7RWV
 
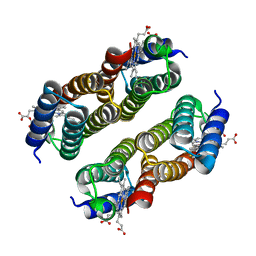 | | Crystal structure of a metal-free RIDC1 variant | | Descriptor: | HEME C, Soluble cytochrome b562 | | Authors: | Kakkis, A, Golub, E. | | Deposit date: | 2021-08-20 | | Release date: | 2022-06-29 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Redox- and metal-directed structural diversification in designed metalloprotein assemblies.
Chem.Commun.(Camb.), 58, 2022
|
|
7RWX
 
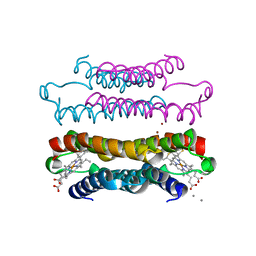 | |
7RWW
 
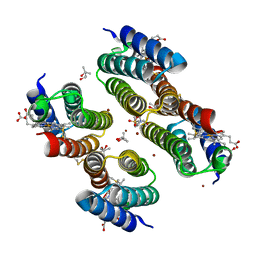 | | Crystal structure of a Zn-bound RIDC1 variant | | Descriptor: | (4R)-2-METHYLPENTANE-2,4-DIOL, (4S)-2-METHYL-2,4-PENTANEDIOL, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, ... | | Authors: | Kakkis, A, Golub, E. | | Deposit date: | 2021-08-20 | | Release date: | 2022-06-29 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Redox- and metal-directed structural diversification in designed metalloprotein assemblies.
Chem.Commun.(Camb.), 58, 2022
|
|
7RWU
 
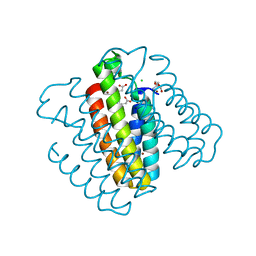 | |
7RWY
 
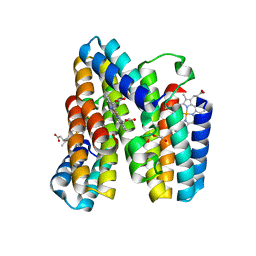 | |
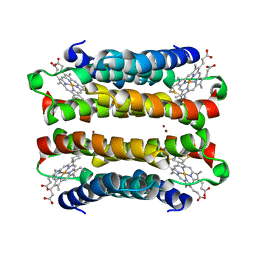
7TEP
 
 | |
6WZ0
 
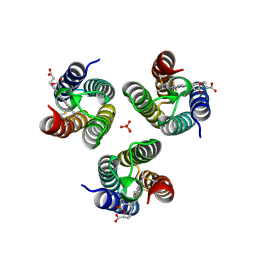 | |
6WZA
 
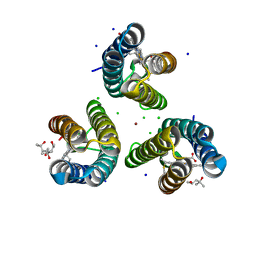 | | Ni-bound structure of an engineered metal-dependent protein trimer, TriCyt1 | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, CHLORIDE ION, HEME C, ... | | Authors: | Tezcan, F.A, Kakkis, A. | | Deposit date: | 2020-05-13 | | Release date: | 2020-09-16 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Metal-Templated Design of Chemically Switchable Protein Assemblies with High-Affinity Coordination Sites.
Angew.Chem.Int.Ed.Engl., 59, 2020
|
|
6WZ3
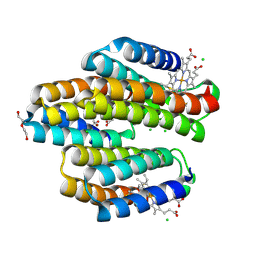 
 | | Cu-bound structure of the engineered protein trimer, TriCyt3 | | Descriptor: | CHLORIDE ION, COPPER (II) ION, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Tezcan, F.A, Kakkis, A. | | Deposit date: | 2020-05-13 | | Release date: | 2020-09-16 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Metal-Templated Design of Chemically Switchable Protein Assemblies with High-Affinity Coordination Sites.
Angew.Chem.Int.Ed.Engl., 59, 2020
|
|
7SU2
 
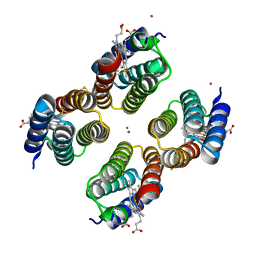 | | Crystal structure of a Co-bound RIDC1 variant | | Descriptor: | COBALT (II) ION, HEME C, Soluble cytochrome b562 | | Authors: | Golub, E, Kakkis, A. | | Deposit date: | 2021-11-15 | | Release date: | 2022-06-29 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Redox- and metal-directed structural diversification in designed metalloprotein assemblies.
Chem.Commun.(Camb.), 58, 2022
|
|
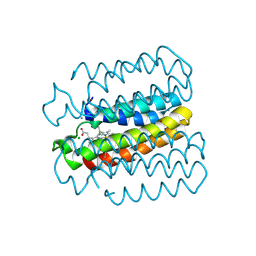
6WZC
 
 | |
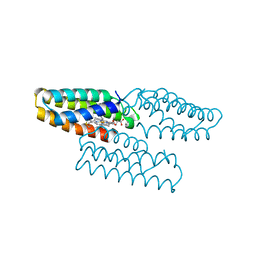
6WZ1
 
 | |
6WYU
 
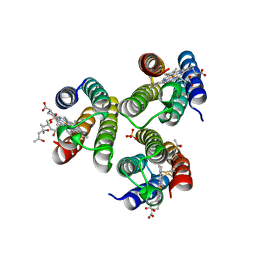 | | Crystallographic trimer of metal-free TriCyt2 | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, HEME C, SULFATE ION, ... | | Authors: | Tezcan, F.A, Kakkis, A. | | Deposit date: | 2020-05-13 | | Release date: | 2020-09-16 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | Metal-Templated Design of Chemically Switchable Protein Assemblies with High-Affinity Coordination Sites.
Angew.Chem.Int.Ed.Engl., 59, 2020
|
|
6X7E
 
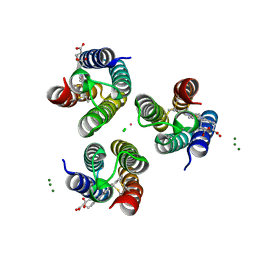 | | Co-bound structure of an engineered protein trimer, TriCyt3, with delta isomerism at the hexahistidine coordination site | | Descriptor: | CHLORIDE ION, COBALT (II) ION, HEME C, ... | | Authors: | Tezcan, F.A, Kakkis, A. | | Deposit date: | 2020-05-29 | | Release date: | 2020-09-16 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.9987 Å) | | Cite: | Metal-Templated Design of Chemically Switchable Protein Assemblies with High-Affinity Coordination Sites.
Angew.Chem.Int.Ed.Engl., 59, 2020
|
|
6X8X
 
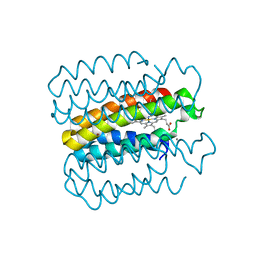 | |
6WZ2
 
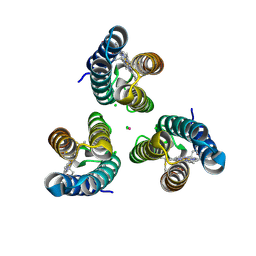 | |
6WZ7
 
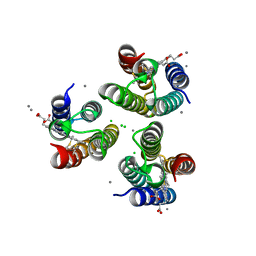 | | Mn-bound structure of a TriCyt3 variant | | Descriptor: | CALCIUM ION, CHLORIDE ION, HEME C, ... | | Authors: | Tezcan, F.A, Kakkis, A. | | Deposit date: | 2020-05-13 | | Release date: | 2020-09-16 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Metal-Templated Design of Chemically Switchable Protein Assemblies with High-Affinity Coordination Sites.
Angew.Chem.Int.Ed.Engl., 59, 2020
|
|
