9YRJ
 
 | |
9YRI
 
 | |
9YTU
 
 | |
9C7O
 
 | |
9C7N
 
 | |
9NG5
 
 | |
9NGH
 
 | |
3I17
 
 | |
8SDB
 
 | | Crystal Structure of E.Coli Branching Enzyme in complex with malto-octose | | Descriptor: | 1,4-alpha-glucan branching enzyme GlgB, alpha-D-glucopyranose, alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose, ... | | Authors: | Bingham, C.R, Nayebi, H, Fawaz, R, Geiger, J.H. | | Deposit date: | 2023-04-06 | | Release date: | 2023-07-12 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | The Structure of Maltooctaose-Bound Escherichia coli Branching Enzyme Suggests a Mechanism for Donor Chain Specificity.
Molecules, 28, 2023
|
|
8W02
 | |
8VZZ
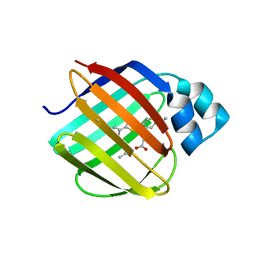 
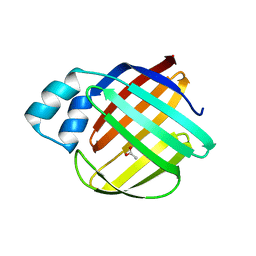
 | | Q108K:K40L:T51V:T53S:Y19W:R58W mutant of hCRBPII bound to synthetic fluorophore TD-1V | | Descriptor: | (2E)-3-{5-[4-(dimethylamino)phenyl]thiophen-2-yl}but-2-enal, ACETATE ION, Retinol-binding protein 2 | | Authors: | Nosssoni, Z, Bingham, C.R, Geiger, J.H, Borhan, B. | | Deposit date: | 2024-02-13 | | Release date: | 2024-04-03 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.22 Å) | | Cite: | Regulation of Absorption and Emission in a Protein/Fluorophore Complex.
Acs Chem.Biol., 19, 2024
|
|
8W00
 | |
8VZY
 
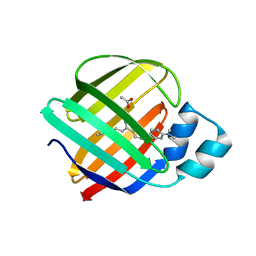
 | | Q108K:K40L:T51V:T53S:R58Y mutant of hCRBPII bound to synthetic fluorophore TD-1V | | Descriptor: | (2E)-3-{5-[4-(dimethylamino)phenyl]thiophen-2-yl}but-2-enal, ACETATE ION, Retinol-binding protein 2 | | Authors: | Nossoni, Z, Bingham, C.R, Geiger, J.H, Borhan, B. | | Deposit date: | 2024-02-13 | | Release date: | 2024-04-03 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.34 Å) | | Cite: | Regulation of Absorption and Emission in a Protein/Fluorophore Complex.
Acs Chem.Biol., 19, 2024
|
|
8VZX
 
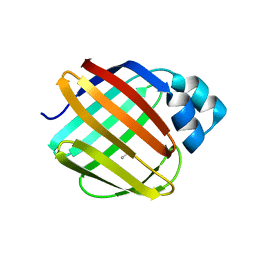 | |
4LPC
 
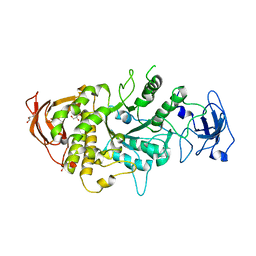 | |
4M6S
 
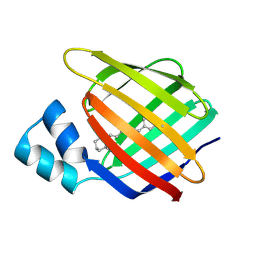 | |
4M7M
 | |
4LQ1
 
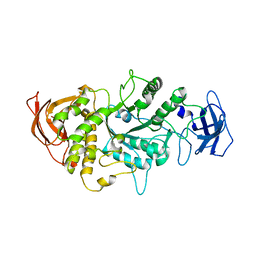 | |
9O2N
 
 | |
4EXZ
 | |
4EFG
 | |
4GKC
 | |
4I9S
 | |
4I9R
 | |
9O3M
 
 | |
