1HWQ
 
 | |
1TJZ
 
 | |
1Q8N
 
 | | Solution Structure of the Malachite Green RNA Binding Aptamer | | Descriptor: | MALACHITE GREEN, RNA Aptamer | | Authors: | Flinders, J, DeFina, S.C, Brackett, D.M, Baugh, C, Wilson, C, Dieckmann, T. | | Deposit date: | 2003-08-21 | | Release date: | 2004-03-23 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | Recognition of planar and nonplanar ligands in the malachite green-RNA aptamer complex.
Chembiochem, 5, 2004
|
|
2NNT
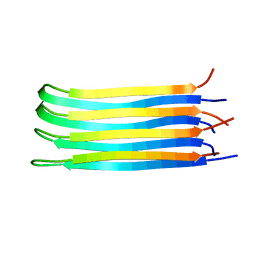 
 | | General structural motifs of amyloid protofilaments | | Descriptor: | Transcription elongation regulator 1 | | Authors: | Ferguson, N, Becker, J, Tidow, H, Tremmel, S, Sharpe, T.D, Krause, G, Flinders, J, Petrovich, M, Berriman, J, Oschkinat, H, Fersht, A.R. | | Deposit date: | 2006-10-24 | | Release date: | 2006-11-14 | | Last modified: | 2023-12-27 | | Method: | SOLID-STATE NMR | | Cite: | General structural motifs of amyloid protofilaments.
Proc.Natl.Acad.Sci.Usa, 103, 2006
|
|
