1VTP
 
 | |
1TIH
 
 | |
2JZM
 
 | |
1QH2
 
 | |
1WN4
 
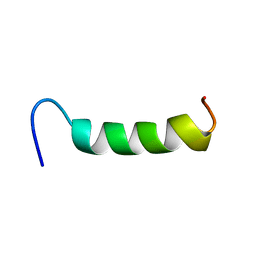 | | NMR Structure of VoNTR | | Descriptor: | VoNTR protein | | Authors: | Dutton, J.L, Renda, R.F, Waine, C, Clark, R.J, Daly, N.L, Jennings, C.V, Anderson, M.A, Craik, D.J. | | Deposit date: | 2004-07-27 | | Release date: | 2004-09-14 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Conserved structural and sequence elements implicated in the processing of gene-encoded circular proteins
J.Biol.Chem., 279, 2004
|
|
1WN8
 
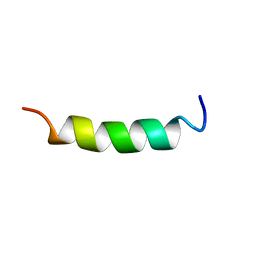 | | NMR Structure of OaNTR | | Descriptor: | Kalata B3/B6 | | Authors: | Dutton, J.L, Renda, R.F, Waine, C, Clark, R.J, Daly, N.L, Jennings, C.V, Anderson, M.A, Craik, D.J. | | Deposit date: | 2004-07-28 | | Release date: | 2004-09-14 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Conserved structural and sequence elements implicated in the processing of gene-encoded circular proteins
J.Biol.Chem., 279, 2004
|
|
1YTP
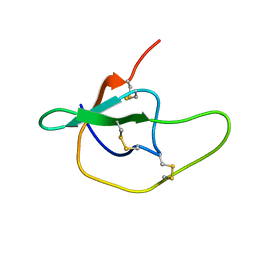 
 | |
2JYY
 
 | |
1N4N
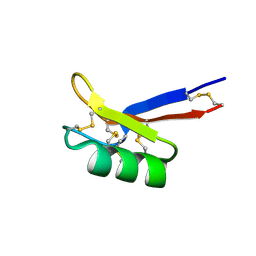 
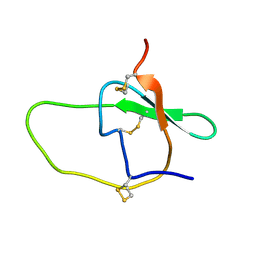
 | | Structure of the Plant Defensin PhD1 from Petunia Hybrida | | Descriptor: | floral defensin-like protein 1 | | Authors: | Janssen, B.J.C, Schirra, H.J, Lay, F.T, Anderson, M.A, Craik, D.J. | | Deposit date: | 2002-11-01 | | Release date: | 2003-11-01 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Structure of Petunia hybrida defensin 1, a novel plant defensin with five disulfide bonds
Biochemistry, 42, 2003
|
|
1MR4
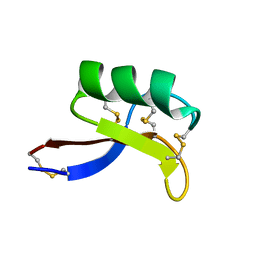 
 | | Solution Structure of NaD1 from Nicotiana alata | | Descriptor: | Nicotiana alata plant defensin 1 (NaD1) | | Authors: | Lay, F.T, Schirra, H.J, Scanlon, M.J, Anderson, M.A, Craik, D.J. | | Deposit date: | 2002-09-18 | | Release date: | 2003-09-18 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | The Three-dimensional Solution Structure of NaD1, a New Floral Defensin from Nicotiana alata and its Application to a Homology Model of the Crop Defense Protein alfAFP
J.MOL.BIOL., 325, 2003
|
|
1PT4
 
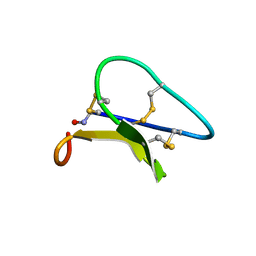 | | Solution structure of the Moebius cyclotide kalata B2 | | Descriptor: | kalata B2 | | Authors: | Jennings, C.V, Anderson, M.A, Daly, N.L, Rosengren, K.J, Craik, D.J. | | Deposit date: | 2003-06-23 | | Release date: | 2004-08-10 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | Isolation, Solution Structure, and Insecticidal Activity of Kalata B2, a Circular Protein with a Twist: Do Mobius Strips Exist in Nature?(,)
Biochemistry, 44, 2005
|
|
1CE3
 | |
1FYB
 
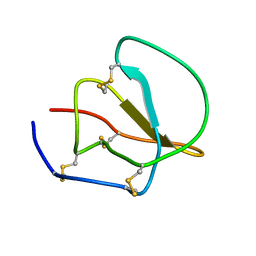
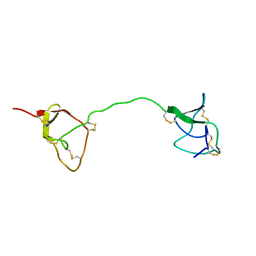 | | SOLUTION STRUCTURE OF C1-T1, A TWO-DOMAIN PROTEINASE INHIBITOR DERIVED FROM THE CIRCULAR PRECURSOR PROTEIN NA-PROPI FROM NICOTIANA ALATA | | Descriptor: | PROTEINASE INHIBITOR | | Authors: | Craik, D.J, Schirra, H.J, Scanlon, M.J, Anderson, M.A. | | Deposit date: | 2000-09-28 | | Release date: | 2001-02-21 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | The solution structure of C1-T1, a two-domain proteinase inhibitor derived from a circular precursor protein from Nicotiana alata.
J.Mol.Biol., 306, 2001
|
|
1JBL
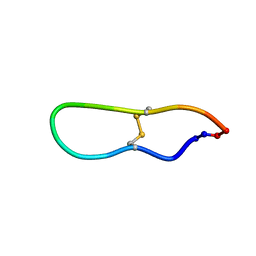 
 | | Solution structure of SFTI-1, A cyclic trypsin inhibitor from sunflower seeds | | Descriptor: | CYCLIC TRYPSIN INHIBITOR | | Authors: | Korsinczky, M.L.J, Schirra, H.J, Rosengren, K.J, West, J, Condie, B.A, Otvos, L, Anderson, M.A, Craik, D.J. | | Deposit date: | 2001-06-05 | | Release date: | 2001-08-22 | | Last modified: | 2015-04-15 | | Method: | SOLUTION NMR | | Cite: | Solution structures by 1H NMR of the novel cyclic trypsin inhibitor SFTI-1 from sunflower seeds and an acyclic permutant.
J.Mol.Biol., 311, 2001
|
|
1JBN
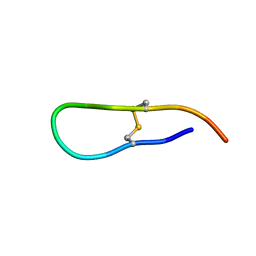 
 | | Solution structure of an acyclic permutant of SFTI-1, A trypsin inhibitor from sunflower seeds | | Descriptor: | CYCLIC TRYPSIN INHIBITOR | | Authors: | Korsinczky, M.L.J, Schirra, H.J, Rosengren, K.J, West, J, Condie, B.A, Otvos, L, Anderson, M.A, Craik, D.J. | | Deposit date: | 2001-06-06 | | Release date: | 2001-08-22 | | Last modified: | 2016-12-28 | | Method: | SOLUTION NMR | | Cite: | Solution structures by 1H NMR of the novel cyclic trypsin inhibitor SFTI-1 from sunflower seeds and an acyclic permutant.
J.Mol.Biol., 311, 2001
|
|
4AAZ
 
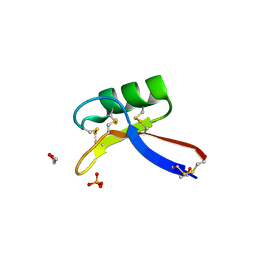 | | X-ray structure of Nicotiana alata Defensin 1 NaD1 | | Descriptor: | 1,2-ETHANEDIOL, FLOWER-SPECIFIC DEFENSIN, PHOSPHATE ION | | Authors: | Lay, F.T, Mills, G.D, Hulett, M.D, Kvansakul, M. | | Deposit date: | 2011-12-06 | | Release date: | 2012-04-25 | | Last modified: | 2012-08-08 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Dimerization of Plant Defensin Nad1 Enhances its Antifungal Activity.
J.Biol.Chem., 287, 2012
|
|
4AB0
 
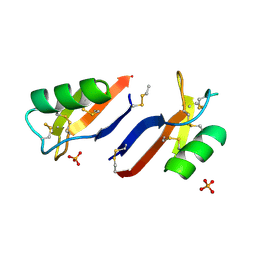 | | X-ray crystal structure of Nicotiana alata defensin NaD1 | | Descriptor: | FLOWER-SPECIFIC DEFENSIN, PHOSPHATE ION | | Authors: | Mills, G.D, Lay, F.T, Hulett, M.D, Kvansakul, M. | | Deposit date: | 2011-12-06 | | Release date: | 2012-04-25 | | Last modified: | 2012-06-20 | | Method: | X-RAY DIFFRACTION (1.636 Å) | | Cite: | Dimerization of Plant Defensin Nad1 Enhances its Antifungal Activity.
J.Biol.Chem., 287, 2012
|
|
8D2J
 
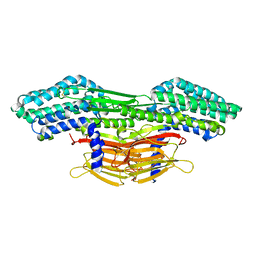 | | A novel insecticidal protein from ferns IPD113_Cow. | | Descriptor: | GLYCEROL, IPD113_Cow | | Authors: | Maher, M.J. | | Deposit date: | 2022-05-30 | | Release date: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Novel insecticidal proteins from ferns resemble insecticidal proteins from Bacillus thuringiensis.
Proc.Natl.Acad.Sci.USA, 120, 2023
|
|
8E7F
 
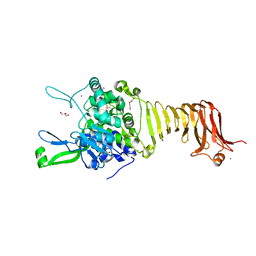 | | Crystal structure of the autotransporter Ssp from Serratia marcescens. | | Descriptor: | CALCIUM ION, DI(HYDROXYETHYL)ETHER, Extracellular serine protease, ... | | Authors: | Hor, L, Pilapitiya, A, Panjikar, S, Paxman, J.J, Heras, B. | | Deposit date: | 2022-08-23 | | Release date: | 2023-03-08 | | Last modified: | 2023-03-15 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of a subtilisin-like autotransporter passenger domain reveals insights into its cytotoxic function.
Nat Commun, 14, 2023
|
|
6DMZ
 
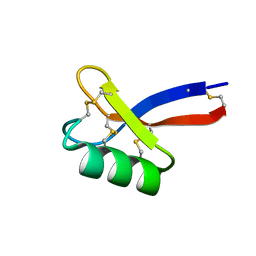 | | Solution structure of ZmD32 | | Descriptor: | Flower-specific gamma-thionin | | Authors: | Harvey, P.J, Craik, D.J. | | Deposit date: | 2018-06-05 | | Release date: | 2019-05-15 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Salt-Tolerant Antifungal and Antibacterial Activities of the Corn Defensin ZmD32.
Front Microbiol, 10, 2019
|
|
4CQK
 
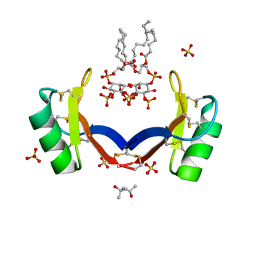 | | Crystal structure of ligand-bound NaD1 | | Descriptor: | (4R)-2-METHYLPENTANE-2,4-DIOL, FLOWER-SPECIFIC DEFENSIN, SULFATE ION, ... | | Authors: | Lay, F.T, Mills, G.M, Poon, I.K.H, Baxter, A.A, Hulett, M.D, Kvansakul, M. | | Deposit date: | 2014-02-17 | | Release date: | 2014-04-16 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Phosphoinositide-Mediated Oligomerization of a Defensin Induces Cell Lysis.
Elife, 3, 2014
|
|
6B55
 
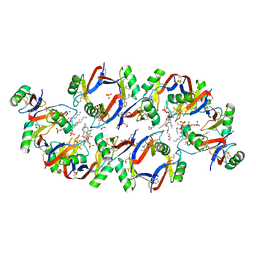 | | Crystal structure of the Plant Defensin NaD1 complexed with phosphatidic acid | | Descriptor: | 1,2-DIOCTANOYL-SN-GLYCERO-3-PHOSPHATE, 1,2-ETHANEDIOL, Flower-specific defensin, ... | | Authors: | Jarva, M, Phan, K, Humble, C, Lay, F.T, Hulett, M, Kvansakul, M. | | Deposit date: | 2017-09-28 | | Release date: | 2018-05-23 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | X-ray structure of a carpet-like antimicrobial defensin-phospholipid membrane disruption complex.
Nat Commun, 9, 2018
|
|
6O0M
 
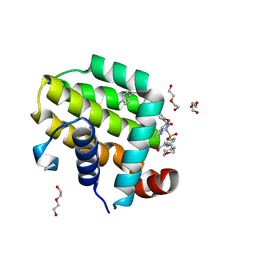 | | crystal structure of BCL-2 F104L mutation with venetoclax | | Descriptor: | 4-{4-[(4'-chloro-5,5-dimethyl[3,4,5,6-tetrahydro[1,1'-biphenyl]]-2-yl)methyl]piperazin-1-yl}-N-[(3-nitro-4-{[(oxan-4-yl )methyl]amino}phenyl)sulfonyl]-2-[(1H-pyrrolo[2,3-b]pyridin-5-yl)oxy]benzamide, Apoptosis regulator Bcl-2,Bcl-2-like protein 1,Apoptosis regulator Bcl-2, DI(HYDROXYETHYL)ETHER | | Authors: | Birkinshaw, R.W, Luo, C.S, Colman, P.M, Czabotar, P.E. | | Deposit date: | 2019-02-16 | | Release date: | 2019-05-22 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Structures of BCL-2 in complex with venetoclax reveal the molecular basis of resistance mutations.
Nat Commun, 10, 2019
|
|
6O0K
 
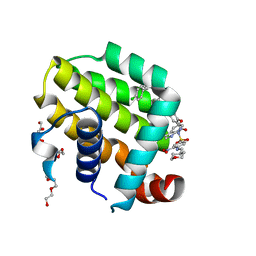 | | crystal structure of BCL-2 with venetoclax | | Descriptor: | 4-{4-[(4'-chloro-5,5-dimethyl[3,4,5,6-tetrahydro[1,1'-biphenyl]]-2-yl)methyl]piperazin-1-yl}-N-[(3-nitro-4-{[(oxan-4-yl )methyl]amino}phenyl)sulfonyl]-2-[(1H-pyrrolo[2,3-b]pyridin-5-yl)oxy]benzamide, Apoptosis regulator Bcl-2, NONAETHYLENE GLYCOL | | Authors: | Birkinshaw, R.W, Luo, C.S, Colman, P.M, Czabotar, P.E. | | Deposit date: | 2019-02-16 | | Release date: | 2019-05-22 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.62 Å) | | Cite: | Structures of BCL-2 in complex with venetoclax reveal the molecular basis of resistance mutations.
Nat Commun, 10, 2019
|
|
6O0L
 
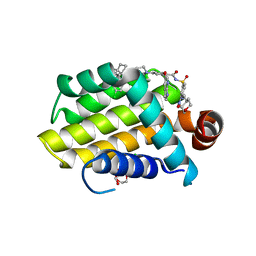 | | crystal structure of BCL-2 G101V mutation with venetoclax | | Descriptor: | 4-{4-[(4'-chloro-5,5-dimethyl[3,4,5,6-tetrahydro[1,1'-biphenyl]]-2-yl)methyl]piperazin-1-yl}-N-[(3-nitro-4-{[(oxan-4-yl )methyl]amino}phenyl)sulfonyl]-2-[(1H-pyrrolo[2,3-b]pyridin-5-yl)oxy]benzamide, Apoptosis regulator Bcl-2,Bcl-2-like protein 1,Apoptosis regulator Bcl-2, CHLORIDE ION, ... | | Authors: | Birkinshaw, R.W, Luo, C.S, Colman, P.M, Czabotar, P.E. | | Deposit date: | 2019-02-16 | | Release date: | 2019-05-22 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structures of BCL-2 in complex with venetoclax reveal the molecular basis of resistance mutations.
Nat Commun, 10, 2019
|
|
