[English] 日本語
 Yorodumi
Yorodumi- EMDB-1557: Cryo-EM density map of Epsilon-15 bacteriophage icosahedral recon... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1557 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM density map of Epsilon-15 bacteriophage icosahedral reconstruction from JADAS automatically acquired data at 7.3 Angstrom resolution. | |||||||||
 Map data Map data | This is the density map of the epsilon-15 bacteriophage reconstructed from JADAS automatically acquired images. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Single-particle / Automated data collection / Reconstruction / Cryo-EM / Epsilon-15 bacteriophage | |||||||||
| Biological species |   Salmonella phage epsilon15 (virus) Salmonella phage epsilon15 (virus) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 7.3 Å | |||||||||
 Authors Authors | Zhang J / Nakamura N / Shimizu Y / Liang N / Liu X / Jakana J / Marsh MP / Booth CR / Shinkawa T / Nakata M / Chiu W | |||||||||
 Citation Citation |  Journal: J Struct Biol / Year: 2009 Journal: J Struct Biol / Year: 2009Title: JADAS: a customizable automated data acquisition system and its application to ice-embedded single particles. Authors: Junjie Zhang / Natsuko Nakamura / Yuko Shimizu / Nathan Liang / Xiangan Liu / Joanita Jakana / Michael P Marsh / Christopher R Booth / Takao Shinkawa / Munetaka Nakata / Wah Chiu /  Abstract: The JEOL Automated Data Acquisition System (JADAS) is a software system built for the latest generation of the JEOL Transmission Electron Microscopes. It is designed to partially or fully automate ...The JEOL Automated Data Acquisition System (JADAS) is a software system built for the latest generation of the JEOL Transmission Electron Microscopes. It is designed to partially or fully automate image acquisition for ice-embedded single particles under low dose conditions. Its built-in flexibility permits users to customize the order of various imaging operations. In this paper, we describe how JADAS is used to accurately locate and image suitable specimen areas on a grid of ice-embedded particles. We also demonstrate the utility of JADAS by imaging the epsilon 15 bacteriophage with the JEM3200FSC electron cryo-microscope, showing that sufficient images can be collected in a single 8h session to yield a subnanometer resolution structure which agrees with the previously determined structure. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1557.map.gz emd_1557.map.gz | 38.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1557-v30.xml emd-1557-v30.xml emd-1557.xml emd-1557.xml | 9.4 KB 9.4 KB | Display Display |  EMDB header EMDB header |
| Images |  1557.jpg 1557.jpg | 282.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1557 http://ftp.pdbj.org/pub/emdb/structures/EMD-1557 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1557 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1557 | HTTPS FTP |
-Validation report
| Summary document |  emd_1557_validation.pdf.gz emd_1557_validation.pdf.gz | 262.4 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_1557_full_validation.pdf.gz emd_1557_full_validation.pdf.gz | 261.5 KB | Display | |
| Data in XML |  emd_1557_validation.xml.gz emd_1557_validation.xml.gz | 7.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1557 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1557 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1557 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1557 | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_1557.map.gz / Format: CCP4 / Size: 173.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1557.map.gz / Format: CCP4 / Size: 173.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | This is the density map of the epsilon-15 bacteriophage reconstructed from JADAS automatically acquired images. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.62 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
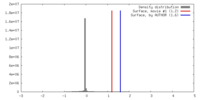
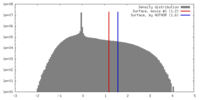
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Epsilon-15 bacteriophage
| Entire | Name: Epsilon-15 bacteriophage |
|---|---|
| Components |
|
-Supramolecule #1000: Epsilon-15 bacteriophage
| Supramolecule | Name: Epsilon-15 bacteriophage / type: sample / ID: 1000 Details: The sample is frozen on carbon support film in vitreous ice. Number unique components: 1 |
|---|---|
| Molecular weight | Experimental: 22 MDa / Theoretical: 22 MDa |
-Supramolecule #1: Salmonella phage epsilon15
| Supramolecule | Name: Salmonella phage epsilon15 / type: virus / ID: 1 / Name.synonym: Epsilon-15 bacteriophage Details: The sample is frozen on carbon support film in vitreous ice. NCBI-ID: 215158 / Sci species name: Salmonella phage epsilon15 / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: No / Syn species name: Epsilon-15 bacteriophage |
|---|---|
| Host (natural) | Organism:  Salmonella (bacteria) / synonym: BACTERIA(EUBACTERIA) Salmonella (bacteria) / synonym: BACTERIA(EUBACTERIA) |
| Molecular weight | Experimental: 22 MDa / Theoretical: 22 MDa |
| Virus shell | Shell ID: 1 / T number (triangulation number): 7 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Grid | Details: 400 mesh copper grid with holy carbon film |
|---|---|
| Vitrification | Cryogen name: ETHANE / Instrument: OTHER |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 3200FSC |
|---|---|
| Temperature | Average: 100 K |
| Specialist optics | Energy filter - Name: JEOL in-column omega filter |
| Details | Data is automatically collected using JEOL Automated Data Acquisition System (JADAS). Microscope used JEM3200FSC |
| Image recording | Category: CCD / Film or detector model: GENERIC GATAN (4k x 4k) / Digitization - Sampling interval: 2.62 µm / Number real images: 181 / Average electron dose: 20 e/Å2 / Bits/pixel: 8 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 56680 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 4.1 mm / Nominal magnification: 40000 |
| Sample stage | Specimen holder: JEM3200FSC cryoholder / Specimen holder model: OTHER |
- Image processing
Image processing
| CTF correction | Details: Each CCD image |
|---|---|
| Final reconstruction | Applied symmetry - Point group: I (icosahedral) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 7.3 Å / Resolution method: OTHER / Software - Name: Multi-path simulated annealing / Number images used: 7543 |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)