3KPF
 
 | |
3KU9
 
 | |
3L1R
 
 | |
7AHW
 
 | |
7ALY
 
 | |
7AMX
 
 | |
7BGO
 
 | |
3MFA
 
 | | Computationally designed endo-1,4-beta-xylanase | | Descriptor: | Endo-1,4-beta-xylanase, SULFATE ION | | Authors: | Morin, A, Harp, J.M. | | Deposit date: | 2010-04-01 | | Release date: | 2010-11-10 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.63 Å) | | Cite: | Computational design of an endo-1,4-{beta}-xylanase ligand binding site.
Protein Eng.Des.Sel., 24, 2011
|
|
3MF9
 | | Computationally designed endo-1,4-beta-xylanase | | Descriptor: | Endo-1,4-beta-xylanase, SULFATE ION | | Authors: | Morin, A, Harp, J.M. | | Deposit date: | 2010-04-01 | | Release date: | 2010-11-10 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Computational design of an endo-1,4-{beta}-xylanase ligand binding site.
Protein Eng.Des.Sel., 24, 2011
|
|
3MFC
 
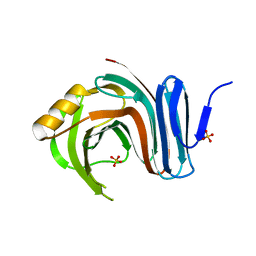 | | Computationally designed end0-1,4-beta,xylanase | | Descriptor: | Endo-1,4-beta-xylanase, SULFATE ION | | Authors: | Morin, A, Harp, J.M. | | Deposit date: | 2010-04-01 | | Release date: | 2010-11-10 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Computational design of an endo-1,4-{beta}-xylanase ligand binding site.
Protein Eng.Des.Sel., 24, 2011
|
|
3MF6
 
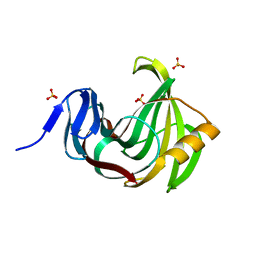 | | Computationally designed endo-1,4-beta-xylanase | | Descriptor: | Endo-1,4-beta-xylanase, SULFATE ION | | Authors: | Morin, A, Harp, J.M. | | Deposit date: | 2010-04-01 | | Release date: | 2010-11-10 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.28 Å) | | Cite: | Computational design of an endo-1,4-{beta}-xylanase ligand binding site.
Protein Eng.Des.Sel., 24, 2011
|
|
4F7R
 
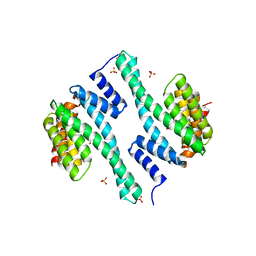 | |
3S9F
 
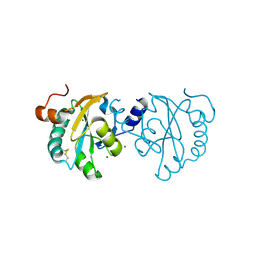 | |
3TUE
 
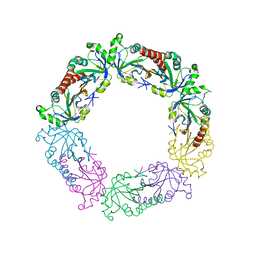 | |
5AIQ
 
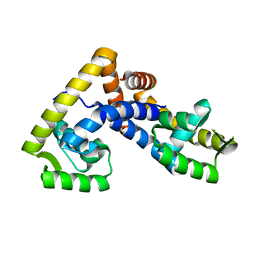 | | Crystal structure of ligand-free NadR | | Descriptor: | TRANSCRIPTIONAL REGULATOR, MARR FAMILY | | Authors: | Liguori, A, Malito, E, Bottomley, M.J. | | Deposit date: | 2015-02-16 | | Release date: | 2016-03-02 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.716 Å) | | Cite: | Molecular Basis of Ligand-Dependent Regulation of Nadr, the Transcriptional Repressor of Meningococcal Virulence Factor Nada.
Plos Pathog., 12, 2016
|
|
5AIP
 
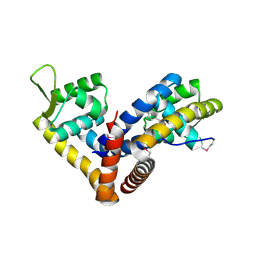 | | Crystal structure of NadR in complex with 4-hydroxyphenylacetate | | Descriptor: | 4-HYDROXYPHENYLACETATE, TRANSCRIPTIONAL REGULATOR, MARR FAMILY | | Authors: | Liguori, A, Malito, E, Bottomley, M.J. | | Deposit date: | 2015-02-16 | | Release date: | 2016-03-02 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Molecular Basis of Ligand-Dependent Regulation of Nadr, the Transcriptional Repressor of Meningococcal Virulence Factor Nada.
Plos Pathog., 12, 2016
|
|
4Z7Y
 
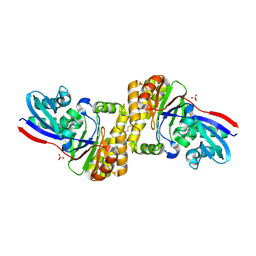 | | diphosphomevalonate decarboxylase from the Sulfolobus solfataricus, space group P21 | | Descriptor: | Diphosphomevalonate decarboxylase, SULFATE ION | | Authors: | Hattori, A, Unno, H, Hemmi, H. | | Deposit date: | 2015-04-08 | | Release date: | 2015-09-16 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | In Vivo Formation of the Protein Disulfide Bond That Enhances the Thermostability of Diphosphomevalonate Decarboxylase, an Intracellular Enzyme from the Hyperthermophilic Archaeon Sulfolobus solfataricus
J.Bacteriol., 197, 2015
|
|
5S9S
 
 | |
5SA2
 
 | |
5S9V
 
 | |
5S9X
 
 | | PanDDA analysis group deposition -- Crystal Structure of Trypanosoma brucei Trypanothione reductase in complex with Z1899842917 | | Descriptor: | 3-[(3,5-dimethyl-1,2-oxazol-4-yl)methyl]-5-methyl-1,3,4-thiadiazol-2(3H)-one, BROMIDE ION, DIMETHYL SULFOXIDE, ... | | Authors: | Fiorillo, A, Ilari, A. | | Deposit date: | 2021-05-18 | | Release date: | 2022-06-15 | | Last modified: | 2023-03-01 | | Method: | X-RAY DIFFRACTION (1.84 Å) | | Cite: | Innovative Approach for a Classic Target: Fragment Screening on Trypanothione Reductase Reveals New Opportunities for Drug Design.
Front Mol Biosci, 9, 2022
|
|
5SA3
 
 | |
5SA0
 
 | | PanDDA analysis group deposition -- Crystal Structure of Trypanosoma brucei Trypanothione reductase in complex with Z1506050651 | | Descriptor: | (1S)-N,2,2-trimethyl-N-(pyridin-3-yl)cyclopropane-1-carboxamide, BROMIDE ION, DIMETHYL SULFOXIDE, ... | | Authors: | Fiorillo, A, Ilari, A. | | Deposit date: | 2021-05-18 | | Release date: | 2022-06-15 | | Last modified: | 2023-03-01 | | Method: | X-RAY DIFFRACTION (1.97 Å) | | Cite: | Innovative Approach for a Classic Target: Fragment Screening on Trypanothione Reductase Reveals New Opportunities for Drug Design.
Front Mol Biosci, 9, 2022
|
|
5S9W
 
 | |
5S9U
 
 | |
