2V4D
 
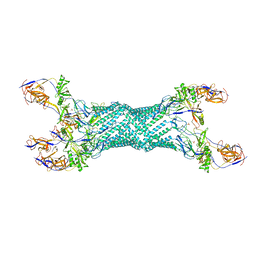 | | Re-refinement of MexA adaptor protein | | Descriptor: | MULTIDRUG RESISTANCE PROTEIN MEXA, SULFATE ION | | Authors: | Symmons, M.F, Bokma, E, Koronakis, E, Hughes, C, Koronakis, V. | | Deposit date: | 2008-09-18 | | Release date: | 2009-04-14 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | The Assembled Structure of a Complete Tripartite Bacterial Multidrug Efflux Pump.
Proc.Natl.Acad.Sci.USA, 106, 2009
|
|
1E3H
 
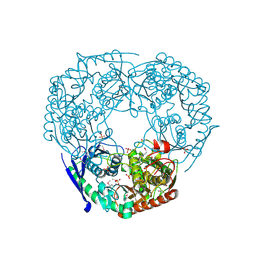 | | SeMet derivative of Streptomyces antibioticus PNPase/GPSI enzyme | | Descriptor: | GUANOSINE PENTAPHOSPHATE SYNTHETASE, SULFATE ION | | Authors: | Symmons, M.F, Jones, G.H, Luisi, B.F. | | Deposit date: | 2000-06-15 | | Release date: | 2000-11-05 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | A Duplicated Fold is the Structural Basis for Polynucleotide Phosphorylase Catalytic Activity, Processivity, and Regulation
Structure, 8, 2000
|
|
1E3P
 
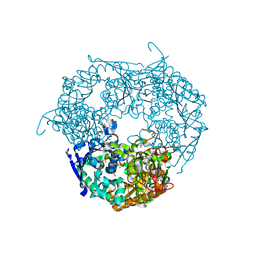 | | tungstate derivative of Streptomyces antibioticus PNPase/GPSI enzyme | | Descriptor: | Polyribonucleotide nucleotidyltransferase, SULFATE ION, TUNGSTATE(VI)ION | | Authors: | Symmons, M.F, Jones, G.H, Luisi, B.F. | | Deposit date: | 2000-06-20 | | Release date: | 2000-11-03 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | A Duplicated Fold is the Structural Basis for Polynucleotide Phosphorylase Catalytic Activity, Processivity, and Regulation
Structure, 8, 2000
|
|
9RLE
 
 | | LolCDE complex with Pal lipoprotein | | Descriptor: | (2S)-3-hydroxypropane-1,2-diyl dihexadecanoate, Lipoprotein-releasing system ATP-binding protein LolD, Lipoprotein-releasing system transmembrane protein LolC, ... | | Authors: | Symmons, M.F, Szewczyk, P, Greene, N.P, Hardwick, S.W, Koronakis, V. | | Deposit date: | 2025-06-16 | | Release date: | 2026-01-21 | | Last modified: | 2026-02-04 | | Method: | ELECTRON MICROSCOPY (3.18 Å) | | Cite: | Liganded LolCDE structures reveal a common substrate-LolE interaction guiding bacterial lipoprotein transport.
Proc.Natl.Acad.Sci.USA, 123, 2026
|
|
9RLH
 
 | | LolCDE complex with del 9-15 LolB lipoprotein | | Descriptor: | (2S)-3-hydroxypropane-1,2-diyl dihexadecanoate, Lipoprotein-releasing system ATP-binding protein LolD, Lipoprotein-releasing system transmembrane protein LolC, ... | | Authors: | Symmons, M.F, Szewczyk, P, Greene, N.P, Hardwick, S.W, Koronakis, V. | | Deposit date: | 2025-06-16 | | Release date: | 2026-01-21 | | Last modified: | 2026-02-04 | | Method: | ELECTRON MICROSCOPY (3.19 Å) | | Cite: | Liganded LolCDE structures reveal a common substrate-LolE interaction guiding bacterial lipoprotein transport.
Proc.Natl.Acad.Sci.USA, 123, 2026
|
|
9RLC
 
 | | LolCDE complex with Lpp lipoprotein | | Descriptor: | (2S)-3-hydroxypropane-1,2-diyl dihexadecanoate, Lipoprotein-releasing system ATP-binding protein LolD, Lipoprotein-releasing system transmembrane protein LolC, ... | | Authors: | Symmons, M.F, Szewczyk, P, Greene, N.P, Hardwick, S.W, Koronakis, V. | | Deposit date: | 2025-06-16 | | Release date: | 2026-01-21 | | Last modified: | 2026-02-04 | | Method: | ELECTRON MICROSCOPY (2.8 Å) | | Cite: | Liganded LolCDE structures reveal a common substrate-LolE interaction guiding bacterial lipoprotein transport.
Proc.Natl.Acad.Sci.USA, 123, 2026
|
|
9RLD
 
 | | LolCDE complex with Lpp lipoprotein | | Descriptor: | (2S)-3-hydroxypropane-1,2-diyl dihexadecanoate, Lipoprotein-releasing system ATP-binding protein LolD, Lipoprotein-releasing system transmembrane protein LolC, ... | | Authors: | Symmons, M.F, Szewczyk, P, Greene, N.P, Hardwick, S.W, Koronakis, V. | | Deposit date: | 2025-06-16 | | Release date: | 2026-01-21 | | Last modified: | 2026-02-04 | | Method: | ELECTRON MICROSCOPY (3 Å) | | Cite: | Liganded LolCDE structures reveal a common substrate-LolE interaction guiding bacterial lipoprotein transport.
Proc.Natl.Acad.Sci.USA, 123, 2026
|
|
9RLF
 
 | | LolCDE complex with Pal lipoprotein | | Descriptor: | (2S)-3-hydroxypropane-1,2-diyl dihexadecanoate, Lipoprotein-releasing system ATP-binding protein LolD, Lipoprotein-releasing system transmembrane protein LolC, ... | | Authors: | Symmons, M.F, Szewczyk, P, Greene, N.P, Hardwick, S.W, Koronakis, V. | | Deposit date: | 2025-06-16 | | Release date: | 2026-01-21 | | Last modified: | 2026-02-04 | | Method: | ELECTRON MICROSCOPY (3.17 Å) | | Cite: | Liganded LolCDE structures reveal a common substrate-LolE interaction guiding bacterial lipoprotein transport.
Proc.Natl.Acad.Sci.USA, 123, 2026
|
|
9RLK
 
 | | LolCDE(delta 235-252) complex with Pal lipoprotein | | Descriptor: | (2S)-3-hydroxypropane-1,2-diyl dihexadecanoate, Lipoprotein-releasing system ATP-binding protein LolD, Lipoprotein-releasing system transmembrane protein LolC, ... | | Authors: | Symmons, M.F, Szewczyk, P, Greene, N.P, Hardwick, S.W, Koronakis, V. | | Deposit date: | 2025-06-16 | | Release date: | 2026-01-21 | | Last modified: | 2026-02-04 | | Method: | ELECTRON MICROSCOPY (3.29 Å) | | Cite: | Liganded LolCDE structures reveal a common substrate-LolE interaction guiding bacterial lipoprotein transport.
Proc.Natl.Acad.Sci.USA, 123, 2026
|
|
9RLI
 
 | | LolCDEdelta(235-252) complex with Pal lipoprotein | | Descriptor: | (2S)-3-hydroxypropane-1,2-diyl dihexadecanoate, Lipoprotein-releasing system ATP-binding protein LolD, Lipoprotein-releasing system transmembrane protein LolC, ... | | Authors: | Symmons, M.F, Szewczyk, P, Greene, N.P, Hardwick, S.W, Koronakis, V. | | Deposit date: | 2025-06-16 | | Release date: | 2026-01-21 | | Last modified: | 2026-02-04 | | Method: | ELECTRON MICROSCOPY (3.63 Å) | | Cite: | Liganded LolCDE structures reveal a common substrate-LolE interaction guiding bacterial lipoprotein transport.
Proc.Natl.Acad.Sci.USA, 123, 2026
|
|
9RLG
 
 | | LolCDE complex with LolB lipoprotein | | Descriptor: | (2S)-3-hydroxypropane-1,2-diyl dihexadecanoate, Lipoprotein-releasing system ATP-binding protein LolD, Lipoprotein-releasing system transmembrane protein LolC, ... | | Authors: | Symmons, M.F, Szewczyk, P, Greene, N.P, Hardwick, S.W, Koronakis, V. | | Deposit date: | 2025-06-16 | | Release date: | 2026-01-21 | | Last modified: | 2026-02-04 | | Method: | ELECTRON MICROSCOPY (3.09 Å) | | Cite: | Liganded LolCDE structures reveal a common substrate-LolE interaction guiding bacterial lipoprotein transport.
Proc.Natl.Acad.Sci.USA, 123, 2026
|
|
9RLJ
 
 | | LolCDE with bound ATPgammaS | | Descriptor: | Lipoprotein-releasing system ATP-binding protein LolD, Lipoprotein-releasing system transmembrane protein LolC, Lipoprotein-releasing system transmembrane protein LolE, ... | | Authors: | Symmons, M.F, Szewczyk, P, Greene, N.P, Hardwick, S.W, Koronakis, V. | | Deposit date: | 2025-06-16 | | Release date: | 2026-01-21 | | Last modified: | 2026-02-04 | | Method: | ELECTRON MICROSCOPY (2.9 Å) | | Cite: | Liganded LolCDE structures reveal a common substrate-LolE interaction guiding bacterial lipoprotein transport.
Proc.Natl.Acad.Sci.USA, 123, 2026
|
|
2C0X
 | | MOLECULAR STRUCTURE OF FD FILAMENTOUS BACTERIOPHAGE REFINED WITH RESPECT TO X-RAY FIBRE DIFFRACTION AND SOLID-STATE NMR DATA | | Descriptor: | COAT PROTEIN B | | Authors: | Marvin, D.A, Welsh, L.C, Symmons, M.F, Scott, W.R.P, Straus, S.K. | | Deposit date: | 2005-09-08 | | Release date: | 2005-12-14 | | Last modified: | 2025-01-22 | | Method: | SOLID-STATE NMR | | Cite: | Molecular Structure of Fd (F1, M13) Filamentous Bacteriophage Refined with Respect to X-Ray Fibre Diffraction and Solid-State NMR Data Supports Specific Models of Phage Assembly at the Bacterial Membrane.
J.Mol.Biol., 355, 2006
|
|
2C0W
 | | Molecular Structure of fd Filamentous Bacteriophage Refined with Respect to X-ray Fibre Diffraction | | Descriptor: | COAT PROTEIN B | | Authors: | Marvin, D.A, Welsh, L.C, Symmons, M.F, Scott, W.R.P, Straus, S.K. | | Deposit date: | 2005-09-08 | | Release date: | 2005-12-14 | | Last modified: | 2024-02-14 | | Method: | FIBER DIFFRACTION (3.2 Å) | | Cite: | Molecular Structure of Fd (F1, M13) Filamentous Bacteriophage Refined with Respect to X-Ray Fibre Diffraction and Solid-State NMR Data Supports Specific Models of Phage Assembly at the Bacterial Membrane.
J.Mol.Biol., 355, 2006
|
|
1IFP
 | |
1QL2
 
 | |
1QL1
 
 | |
2WMZ
 | | Structure of a mutated TolC | | Descriptor: | CHLORIDE ION, OUTER MEMBRANE PROTEIN TOLC, SODIUM ION, ... | | Authors: | Hinchliffe, P, Pei, X.Y, Symmons, M.F, Koronakis, E, Hughes, C, Koronakis, V. | | Deposit date: | 2009-07-03 | | Release date: | 2011-01-26 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Structures of Sequential Open States in a Symmetrical Opening Transition of the Tolc Exit Duct.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
1HH0
 | | Filamentous Bacteriophage PH75 | | Descriptor: | PH75 INOVIRUS MAJOR COAT PROTEIN | | Authors: | Pederson, D.M, Welsh, L.C, Marvin, D.A, Sampson, M, Perham, R.N, Yu, M, Slater, M.R. | | Deposit date: | 2000-12-17 | | Release date: | 2001-06-01 | | Last modified: | 2024-02-14 | | Method: | FIBER DIFFRACTION (2.4 Å) | | Cite: | The Protein Capsid of Filamentous Bacteriophage Ph75 from Thermus Thermophilus
J.Mol.Biol., 309, 2001
|
|
1HGZ
 | | Filamentous Bacteriophage PH75 | | Descriptor: | PH75 INOVIRUS MAJOR COAT PROTEIN | | Authors: | Pederson, D.M, Welsh, L.C, Marvin, D.A, Sampson, M, Perham, R.N, Yu, M, Slater, M.R. | | Deposit date: | 2000-12-17 | | Release date: | 2001-06-01 | | Last modified: | 2024-02-14 | | Method: | FIBER DIFFRACTION (2.4 Å) | | Cite: | The Protein Capsid of Filamentous Bacteriophage Ph75 from Thermus Thermophilus
J.Mol.Biol., 309, 2001
|
|
1HGV
 | | Filamentous Bacteriophage PH75 | | Descriptor: | PH75 INOVIRUS MAJOR COAT PROTEIN | | Authors: | Pederson, D.M, Welsh, L.C, Marvin, D.A, Sampson, M, Perham, R.N, Yu, M, Slater, M.R. | | Deposit date: | 2000-12-15 | | Release date: | 2001-06-01 | | Last modified: | 2023-12-13 | | Method: | FIBER DIFFRACTION (2.4 Å) | | Cite: | The Protein Capsid of Filamentous Bacteriophage Ph75 from Thermus Thermophilus
J.Mol.Biol., 309, 2001
|
|
2XMN
 | | High resolution snapshots of defined TolC open states present an iris- like movement of periplasmic entrance helices | | Descriptor: | CHLORIDE ION, DODECYL-BETA-D-MALTOSIDE, Outer membrane protein TolC | | Authors: | Pei, X.Y, Koronakis, E, Hughes, C, Koronakis, V. | | Deposit date: | 2010-07-28 | | Release date: | 2011-01-26 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | Structures of sequential open states in a symmetrical opening transition of the TolC exit duct.
Proc. Natl. Acad. Sci. U.S.A., 108, 2011
|
|
