1ZZH
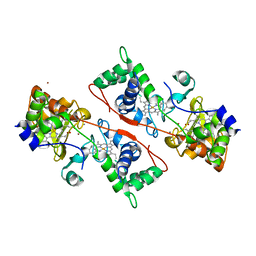 
 | | Structure of the fully oxidized di-heme cytochrome c peroxidase from R. capsulatus | | Descriptor: | CALCIUM ION, HEME C, ZINC ION, ... | | Authors: | De Smet, L, Savvides, S.N, Van Horen, E, Pettigrew, G, Van Beeumen, J.J. | | Deposit date: | 2005-06-14 | | Release date: | 2005-11-29 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structural and mutagenesis studies on the cytochrome c peroxidase from Rhodobacter capsulatus provide new insights into structure-function relationships of bacterial di-heme peroxidases
J.Biol.Chem., 281, 2006
|
|
7BFX
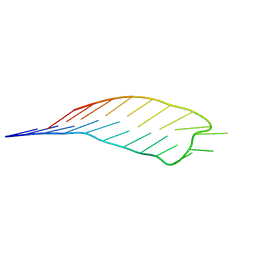 
 | | deoxyxylose nucleic acid hairpin | | Descriptor: | dXyNA (5'-D(*(XA)P*(XG)P*(XC)P*(XA)P*(XA)P*(XT)P*(XC)P*(XC)P*(XC)P*(XC)P*(XC)P*(XC)P*(XG)P*(XG)P*(XA)P*(XT)P*(XT)P*(XG)P*(XC)P*T)-3') | | Authors: | Mattelaer, C.-A, Mohitosh, M, Smets, L, Maiti, M, Schepers, G, Mattelaer, H.-P, Rosemeyer, H, Herdewijn, P, Lescrinier, E. | | Deposit date: | 2021-01-05 | | Release date: | 2021-01-27 | | Last modified: | 2024-01-31 | | Method: | SOLUTION NMR | | Cite: | Stable Hairpin Structures Formed by Xylose-Based Nucleic Acids.
Chembiochem, 22, 2021
|
|
7BFS
 
 | | deoxyxylose nucleic acid hairpin | | Descriptor: | DNA (5'-D(*AP*GP*CP*AP*AP*TP*CP*CP*(XC)P*(XC)P*(XC)P*(XC)P*GP*GP*AP*TP*TP*GP*CP*T)-3') | | Authors: | Mattelaer, C.-A, Mohitosh, M, Smets, L, Maiti, M, Schepers, G, Mattelaer, H.-P, Rosemeyer, H, Herdewijn, P, Lescrinier, E. | | Deposit date: | 2021-01-04 | | Release date: | 2021-01-27 | | Last modified: | 2021-05-12 | | Method: | SOLUTION NMR | | Cite: | Stable Hairpin Structures Formed by Xylose-Based Nucleic Acids.
Chembiochem, 22, 2021
|
|
