+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the human CCAN CENP-A alpha-satellite complex | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | Chromosome / kinetochore / cell division / centromere / CELL CYCLE | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationspindle attachment to meiosis I kinetochore / centromeric DNA binding / CENP-A containing chromatin assembly / protein localization to chromosome, centromeric region / kinetochore assembly / inner kinetochore / condensed chromosome, centromeric region / attachment of mitotic spindle microtubules to kinetochore / establishment of mitotic spindle orientation / chromosome, centromeric region ...spindle attachment to meiosis I kinetochore / centromeric DNA binding / CENP-A containing chromatin assembly / protein localization to chromosome, centromeric region / kinetochore assembly / inner kinetochore / condensed chromosome, centromeric region / attachment of mitotic spindle microtubules to kinetochore / establishment of mitotic spindle orientation / chromosome, centromeric region / mitotic cytokinesis / negative regulation of megakaryocyte differentiation / protein localization to CENP-A containing chromatin / pericentric heterochromatin / Amplification of signal from unattached kinetochores via a MAD2 inhibitory signal / Replacement of protamines by nucleosomes in the male pronucleus / CENP-A containing nucleosome / heterochromatin organization / Packaging Of Telomere Ends / Mitotic Prometaphase / Recognition and association of DNA glycosylase with site containing an affected purine / Cleavage of the damaged purine / EML4 and NUDC in mitotic spindle formation / Deposition of new CENPA-containing nucleosomes at the centromere / nucleosomal DNA binding / Recognition and association of DNA glycosylase with site containing an affected pyrimidine / Cleavage of the damaged pyrimidine / Resolution of Sister Chromatid Cohesion / Inhibition of DNA recombination at telomere / Meiotic synapsis / telomere organization / RNA Polymerase I Promoter Opening / Assembly of the ORC complex at the origin of replication / SUMOylation of chromatin organization proteins / DNA methylation / Condensation of Prophase Chromosomes / ERCC6 (CSB) and EHMT2 (G9a) positively regulate rRNA expression / SIRT1 negatively regulates rRNA expression / Chromatin modifications during the maternal to zygotic transition (MZT) / HCMV Late Events / PRC2 methylates histones and DNA / innate immune response in mucosa / Defective pyroptosis / chromosome segregation / HDACs deacetylate histones / RHO GTPases Activate Formins / RNA Polymerase I Promoter Escape / Nonhomologous End-Joining (NHEJ) / Transcriptional regulation by small RNAs / Formation of the beta-catenin:TCF transactivating complex / RUNX1 regulates genes involved in megakaryocyte differentiation and platelet function / NoRC negatively regulates rRNA expression / Activated PKN1 stimulates transcription of AR (androgen receptor) regulated genes KLK2 and KLK3 / B-WICH complex positively regulates rRNA expression / G2/M DNA damage checkpoint / HDMs demethylate histones / DNA Damage/Telomere Stress Induced Senescence / Metalloprotease DUBs / PKMTs methylate histone lysines / Meiotic recombination / kinetochore / RMTs methylate histone arginines / Pre-NOTCH Transcription and Translation / Activation of anterior HOX genes in hindbrain development during early embryogenesis / HCMV Early Events / Transcriptional regulation of granulopoiesis / structural constituent of chromatin / Separation of Sister Chromatids / antimicrobial humoral immune response mediated by antimicrobial peptide / UCH proteinases / nucleosome / nucleosome assembly / E3 ubiquitin ligases ubiquitinate target proteins / Recruitment and ATM-mediated phosphorylation of repair and signaling proteins at DNA double strand breaks / chromatin organization / RUNX1 regulates transcription of genes involved in differentiation of HSCs / mitotic cell cycle / HATs acetylate histones / Processing of DNA double-strand break ends / midbody / antibacterial humoral response / Senescence-Associated Secretory Phenotype (SASP) / Oxidative Stress Induced Senescence / Estrogen-dependent gene expression / chromosome, telomeric region / nuclear body / Ub-specific processing proteases / defense response to Gram-positive bacterium / protein heterodimerization activity / Amyloid fiber formation / negative regulation of cell population proliferation / cell division / chromatin binding / protein-containing complex / DNA binding / RNA binding / extracellular space / extracellular exosome / extracellular region / nucleoplasm Similarity search - Function | ||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.44 Å | ||||||||||||
 Authors Authors | Yatskevich S / Muir KW | ||||||||||||
| Funding support |  United Kingdom, United Kingdom,  Germany, 3 items Germany, 3 items
| ||||||||||||
 Citation Citation |  Journal: Science / Year: 2022 Journal: Science / Year: 2022Title: Structure of the human inner kinetochore bound to a centromeric CENP-A nucleosome. Authors: Stanislau Yatskevich / Kyle W Muir / Dom Bellini / Ziguo Zhang / Jing Yang / Thomas Tischer / Masa Predin / Tom Dendooven / Stephen H McLaughlin / David Barford /  Abstract: Kinetochores assemble onto specialized centromeric CENP-A (centromere protein A) nucleosomes (CENP-A) to mediate attachments between chromosomes and the mitotic spindle. We describe cryo-electron ...Kinetochores assemble onto specialized centromeric CENP-A (centromere protein A) nucleosomes (CENP-A) to mediate attachments between chromosomes and the mitotic spindle. We describe cryo-electron microscopy structures of the human inner kinetochore constitutive centromere associated network (CCAN) complex bound to CENP-A reconstituted onto α-satellite DNA. CCAN forms edge-on contacts with CENP-A, and a linker DNA segment of the α-satellite repeat emerges from the fully wrapped end of the nucleosome to thread through the central CENP-LN channel that tightly grips the DNA. The CENP-TWSX histone-fold module further augments DNA binding and partially wraps the linker DNA in a manner reminiscent of canonical nucleosomes. Our study suggests that the topological entrapment of the linker DNA by CCAN provides a robust mechanism by which kinetochores withstand both pushing and pulling forces exerted by the mitotic spindle. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14334.map.gz emd_14334.map.gz | 91.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14334-v30.xml emd-14334-v30.xml emd-14334.xml emd-14334.xml | 17.1 KB 17.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_14334_fsc.xml emd_14334_fsc.xml | 12.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_14334.png emd_14334.png | 56.4 KB | ||
| Filedesc metadata |  emd-14334.cif.gz emd-14334.cif.gz | 6.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14334 http://ftp.pdbj.org/pub/emdb/structures/EMD-14334 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14334 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14334 | HTTPS FTP |
-Validation report
| Summary document |  emd_14334_validation.pdf.gz emd_14334_validation.pdf.gz | 475.7 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_14334_full_validation.pdf.gz emd_14334_full_validation.pdf.gz | 475.2 KB | Display | |
| Data in XML |  emd_14334_validation.xml.gz emd_14334_validation.xml.gz | 13.2 KB | Display | |
| Data in CIF |  emd_14334_validation.cif.gz emd_14334_validation.cif.gz | 17.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14334 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14334 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14334 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14334 | HTTPS FTP |
-Related structure data
| Related structure data |  7r5rMC  7pb4C  7pb8C  7piiC  7pknC  7r5sC  7r5vC  7ywxC  7yyhC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_14334.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14334.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.853 Å | ||||||||||||||||||||||||||||||||||||
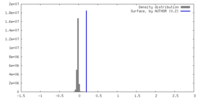
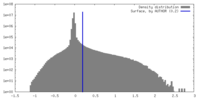
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : Human CCAN complex with bound alpha-satellite DNA
| Entire | Name: Human CCAN complex with bound alpha-satellite DNA |
|---|---|
| Components |
|
-Supramolecule #1: Human CCAN complex with bound alpha-satellite DNA
| Supramolecule | Name: Human CCAN complex with bound alpha-satellite DNA / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#7 |
|---|
-Macromolecule #1: Histone H3-like centromeric protein A
| Macromolecule | Name: Histone H3-like centromeric protein A / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 16.02363 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGPRRRSRKP EAPRRRSPSP TPTPGPSRRG PSLGASSHQH SRRRQGWLKE IRKLQKSTHL LIRKLPFSRL AREICVKFTR GVDFNWQAQ ALLALQEAAE AFLVHLFEDA YLLTLHAGRV TLFPKDVQLA RRIRGLEEGL G UniProtKB: Histone H3-like centromeric protein A |
-Macromolecule #2: Histone H4
| Macromolecule | Name: Histone H4 / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 11.394426 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSGRGKGGKG LGKGGAKRHR KVLRDNIQGI TKPAIRRLAR RGGVKRISGL IYEETRGVLK VFLENVIRDA VTYTEHAKRK TVTAMDVVY ALKRQGRTLY GFGG UniProtKB: Histone H4 |
-Macromolecule #3: Histone H2A type 1-C
| Macromolecule | Name: Histone H2A type 1-C / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 14.135523 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSGRGKQGGK ARAKAKSRSS RAGLQFPVGR VHRLLRKGNY AERVGAGAPV YLAAVLEYLT AEILELAGNA ARDNKKTRII PRHLQLAIR NDEELNKLLG RVTIAQGGVL PNIQAVLLPK KTESHHKAKG K UniProtKB: Histone H2A type 1-C |
-Macromolecule #4: Histone H2B type 1-C/E/F/G/I
| Macromolecule | Name: Histone H2B type 1-C/E/F/G/I / type: protein_or_peptide / ID: 4 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 13.937213 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MPEPAKSAPA PKKGSKKAVT KAQKKDGKKR KRSRKESYSV YVYKVLKQVH PDTGISSKAM GIMNSFVNDI FERIAGEASR LAHYNKRST ITSREIQTAV RLLLPGELAK HAVSEGTKAV TKYTSSK UniProtKB: Histone H2B type 1-C/E/F/G/I |
-Macromolecule #7: Centromere protein C
| Macromolecule | Name: Centromere protein C / type: protein_or_peptide / ID: 7 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 61.724816 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: AASGLDHLKN GYRRRFCRPS RARDINTEQG QNVLEILQDC FEEKSLANDF STNSTKSVPN STRKIKDTCI QSPSKECQKS HPKSVPVSS KKKEASLQFV VEPSEATNRS VQAHEVHQKI LATDVSSKNT PDSKKISSRN INDHHSEADE EFYLSVGSPS V LLDAKTSV ...String: AASGLDHLKN GYRRRFCRPS RARDINTEQG QNVLEILQDC FEEKSLANDF STNSTKSVPN STRKIKDTCI QSPSKECQKS HPKSVPVSS KKKEASLQFV VEPSEATNRS VQAHEVHQKI LATDVSSKNT PDSKKISSRN INDHHSEADE EFYLSVGSPS V LLDAKTSV SQNVIPSSAQ KRETYTFENS VNMLPSSTEV SVKTKKRLNF DDKVMLKKIE IDNKVSDEED KTSEGQERKP SG SSQNRIR DSEYEIQRQA KKSFSTLFLE TVKRKSESSP IVRHAATAPP HSCPPDDTKL IEDEFIIDES DQSFASRSWI TIP RKAGSL KQRTISPAES TALLQGRKSR EKHHNILPKT LANDKHSHKP HPVETSQPSD KTVLDTSYAL IGETVNNYRS TKYE MYSKN AEKPSRSKRT IKQKQRRKFM AKPAEEQLDV GQSKDENIHT SHITQDEFQR NSDRNMEEHE EMGNDCVSKK QMPPV GSKK SSTRKDKEES KKKRFSSESK NKLVPEEVTS TVTKSRRISR RPSDWWVVKS EESPVYSNS UniProtKB: Centromere protein C |
-Macromolecule #5: DNA (171-MER)
| Macromolecule | Name: DNA (171-MER) / type: dna / ID: 5 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 52.725812 KDa |
| Sequence | String: (DT)(DC)(DC)(DA)(DA)(DA)(DT)(DG)(DT)(DC) (DC)(DA)(DA)(DT)(DT)(DC)(DC)(DA)(DG)(DA) (DT)(DA)(DC)(DT)(DA)(DC)(DA)(DA)(DA) (DA)(DA)(DG)(DA)(DG)(DT)(DG)(DT)(DT)(DT) (DC) (DA)(DA)(DA)(DA)(DC)(DT) ...String: (DT)(DC)(DC)(DA)(DA)(DA)(DT)(DG)(DT)(DC) (DC)(DA)(DA)(DT)(DT)(DC)(DC)(DA)(DG)(DA) (DT)(DA)(DC)(DT)(DA)(DC)(DA)(DA)(DA) (DA)(DA)(DG)(DA)(DG)(DT)(DG)(DT)(DT)(DT) (DC) (DA)(DA)(DA)(DA)(DC)(DT)(DG)(DC) (DT)(DC)(DT)(DA)(DT)(DG)(DA)(DA)(DA)(DA) (DG)(DG) (DA)(DA)(DT)(DG)(DT)(DT)(DC) (DA)(DA)(DC)(DT)(DC)(DT)(DA)(DT)(DG)(DA) (DG)(DT)(DT) (DG)(DA)(DA)(DT)(DG)(DC) (DA)(DA)(DA)(DC)(DA)(DT)(DC)(DA)(DC)(DA) (DT)(DA)(DG)(DA) (DA)(DG)(DT)(DT)(DT) (DC)(DT)(DG)(DA)(DG)(DA)(DA)(DT)(DG)(DC) (DT)(DT)(DC)(DT)(DG) (DT)(DC)(DT)(DA) (DG)(DT)(DT)(DT)(DT)(DT)(DA)(DT)(DG)(DT) (DG)(DA)(DA)(DC)(DA)(DT) (DA)(DT)(DT) (DC)(DC)(DC)(DG)(DT)(DT)(DT)(DC)(DC)(DA) (DA)(DC)(DG)(DA)(DA)(DG)(DG) (DC)(DC) (DT)(DC)(DA)(DA)(DA)(DG)(DC)(DG)(DG) |
-Macromolecule #6: DNA (171-MER)
| Macromolecule | Name: DNA (171-MER) / type: dna / ID: 6 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 52.822809 KDa |
| Sequence | String: (DC)(DC)(DG)(DC)(DT)(DT)(DT)(DG)(DA)(DG) (DG)(DC)(DC)(DT)(DT)(DC)(DG)(DT)(DT)(DG) (DG)(DA)(DA)(DA)(DC)(DG)(DG)(DG)(DA) (DA)(DT)(DA)(DT)(DG)(DT)(DT)(DC)(DA)(DC) (DA) (DT)(DA)(DA)(DA)(DA)(DA) ...String: (DC)(DC)(DG)(DC)(DT)(DT)(DT)(DG)(DA)(DG) (DG)(DC)(DC)(DT)(DT)(DC)(DG)(DT)(DT)(DG) (DG)(DA)(DA)(DA)(DC)(DG)(DG)(DG)(DA) (DA)(DT)(DA)(DT)(DG)(DT)(DT)(DC)(DA)(DC) (DA) (DT)(DA)(DA)(DA)(DA)(DA)(DC)(DT) (DA)(DG)(DA)(DC)(DA)(DG)(DA)(DA)(DG)(DC) (DA)(DT) (DT)(DC)(DT)(DC)(DA)(DG)(DA) (DA)(DA)(DC)(DT)(DT)(DC)(DT)(DA)(DT)(DG) (DT)(DG)(DA) (DT)(DG)(DT)(DT)(DT)(DG) (DC)(DA)(DT)(DT)(DC)(DA)(DA)(DC)(DT)(DC) (DA)(DT)(DA)(DG) (DA)(DG)(DT)(DT)(DG) (DA)(DA)(DC)(DA)(DT)(DT)(DC)(DC)(DT)(DT) (DT)(DT)(DC)(DA)(DT) (DA)(DG)(DA)(DG) (DC)(DA)(DG)(DT)(DT)(DT)(DT)(DG)(DA)(DA) (DA)(DC)(DA)(DC)(DT)(DC) (DT)(DT)(DT) (DT)(DT)(DG)(DT)(DA)(DG)(DT)(DA)(DT)(DC) (DT)(DG)(DG)(DA)(DA)(DT)(DT) (DG)(DG) (DA)(DC)(DA)(DT)(DT)(DT)(DG)(DG)(DA) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)