4DJI
 
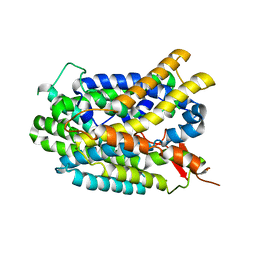 | | Structure of glutamate-GABA antiporter GadC | | Descriptor: | Probable glutamate/gamma-aminobutyrate antiporter | | Authors: | Ma, D, Lu, P.L, Yan, C.Y, Fan, C, Yin, P, Wang, J.W, Shi, Y.G. | | Deposit date: | 2012-02-02 | | Release date: | 2012-03-14 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (3.187 Å) | | Cite: | Structure and mechanism of a glutamate-GABA antiporter
Nature, 483, 2012
|
|
9XNH
 
 | | pilus-like-gamma, a bacteria pilus-Like structure obtained from a Karstcave from Guilin city, Guangxi ZhuangAutonomous Region, China | | Descriptor: | pilus-like-gamma | | Authors: | Hu, M.X, Zhang, Q, Chen, S, Ge, Q.J, Wang, J.W, Yan, N, Qin, L.J. | | Deposit date: | 2025-11-12 | | Release date: | 2026-02-18 | | Method: | ELECTRON MICROSCOPY (3.66 Å) | | Cite: | Absolute hand determination ofglycofibrils from natural sources incryo-EM
To Be Published
|
|
9XFV
 
 | | Pilus-like-alpha, a bacteria pilus-like structure obtained from a Karst cave from Guilin City, Guangxi Zhuang Autonomous Region, China | | Descriptor: | bacteria pilus (like) | | Authors: | Hu, M.X, Zhang, Q, Chen, S, Ge, Q.J, Wang, J.W, Yan, N, Qin, L.J. | | Deposit date: | 2025-10-29 | | Release date: | 2026-02-18 | | Method: | ELECTRON MICROSCOPY (3.07 Å) | | Cite: | Absolute hand determination of glycofibrils from natural sources in cryo-EM
To Be Published
|
|
9XNC
 
 | | pilus-like-beta, a bacteria pilus-like structure obtained from a Karst cave from Guilin City, Guangxi Zhuang Autonomous Region, China | | Descriptor: | Bacteria pilus (like) | | Authors: | Hu, M.X, Zhang, Q, Chen, S, Ge, Q.J, Wang, J.W, Yan, N, Qin, L.J. | | Deposit date: | 2025-11-12 | | Release date: | 2026-02-18 | | Method: | ELECTRON MICROSCOPY (3.37 Å) | | Cite: | Absolute hand determination of glycofibrils from natural sources in cryo-EM
To Be Published
|
|
4DXW
 
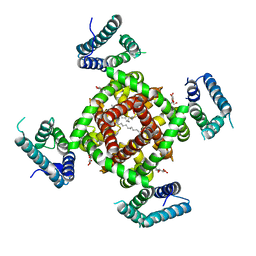 | | Crystal structure of NavRh, a voltage-gated sodium channel | | Descriptor: | 1,2-DIMYRISTOYL-SN-GLYCERO-3-PHOSPHOCHOLINE, CALCIUM ION, Ion transport protein, ... | | Authors: | Zhang, X, Ren, W.L, Yan, C.Y, Wang, J.W, Yan, N. | | Deposit date: | 2012-02-28 | | Release date: | 2012-05-23 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (3.052 Å) | | Cite: | Crystal structure of an orthologue of the NaChBac voltage-gated sodium channel
Nature, 486, 2012
|
|
3V6T
 
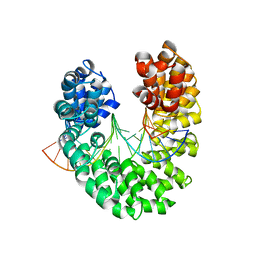 | | Crystal structure of the DNA-bound dHax3, a TAL effector, at 1.85 angstrom | | Descriptor: | DNA (5'-D(*AP*GP*AP*GP*AP*GP*AP*TP*AP*AP*AP*GP*GP*GP*AP*CP*A)-3'), DNA (5'-D(*TP*GP*TP*CP*CP*CP*TP*TP*TP*AP*TP*CP*TP*CP*TP*CP*T)-3'), dHax3 | | Authors: | Deng, D, Yan, C.Y, Pan, X.J, Wang, J.W, Yan, N, Shi, Y.G. | | Deposit date: | 2011-12-20 | | Release date: | 2012-01-18 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Structural Basis for Sequence-Specific Recognition of DNA by TAL Effectors
Science, 2012
|
|
3V6P
 
 | | Crystal structure of the DNA-binding domain of dHax3, a TAL effector | | Descriptor: | dHax3 | | Authors: | Deng, D, Yan, C.Y, Pan, X.J, Wang, J.W, Shi, Y.G, Yan, N. | | Deposit date: | 2011-12-20 | | Release date: | 2012-01-18 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.401 Å) | | Cite: | Structural Basis for Sequence-Specific Recognition of DNA by TAL Effectors
Science, 2012
|
|
4F1Z
 
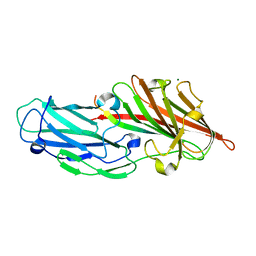 | | Crystal structures reveal the multi-ligand binding mechanism of the Staphylococcus aureus ClfB | | Descriptor: | Clumping factor B, MAGNESIUM ION, peptide from Keratin, ... | | Authors: | Yang, M.J, Xiang, H, Wang, J.W, Liu, B, Chen, Y.G, Liu, L, Deng, X.M, Feng, Y. | | Deposit date: | 2012-05-07 | | Release date: | 2012-08-08 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal Structures Reveal the Multi-Ligand Binding Mechanism of Staphylococcus aureus ClfB
Plos Pathog., 8, 2012
|
|
4F24
 
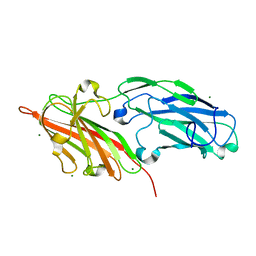 | | Crystal structures reveal the multi-ligand binding mechanism of the Staphylococcus aureus ClfB | | Descriptor: | Clumping factor B, MAGNESIUM ION | | Authors: | Yang, M.J, Xiang, H, Wang, J.W, Liu, B, Chen, Y.G, Liu, L, Deng, X.M, Feng, Y. | | Deposit date: | 2012-05-07 | | Release date: | 2012-08-08 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.509 Å) | | Cite: | Crystal Structures Reveal the Multi-Ligand Binding Mechanism of Staphylococcus aureus ClfB
Plos Pathog., 8, 2012
|
|
4F27
 
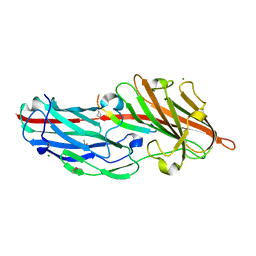 | | Crystal structures reveal the multi-ligand binding mechanism of the Staphylococcus aureus ClfB | | Descriptor: | Clumping factor B, MAGNESIUM ION, peptide from Fibrinogen alpha chain | | Authors: | Yang, M.J, Xiang, H, Wang, J.W, Liu, B, Chen, Y.G, Liu, L, Deng, X.M, Feng, Y. | | Deposit date: | 2012-05-07 | | Release date: | 2012-08-08 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.917 Å) | | Cite: | Crystal Structures Reveal the Multi-Ligand Binding Mechanism of Staphylococcus aureus ClfB
Plos Pathog., 8, 2012
|
|
4F20
 
 | | Crystal structures reveal the multi-ligand binding mechanism of the Staphylococcus aureus ClfB | | Descriptor: | Clumping factor B, MAGNESIUM ION, peptide from Dermokine | | Authors: | Yang, M.J, Xiang, H, Wang, J.W, Liu, B, Chen, Y.G, Liu, L, Deng, X.M, Feng, Y. | | Deposit date: | 2012-05-07 | | Release date: | 2012-08-08 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.502 Å) | | Cite: | Crystal Structures Reveal the Multi-Ligand Binding Mechanism of Staphylococcus aureus ClfB
Plos Pathog., 8, 2012
|
|
8XAT
 
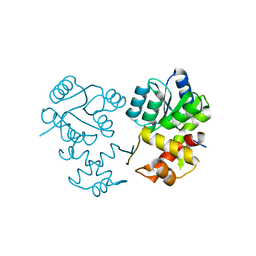 | |
8XAS
 
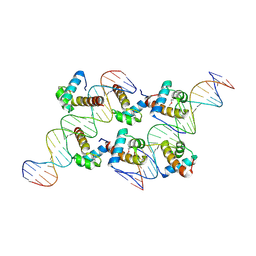 | | Crystal structure of AtARR1-DBD in complex with a DNA fragment | | Descriptor: | DNA (50-MER), Two-component response regulator ARR1 | | Authors: | Li, J.X, Zhou, C.M, zhang, P, Wang, J.W. | | Deposit date: | 2023-12-05 | | Release date: | 2024-01-24 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.346 Å) | | Cite: | The structure of B-ARR reveals the molecular basis of transcriptional activation by cytokinin.
Proc.Natl.Acad.Sci.USA, 121, 2024
|
|
3PZ7
 
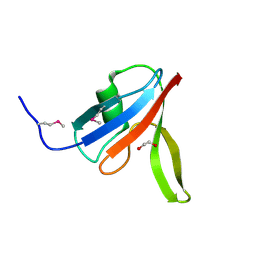 | |
21XL
 
 | | TLP-2g, a glycofibril obtained from a Karst cave from Guilin City, Guangxi Zhuang Autonomous Region, China | | Descriptor: | TLP-2g, beta-D-mannopyranose-(1-4)-beta-D-mannopyranose-(1-3)-beta-D-mannopyranose-(1-3)-[beta-D-mannopyranose-(1-4)]2-acetamido-2-deoxy-alpha-D-glucopyranose-(1-2)-[alpha-D-mannopyranose-(1-4)-beta-D-mannopyranose-(1-3)]beta-D-mannopyranose-(1-2)-[alpha-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-alpha-D-mannopyranose-(1-4)-alpha-D-mannopyranose-(1-3)]alpha-D-mannopyranose-(1-2)-beta-D-mannopyranose-(1-4)-[2-acetamido-2-deoxy-alpha-D-glucopyranose-(1-3)-[beta-D-mannopyranose-(1-4)]beta-D-mannopyranose-(1-2)-beta-D-mannopyranose-(1-4)-beta-D-mannopyranose-(1-2)]beta-D-mannopyranose-(1-3)-2-acetamido-2-deoxy-alpha-D-glucopyranose-(1-6)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-2)-alpha-D-mannopyranose-(1-2)-[beta-D-mannopyranose-(1-3)-2-acetamido-2-deoxy-alpha-D-glucopyranose-(1-3)]beta-D-mannopyranose | | Authors: | Hu, M.X, Zhang, Q, Chen, S, Ge, Q.J, Wang, J.W, Yan, N, Qin, L.J. | | Deposit date: | 2026-01-03 | | Release date: | 2026-02-18 | | Method: | ELECTRON MICROSCOPY (2.58 Å) | | Cite: | Absolute hand determination of glycofibrils from natural sources in cryo-EM
To Be Published
|
|
21XJ
 
 | | TLP-2a, a glycofibril obtained from a Karst cave from Guilin City, Guangxi Zhuang Autonomous Region, China | | Descriptor: | TLP-2a, beta-D-mannopyranose-(1-2)-beta-D-mannopyranose-(1-3)-[beta-D-mannopyranose-(1-4)]alpha-D-mannopyranose-(1-2)-[beta-D-mannopyranose-(1-4)-beta-D-mannopyranose-(1-2)-[beta-D-mannopyranose-(1-3)]beta-D-mannopyranose-(1-4)][beta-D-mannopyranose-(1-3)]beta-D-mannopyranose-(1-4)-beta-D-mannopyranose-(1-3)-[beta-D-mannopyranose-(1-3)-[2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)]alpha-D-mannopyranose-(1-3)-beta-D-mannopyranose-(1-2)-beta-D-mannopyranose-(1-4)-[beta-D-mannopyranose-(1-2)-[beta-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-3)]beta-D-mannopyranose-(1-4)]2-acetamido-2-deoxy-beta-D-glucopyranose | | Authors: | Hu, M.X, Zhang, Q, Chen, S, Ge, Q.J, Wang, J.W, Yan, N, Qin, L.J. | | Deposit date: | 2026-01-03 | | Release date: | 2026-02-18 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | Absolute hand determination of glycofibrils from natural sources in cryo-EM
To Be Published
|
|
21XM
 
 | | TLP-2h, a glycofibril obtained from a Karst cave from Guilin City, Guangxi Zhuang Autonomous Region, China | | Descriptor: | TLP-2h, beta-D-mannopyranose-(1-3)-[beta-D-mannopyranose-(1-4)]alpha-D-mannopyranose-(1-3)-beta-D-mannopyranose-(1-2)-beta-D-mannopyranose-(1-4)-[beta-D-mannopyranose-(1-2)-[beta-D-mannopyranose-(1-4)]beta-D-mannopyranose-(1-3)]beta-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-3)-beta-D-mannopyranose-(1-3)-beta-D-mannopyranose-(1-4)-beta-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-2)-beta-D-mannopyranose-(1-4)-[alpha-D-mannopyranose-(1-2)]beta-D-mannopyranose-(1-4)]2-acetamido-2-deoxy-beta-D-glucopyranose, beta-D-mannopyranose-(1-3)-beta-D-mannopyranose-(1-6)-beta-D-mannopyranose-(1-2)-[beta-D-mannopyranose-(1-3)]beta-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-4)]alpha-D-mannopyranose | | Authors: | Hu, M.X, Zhang, Q, Chen, S, Ge, Q.J, Wang, J.W, Yan, N, Qin, L.J. | | Deposit date: | 2026-01-03 | | Release date: | 2026-02-18 | | Method: | ELECTRON MICROSCOPY (2.8 Å) | | Cite: | Absolute hand determination of glycofibrils from natural sources in cryo-EM
To Be Published
|
|
4DJK
 
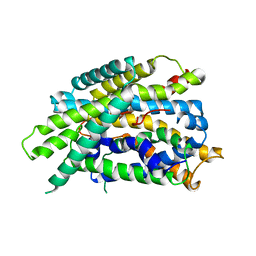 | | Structure of glutamate-GABA antiporter GadC | | Descriptor: | Probable glutamate/gamma-aminobutyrate antiporter | | Authors: | Ma, D, Lu, P.L, Yan, C.Y, Fan, C, Yin, P, Wang, J.W, Shi, Y.G. | | Deposit date: | 2012-02-02 | | Release date: | 2012-03-14 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (3.097 Å) | | Cite: | Structure and mechanism of a glutamate-GABA antiporter
Nature, 483, 2012
|
|
3PXI
 
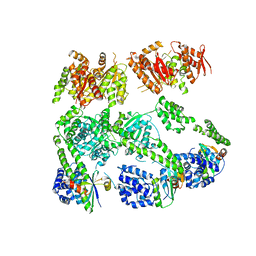 | | Structure of MecA108:ClpC | | Descriptor: | Adapter protein mecA 1, Negative regulator of genetic competence ClpC/MecB | | Authors: | Wang, F, Mei, Z.Q, Wang, J.W, Shi, Y.G. | | Deposit date: | 2010-12-09 | | Release date: | 2011-03-16 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (6.926 Å) | | Cite: | Structure and mechanism of the hexameric MecA-ClpC molecular machine.
Nature, 471, 2011
|
|
21XK
 
 | | TLP-2f, a glycofibril obtained from a Karst cave from Guilin City, Guangxi Zhuang Autonomous Region, China | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-2)-alpha-D-mannopyranose-(1-2)-beta-D-mannopyranose-(1-4)-[beta-D-mannopyranose-(1-2)-beta-D-mannopyranose-(1-3)][beta-D-mannopyranose-(1-2)][beta-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-2)-[beta-D-mannopyranose-(1-6)-beta-D-mannopyranose-(1-4)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, TLP-2f, alpha-D-mannopyranose-(1-6)-alpha-D-mannopyranose-(1-3)-beta-D-mannopyranose-(1-6)-[beta-D-mannopyranose-(1-4)]beta-D-mannopyranose-(1-2)-beta-D-mannopyranose-(1-6)-beta-D-mannopyranose-(1-4)-[beta-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-2)-beta-D-mannopyranose-(1-3)-beta-D-mannopyranose-(1-6)-beta-D-mannopyranose-(1-6)]2-acetamido-2-deoxy-alpha-D-glucopyranose-(1-3)-[2-acetamido-2-deoxy-beta-D-glucopyranose-(1-2)-alpha-D-mannopyranose-(1-2)]alpha-D-mannopyranose, ... | | Authors: | Hu, M.X, Zhang, Q, Chen, S, Ge, Q.J, Wang, J.W, Yan, N, Qin, L.J. | | Deposit date: | 2026-01-03 | | Release date: | 2026-02-18 | | Method: | ELECTRON MICROSCOPY (2.64 Å) | | Cite: | Absolute hand determination of glycofibrils from natural sources in cryo-EM
To Be Published
|
|
9WJU
 
 | |
9WJR
 
 | | Cryo-EM structure of the L. garvieae Man-PTS | | Descriptor: | Mannose/fructose/sorbose family PTS transporter subunit IIC, PTS mannose transporter subunit IID, alpha-D-mannopyranose | | Authors: | Wang, J.W. | | Deposit date: | 2025-08-31 | | Release date: | 2025-11-12 | | Last modified: | 2026-03-11 | | Method: | ELECTRON MICROSCOPY (2.87 Å) | | Cite: | Structural basis for the extended-spectrum antimicrobial activity of Garvieacin Q.
Appl.Environ.Microbiol., 92, 2026
|
|
9WJW
 
 | | Cryo-EM structure of the GarQ-lmYZ complex | | Descriptor: | Mannose/fructose/sorbose family PTS transporter subunit IIC, PTS mannose family transporter subunit IID, Prepeptide GarQ, ... | | Authors: | Wang, J.W. | | Deposit date: | 2025-08-31 | | Release date: | 2025-11-12 | | Last modified: | 2026-03-11 | | Method: | ELECTRON MICROSCOPY (2.96 Å) | | Cite: | Structural basis for the extended-spectrum antimicrobial activity of Garvieacin Q.
Appl.Environ.Microbiol., 92, 2026
|
|
6K1H
 
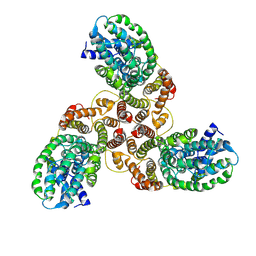 | | Structure of membrane protein | | Descriptor: | PTS mannose transporter subunit IID, PTS system mannose-specific EIIC component, alpha-D-mannopyranose | | Authors: | Wang, J.W, Zeng, J.W. | | Deposit date: | 2019-05-10 | | Release date: | 2019-07-10 | | Last modified: | 2025-07-02 | | Method: | ELECTRON MICROSCOPY (3.52 Å) | | Cite: | Structure of the mannose transporter of the bacterial phosphotransferase system.
Cell Res., 29, 2019
|
|
7DYR
 
 | |
