the injectisome and bacterial envelope and demonstrate that their deployment results in major membrane remodeling and thinning, which may be central for the protein translocation process. We also show that InvA, a critical export apparatus component, forms a multiring cytoplasmic conduit that provides a pathway for the type III secretion substrates to reach the entrance of the export gate. Combined with structure-guided mutagenesis, our studies provide major insight into potential mechanisms of protein translocation and injectisome assembly. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_20832.map.gz emd_20832.map.gz | 78.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-20832-v30.xml emd-20832-v30.xml emd-20832.xml emd-20832.xml | 11.3 KB 11.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_20832_fsc.xml emd_20832_fsc.xml | 9.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_20832.png emd_20832.png | 288.1 KB | ||
| Filedesc metadata |  emd-20832.cif.gz emd-20832.cif.gz | 5.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-20832 http://ftp.pdbj.org/pub/emdb/structures/EMD-20832 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20832 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20832 | HTTPS FTP |
-Validation report
| Summary document |  emd_20832_validation.pdf.gz emd_20832_validation.pdf.gz | 644.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_20832_full_validation.pdf.gz emd_20832_full_validation.pdf.gz | 644.3 KB | Display | |
| Data in XML |  emd_20832_validation.xml.gz emd_20832_validation.xml.gz | 11.5 KB | Display | |
| Data in CIF |  emd_20832_validation.cif.gz emd_20832_validation.cif.gz | 15.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20832 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20832 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20832 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20832 | HTTPS FTP |
-Related structure data
| Related structure data |  6uotMC  6uovC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_20832.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_20832.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | primary map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.32 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Complex of the periplasmic domains of PrgH and PrgK from the Salm...
| Entire | Name: Complex of the periplasmic domains of PrgH and PrgK from the Salmonella SPI-1 type III secretion needle complex |
|---|---|
| Components |
|
-Supramolecule #1: Complex of the periplasmic domains of PrgH and PrgK from the Salm...
| Supramolecule | Name: Complex of the periplasmic domains of PrgH and PrgK from the Salmonella SPI-1 type III secretion needle complex type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) |
-Macromolecule #1: Protein PrgH
| Macromolecule | Name: Protein PrgH / type: protein_or_peptide / ID: 1 / Number of copies: 24 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) |
| Molecular weight | Theoretical: 44.509367 KDa |
| Sequence | String: METSKEKTIT SPGPYIVRLL NSSLNGCEFP LLTGRTLFVV GQSDALTASG QLPDIPADSF FIPLDHGGVN FEIQVDTDAT EIILHELKE GNSESRSVQL NTPIQVGELL ILIRPESEPW VPEQPEKLET SAKKNEPRFK NGIVAALAGF FILGIGTVGT L WILNSPQR ...String: METSKEKTIT SPGPYIVRLL NSSLNGCEFP LLTGRTLFVV GQSDALTASG QLPDIPADSF FIPLDHGGVN FEIQVDTDAT EIILHELKE GNSESRSVQL NTPIQVGELL ILIRPESEPW VPEQPEKLET SAKKNEPRFK NGIVAALAGF FILGIGTVGT L WILNSPQR QAAELDSLLG QEKERFQVLP GRDKMLYVAA QNERDTLWAR QVLARGDYDK NARVINENEE NKRISIWLDT YY PQLAYYR IHFDEPRKPV FWLSRQRNTM SKKELEVLSQ KLRALMPYAD SVNITLMDDV TAAGQAEAGL KQQALPYSRR NHK GGVTFV IQGALDDVEI LRARQFVDSY YRTWGGRYVQ FAIELKDDWL KGRSFQYGAE GYIKMSPGHW YFPSPL UniProtKB: Protein PrgH |
-Macromolecule #2: Lipoprotein PrgK
| Macromolecule | Name: Lipoprotein PrgK / type: protein_or_peptide / ID: 2 / Number of copies: 24 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) |
| Molecular weight | Theoretical: 28.245287 KDa |
| Sequence | String: MIRRYLYTFL LVMTLAGCKD KDLLKGLDQE QANEVIAVLQ MHNIEANKID SGKLGYSITV AEPDFTAAVY WIKTYQLPPR PRVEIAQMF PADSLVSSPR AEKARLYSAI EQRLEQSLQT MEGVLSARVH ISYDIDAGEN GRPPKPVHLS ALAVYERGSP L AHQISDIK ...String: MIRRYLYTFL LVMTLAGCKD KDLLKGLDQE QANEVIAVLQ MHNIEANKID SGKLGYSITV AEPDFTAAVY WIKTYQLPPR PRVEIAQMF PADSLVSSPR AEKARLYSAI EQRLEQSLQT MEGVLSARVH ISYDIDAGEN GRPPKPVHLS ALAVYERGSP L AHQISDIK RFLKNSFADV DYDNISVVLS ERSDAQLQAP GTPVKRNSFA TSWIVLIILL SVMSAGFGVW YYKNHYARNK KG ITADDKA KSSNE UniProtKB: Lipoprotein PrgK |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 48.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller