1S58
 
 | |
1B1A
 
 | | GLUTAMATE MUTASE (B12-BINDING SUBUNIT), NMR, MINIMIZED AVERAGE STRUCTURE | | Descriptor: | GLUTAMATE MUTASE | | Authors: | Hoffmann, B, Konrat, R, Bothe, H, Buckel, W, Kraeutler, B. | | Deposit date: | 1998-11-19 | | Release date: | 1999-07-13 | | Last modified: | 2024-04-10 | | Method: | SOLUTION NMR | | Cite: | Structure and dynamics of the B12-binding subunit of glutamate mutase from Clostridium cochlearium.
Eur.J.Biochem., 263, 1999
|
|
1A88
 
 | | CHLOROPEROXIDASE L | | Descriptor: | CHLOROPEROXIDASE L | | Authors: | Hofmann, B, Toelzer, S, Pelletier, I, Altenbuchner, J, Van Pee, K.-H, Hecht, H.-J. | | Deposit date: | 1998-04-03 | | Release date: | 1998-10-14 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural investigation of the cofactor-free chloroperoxidases.
J.Mol.Biol., 279, 1998
|
|
1A8S
 
 | | CHLOROPEROXIDASE F/PROPIONATE COMPLEX | | Descriptor: | CHLOROPEROXIDASE F, PROPANOIC ACID, SULFATE ION | | Authors: | Hofmann, B, Toelzer, S, Pelletier, I, Altenbuchner, J, Van Pee, K.-H, Hecht, H.-J. | | Deposit date: | 1998-03-27 | | Release date: | 1998-06-17 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural investigation of the cofactor-free chloroperoxidases.
J.Mol.Biol., 279, 1998
|
|
1A8U
 
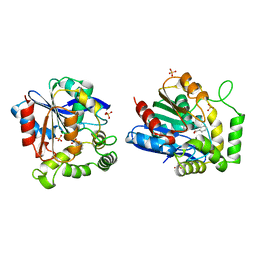 | | CHLOROPEROXIDASE T/BENZOATE COMPLEX | | Descriptor: | BENZOIC ACID, CHLOROPEROXIDASE T, SULFATE ION | | Authors: | Hofmann, B, Toelzer, S, Pelletier, I, Altenbuchner, J, Van Pee, K.-H, Hecht, H.-J. | | Deposit date: | 1998-03-26 | | Release date: | 1998-06-17 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structural investigation of the cofactor-free chloroperoxidases.
J.Mol.Biol., 279, 1998
|
|
1A8Q
 
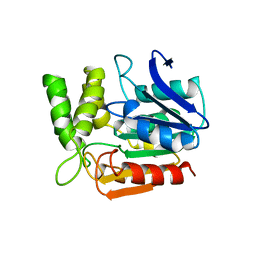 | | BROMOPEROXIDASE A1 | | Descriptor: | BROMOPEROXIDASE A1 | | Authors: | Hofmann, B, Toelzer, S, Pelletier, I, Altenbuchner, J, Van Pee, K.-H, Hecht, H.-J. | | Deposit date: | 1998-03-27 | | Release date: | 1998-06-17 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Structural investigation of the cofactor-free chloroperoxidases.
J.Mol.Biol., 279, 1998
|
|
1BRT
 
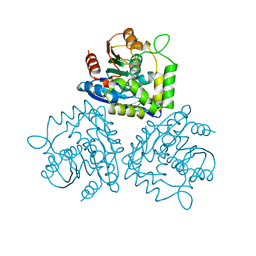 | | BROMOPEROXIDASE A2 MUTANT M99T | | Descriptor: | BROMOPEROXIDASE A2, CHLORIDE ION | | Authors: | Hofmann, B, Toelzer, S, Pelletier, I, Altenbuchner, J, Van Pee, K.H, Hecht, H.J. | | Deposit date: | 1998-03-30 | | Release date: | 1998-06-17 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Structural investigation of the cofactor-free chloroperoxidases.
J.Mol.Biol., 279, 1998
|
|
1A7U
 
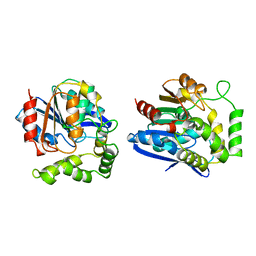 | | CHLOROPEROXIDASE T | | Descriptor: | CHLOROPEROXIDASE T | | Authors: | Hofmann, B, Toelzer, S, Pelletier, I, Altenbuchner, J, Van Pee, K.-H, Hecht, H.-J. | | Deposit date: | 1998-03-17 | | Release date: | 1998-06-17 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Structural investigation of the cofactor-free chloroperoxidases.
J.Mol.Biol., 279, 1998
|
|
3P0S
 
 | |
3N9G
 
 | |
3N7X
 
 | |
1FMF
 
 | | REFINED SOLUTION STRUCTURE OF THE (13C,15N-LABELED) B12-BINDING SUBUNIT OF GLUTAMATE MUTASE FROM CLOSTRIDIUM TETANOMORPHUM | | Descriptor: | METHYLASPARTATE MUTASE S CHAIN | | Authors: | Hoffmann, B, Konrat, R, Tollinger, M, Huhta, M, Marsh, E.N.G, Kraeutler, B. | | Deposit date: | 2000-08-17 | | Release date: | 2002-02-15 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | A protein pre-organized to trap the nucleotide moiety of coenzyme B(12): refined solution structure of the B(12)-binding subunit of glutamate mutase from Clostridium tetanomorphum.
Chembiochem, 2, 2001
|
|
4RKM
 
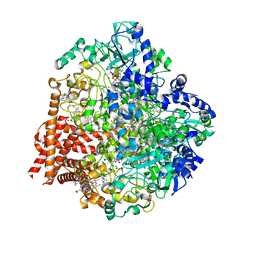 | | Wolinella succinogenes octaheme sulfite reductase MccA, form I | | Descriptor: | (R,R)-2,3-BUTANEDIOL, ACETATE ION, COPPER (I) ION, ... | | Authors: | Hermann, B, Kern, M, La Pietra, L, Simon, J, Einsle, O. | | Deposit date: | 2014-10-13 | | Release date: | 2015-02-04 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The octahaem MccA is a haem c-copper sulfite reductase.
Nature, 520, 2015
|
|
4RKN
 
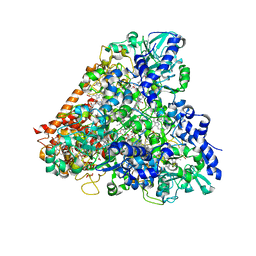 | | Wolinella succinogenes octaheme sulfite reductase MccA, form II | | Descriptor: | COPPER (II) ION, DITHIONITE, MccA, ... | | Authors: | Hermann, B, Kern, M, La Pietra, L, Simon, J, Einsle, O. | | Deposit date: | 2014-10-13 | | Release date: | 2015-02-04 | | Last modified: | 2017-11-22 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The octahaem MccA is a haem c-copper sulfite reductase.
Nature, 520, 2015
|
|
1EWX
 
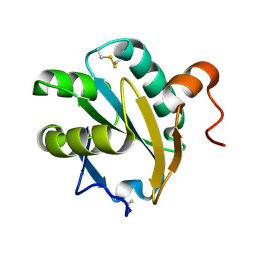 | | Crystal structure of native tryparedoxin I from Crithidia fasciculata | | Descriptor: | TRYPAREDOXIN I | | Authors: | Hofmann, B, Guerrero, S.A, Kalisz, H.M, Menge, U, Nogoceke, E, Montemartini, M, Singh, M, Flohe, L, Hecht, H.J. | | Deposit date: | 2000-04-28 | | Release date: | 2000-05-10 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structures of tryparedoxins revealing interaction with trypanothione.
Biol.Chem., 382, 2001
|
|
1I5G
 
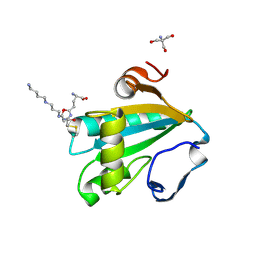 | | TRYPAREDOXIN II COMPLEXED WITH GLUTATHIONYLSPERMIDINE | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, GLUTATHIONYLSPERMIDINE, TRYPAREDOXIN II | | Authors: | Hofmann, B, Budde, H, Bruns, K, Guerrero, S.A, Kalisz, H.M, Menge, U, Montemartini, M, Nogoceke, E, Steinert, P, Wissing, J.B, Flohe, L, Hecht, H.-J. | | Deposit date: | 2001-02-27 | | Release date: | 2001-03-14 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Structures of tryparedoxins revealing interaction with trypanothione.
Biol.Chem., 382, 2001
|
|
1LIP
 
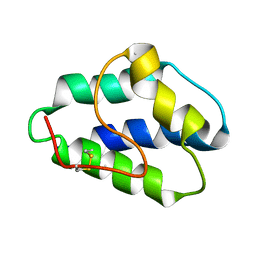 | |
1OW9
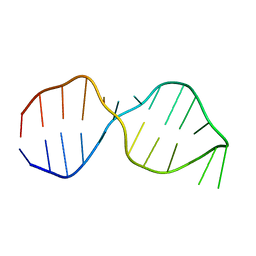 
 | | NMR Structure of the Active Conformation of the VS Ribozyme Cleavage Site | | Descriptor: | A mimic of the VS Ribozyme Hairpin Substrate | | Authors: | Hoffmann, B, Mitchell, G.T, Gendron, P, Major, F, Andersen, A.A, Collins, R.A, Legault, P. | | Deposit date: | 2003-03-28 | | Release date: | 2003-05-20 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | NMR Structure of the Active Conformation of the Varkud satellite Ribozyme Cleavage Site
Proc.Natl.Acad.Sci.USA, 100, 2003
|
|
1UB1
 
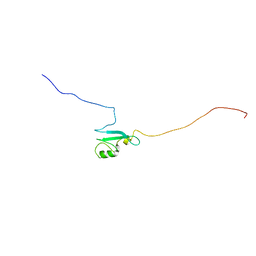 | | Solution structure of the matrix attachment region-binding domain of chicken MeCP2 | | Descriptor: | attachment region binding protein | | Authors: | Heitmann, B, Maurer, T, Weitzel, J.M, Stratling, W.H, Kalbitzer, H.R, Brunner, E, Structural Proteomics in Europe (SPINE) | | Deposit date: | 2003-03-27 | | Release date: | 2003-08-05 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the matrix attachment region-binding domain of chicken MeCP2
EUR.J.BIOCHEM., 270, 2003
|
|
1FG4
 
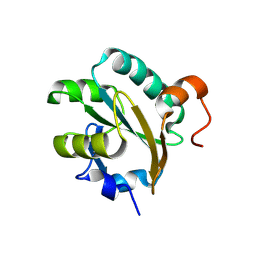 | | STRUCTURE OF TRYPAREDOXIN II | | Descriptor: | TRYPAREDOXIN II | | Authors: | Hofmann, B, Budde, H, Bruns, K, Guerrero, S.A, Kalisz, H.M, Menge, U, Montemartini, M, Nogoceke, E, Steinert, P, Wissing, J.B, Flohe, L, Hecht, H.J. | | Deposit date: | 2000-07-28 | | Release date: | 2001-04-25 | | Last modified: | 2017-10-04 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structures of tryparedoxins revealing interaction with trypanothione.
Biol.Chem., 382, 2001
|
|
1CV8
 
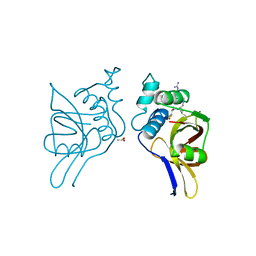 | | STAPHOPAIN, CYSTEINE PROTEINASE FROM STAPHYLOCOCCUS AUREUS V8 | | Descriptor: | ACETATE ION, N-[N-[1-HYDROXYCARBOXYETHYL-CARBONYL]LEUCYLAMINO-BUTYL]-GUANIDINE, STAPHOPAIN | | Authors: | Hofmann, B, Schomburg, D, Hecht, H.-J. | | Deposit date: | 1998-05-08 | | Release date: | 1998-10-14 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Crystal Structure of a Thiol Proteinase from Staphylococcus Aureus V-8 in the E-64 Inhibitor Complex
Acta Crystallogr.,Sect.A, 49, 1993
|
|
1EZK
 
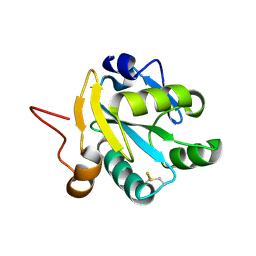 | | Crystal structure of recombinant tryparedoxin I | | Descriptor: | TRYPAREDOXIN I | | Authors: | Hofmann, B, Guerrero, S.A, Kalisz, H.M, Menge, U, Nogoceke, E, Montemartini, M, Singh, M, Flohe, L, Hecht, H.J. | | Deposit date: | 2000-05-11 | | Release date: | 2000-05-24 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structures of tryparedoxins revealing interaction with trypanothione.
Biol.Chem., 382, 2001
|
|
4BG7
 
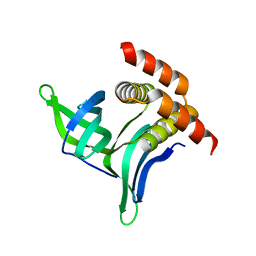 | |
6I5B
 
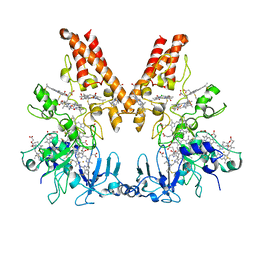 | | Crystal Structure of Outer Cell Wall Cytochrome OcwA | | Descriptor: | (R,R)-2,3-BUTANEDIOL, CHLORIDE ION, HEME C, ... | | Authors: | Hermann, B, Einsle, O. | | Deposit date: | 2018-11-13 | | Release date: | 2019-09-25 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | How Thermophilic Gram-Positive Organisms Perform Extracellular Electron Transfer: Characterization of the Cell Surface Terminal Reductase OcwA.
Mbio, 10, 2019
|
|
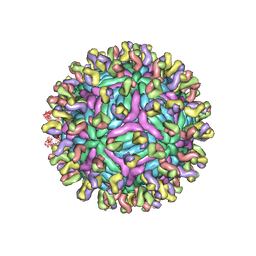
3IYW
 
 | |
