[English] 日本語
 Yorodumi
Yorodumi- EMDB-10464: Structure of Drosophila melanogaster Dispatched bound to a modifi... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-10464 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of Drosophila melanogaster Dispatched bound to a modified Hedgehog ligand, HhN-C85II | |||||||||
 Map data Map data | Drosophila melanogaster Dispatched bound to modified Hedgehog ligand HhN-C85II | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | RND transporter / Dispatched / Hedgehog / transmembrane domain / ectodomain / cholesteryl hemisuccinate / detergent micelle / digitonin / monomer / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationprogression of morphogenetic furrow involved in compound eye morphogenesis / terminal cell fate specification, open tracheal system / cytoneme assembly / germ cell attraction / wing disc proximal/distal pattern formation / labial disc development / regulation of cell proliferation involved in compound eye morphogenesis / Bolwig's organ morphogenesis / Release of Hh-Np from the secreting cell / Ligand-receptor interactions ...progression of morphogenetic furrow involved in compound eye morphogenesis / terminal cell fate specification, open tracheal system / cytoneme assembly / germ cell attraction / wing disc proximal/distal pattern formation / labial disc development / regulation of cell proliferation involved in compound eye morphogenesis / Bolwig's organ morphogenesis / Release of Hh-Np from the secreting cell / Ligand-receptor interactions / leg disc morphogenesis / Formation and transport of the N-HH ligand / cytoneme / regulation of epithelial cell migration, open tracheal system / morphogenesis of larval imaginal disc epithelium / Assembly of the 'signalling complexes' / wing disc pattern formation / compound eye photoreceptor cell differentiation / Hedgehog ligand biogenesis / gonadal mesoderm development / patched ligand maturation / analia development / anterior head segmentation / anterior/posterior lineage restriction, imaginal disc / epithelial cell migration, open tracheal system / compound eye morphogenesis / genital disc development / genital disc anterior/posterior pattern formation / posterior head segmentation / trunk segmentation / imaginal disc growth / spiracle morphogenesis, open tracheal system / wing disc anterior/posterior pattern formation / morphogen activity / hindgut morphogenesis / segment polarity determination / foregut morphogenesis / ventral midline development / cholesterol-protein transferase activity / mucosal immune response / imaginal disc-derived wing morphogenesis / compartment pattern specification / glial cell migration / negative regulation of homotypic cell-cell adhesion / developmental pigmentation / patched binding / self proteolysis / germ cell migration / embryonic pattern specification / intein-mediated protein splicing / positive regulation of protein localization to cell surface / cell fate specification / smoothened signaling pathway / positive regulation of neuroblast proliferation / protein autoprocessing / epidermis development / endocytic vesicle / negative regulation of proteolysis / regulation of mitotic cell cycle / cell-cell signaling / peptidase activity / heart development / regulation of gene expression / cytoplasmic vesicle / Hydrolases; Acting on ester bonds / endosome / calcium ion binding / : / extracellular region / membrane / nucleus / plasma membrane / cytosol Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.76 Å | |||||||||
 Authors Authors | Korkhov VM / Cannac F | |||||||||
| Funding support |  Switzerland, 1 items Switzerland, 1 items
| |||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2020 Journal: Sci Adv / Year: 2020Title: Cryo-EM structure of the Hedgehog release protein Dispatched. Authors: Fabien Cannac / Chao Qi / Julia Falschlunger / George Hausmann / Konrad Basler / Volodymyr M Korkhov /  Abstract: The Hedgehog (Hh) signaling pathway controls embryonic development and adult tissue homeostasis in multicellular organisms. In , the pathway is primed by secretion of a dually lipid-modified ...The Hedgehog (Hh) signaling pathway controls embryonic development and adult tissue homeostasis in multicellular organisms. In , the pathway is primed by secretion of a dually lipid-modified morphogen, Hh, a process dependent on a membrane-integral protein Dispatched. Although Dispatched is a critical component of the pathway, the structural basis of its activity has, so far, not been described. Here, we describe a cryo-electron microscopy structure of the Dispatched at 3.2-Å resolution. The ectodomains of Dispatched adopt an open conformation suggestive of a receptor-chaperone role. A three-dimensional reconstruction of Dispatched bound to Hh confirms the ability of Dispatched to bind Hh but using a unique mode distinct from those previously observed in structures of Hh complexes. The structure may represent the state of the complex that precedes shedding of Hh from the surface of the morphogen-releasing cell. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_10464.map.gz emd_10464.map.gz | 99.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-10464-v30.xml emd-10464-v30.xml emd-10464.xml emd-10464.xml | 13.8 KB 13.8 KB | Display Display |  EMDB header EMDB header |
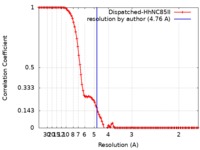
| FSC (resolution estimation) |  emd_10464_fsc.xml emd_10464_fsc.xml | 10.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_10464.png emd_10464.png | 61.8 KB | ||
| Filedesc metadata |  emd-10464.cif.gz emd-10464.cif.gz | 6.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-10464 http://ftp.pdbj.org/pub/emdb/structures/EMD-10464 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10464 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10464 | HTTPS FTP |
-Related structure data
| Related structure data |  6td6MC  6tbuC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_10464.map.gz / Format: CCP4 / Size: 107.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_10464.map.gz / Format: CCP4 / Size: 107.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Drosophila melanogaster Dispatched bound to modified Hedgehog ligand HhN-C85II | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.88 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : A complex of Dispatched and HhN-C85II in digitonin micelle
| Entire | Name: A complex of Dispatched and HhN-C85II in digitonin micelle |
|---|---|
| Components |
|
-Supramolecule #1: A complex of Dispatched and HhN-C85II in digitonin micelle
| Supramolecule | Name: A complex of Dispatched and HhN-C85II in digitonin micelle type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|
-Supramolecule #2: Drosophila melanogaster protein Dispatched
| Supramolecule | Name: Drosophila melanogaster protein Dispatched / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #3: Drosophila melanogaster Hedgehog, HhN-C85II
| Supramolecule | Name: Drosophila melanogaster Hedgehog, HhN-C85II / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Protein dispatched
| Macromolecule | Name: Protein dispatched / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 139.149875 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MLCFDSERMN WYYHVLARRP YLVVVSIAVY CVACIIVALV LNKLPDFSDP TLGFETRGTK IGERLTAWYN LLQETDHHGA LFSNPSDLW ERRRVEQGYV ETKLHPNHRR RKNKHKNRNK NKRRKEQNQS SHEHHDVAQK MMQFKKRLKA TSSPSPNLGF D TWIGDSGV ...String: MLCFDSERMN WYYHVLARRP YLVVVSIAVY CVACIIVALV LNKLPDFSDP TLGFETRGTK IGERLTAWYN LLQETDHHGA LFSNPSDLW ERRRVEQGYV ETKLHPNHRR RKNKHKNRNK NKRRKEQNQS SHEHHDVAQK MMQFKKRLKA TSSPSPNLGF D TWIGDSGV FRDYEITNDS ASSSLEPTRR TEQIEYGHNT TSVDEEEHQQ RVQTKKSTWR LLKQAATLPT DGWADMHRRQ PI EGFFCDS SPRKEYSHFV VQRIGPNATD SLFDLNGLLA MCQLQDQITE VPSYRAFCEP EMLTTECCRP WSLPNYAAML ANK SSCFDL TTEDVTSLHT LLLGCYEYFH DLKMDNHCNE IPHCRAPEEC KRLNIVFNVL NFLTDFSFIK SNDSNVYLKY AMIF IPVAQ SNRLLPLFHE WEDVELINEL VEVVAMDLGL ENELFNELLL TDVWLVSLGG TFVMASVWLY TGSAFITLMS CVAIC FSLG LAYFFYAIVL EFEFFPYMNL LAVVVIIGIG ADDVFLFLKI WHCVLTERFS NRCTLTTQSQ SALPTLENSD HTESLE NIM ALTMRHAAAS MFVTSLTTAG AFYASYSSSI TAIKCFGIFA GTVVVTNYLL MITWLPASVS IMERLFATRM SCHHPMS IK LIHACKKSIN RFCQMFEECI TKSIMNYAYL WLLIFGALGA SSAVIVFWYP GLQLPEKSHF QLFVSKHPFE VYSSLKQQ F WFEKPLQAYY NFKMHMHFVW GVQAVDDGDY TNPNSYGHLH YDNNFNVSSR PAQLWILDFC QSVRQQPFYK ETLGMLLPN CFIENLIDYM KRRCIDDMDS TRKDRSPCCD AQFPFEPHIF EYCLPQSISN MYDTTFFRPG VAGPKFAEAP RLETEDYLGM SGNESAEYS TNGSFTPLLV KALVIEFESN VAYSTIYANI RQFYESVEHW FQMQLKTAPP ELQGGWFTSD LKFYNVQDTL S HDTFVAIC LAMAASLAVL LCFTVNILIS IYAVLTVSLS IFNTVAVLIL LGWQLNILES IAVSTAIGLA VDFSLHYGIH YR MSPVKER LAATQFVLSR IIGPTVMAAT TTGLAGGIMM ASNILPYIQI GVFLVVVMIV SWFYATFFLM SLLRVAGPQH GFL ELKWPL WSKRSSGSSK FYERKPSQVI ASEQLLTPTS SAIVELANSE THELESLNSN SLIKTISGIE SAHALSSLPR DFEH SFQTM HECKYQTYPS TSN UniProtKB: Protein dispatched |
-Macromolecule #2: Protein hedgehog
| Macromolecule | Name: Protein hedgehog / type: protein_or_peptide / ID: 2 Details: 6xHis-Sumo-tagged HhN-C85II, N terminal fragment of Hedgehog Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 52.217121 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MDNHSSVPWA SAASVTCLSL DAKCHSSSSS SSSKSAASSI SAIPQEETQT MRHIAHTQRC LSRLTSLVAL LLIVLPMVFS PAHSCGPGR GLGRHRARNL YPLVLKQTIP NLSEYTNSAS GPLEGVIRRD SPKFKDLVPN YNRDILFRDE EGTGADRLMS K RCKEKLNV ...String: MDNHSSVPWA SAASVTCLSL DAKCHSSSSS SSSKSAASSI SAIPQEETQT MRHIAHTQRC LSRLTSLVAL LLIVLPMVFS PAHSCGPGR GLGRHRARNL YPLVLKQTIP NLSEYTNSAS GPLEGVIRRD SPKFKDLVPN YNRDILFRDE EGTGADRLMS K RCKEKLNV LAYSVMNEWP GIRLLVTESW DEDYHHGQES LHYEGRAVTI ATSDRDQSKY GMLARLAVEA GFDWVSYVSR RH IYCSVKS DSSISSHVHG CFTPESTALL ESGVRKPLGE LSIGDRVLSM TANGQAVYSE VILFMDRNLE QMQNFVQLHT DGG AVLTVT PAHLVSVWQP ESQKLTFVFA DRIEEKNQVL VRDVETGELR PQRVVKVGSV RSKGVVAPLT REGTIVVNSV AASC YAVIN SQSLAHWGLA PMRLLSTLEA WLPAKEQLHS SPKVVSSAQQ QNGIHWYANA LYKVKDYVLP QSWRHD UniProtKB: Protein hedgehog |
-Macromolecule #4: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 4 / Number of copies: 1 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Average electron dose: 51.5 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)