2LC6
 | |
2LVX
 
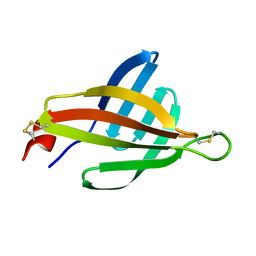 | |
2MBF
 
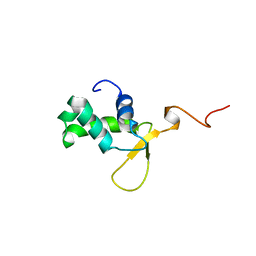 | |
2LWX
 
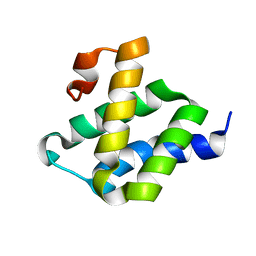 | |
2MP1
 
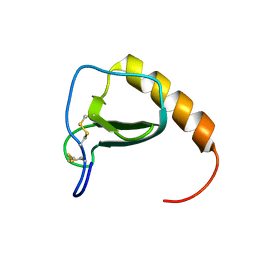 | |
2N1H
 | |
2N55
 
 | |
2GOV
 
 | | Solution structure of Murine p22HBP | | Descriptor: | Heme-binding protein 1 | | Authors: | Volkman, B.F, Dias, J.S, Goodfellow, B.J, Peterson, F.C, Center for Eukaryotic Structural Genomics (CESG) | | Deposit date: | 2006-04-14 | | Release date: | 2006-05-09 | | Last modified: | 2022-03-09 | | Method: | SOLUTION NMR | | Cite: | The First Structure from the SOUL/HBP Family of Heme-binding Proteins, Murine P22HBP.
J.Biol.Chem., 281, 2006
|
|
2G0Q
 
 | | Solution structure of At5g39720.1 from Arabidopsis thaliana | | Descriptor: | AT5G39720.1 protein | | Authors: | Volkman, B.F, Peterson, F.C, Lytle, B.L, Center for Eukaryotic Structural Genomics (CESG) | | Deposit date: | 2006-02-13 | | Release date: | 2006-02-28 | | Last modified: | 2022-03-09 | | Method: | SOLUTION NMR | | Cite: | Solution structure of Arabidopsis thaliana protein At5g39720.1, a member of the AIG2-like protein family.
Acta Crystallogr.,Sect.F, 62, 2006
|
|
2GOW
 
 | | Solution structure of BC059385 from Homo sapiens | | Descriptor: | Ubiquitin-like protein 3 | | Authors: | Volkman, B.F, de la Cruz, N.B, Lytle, B.L, Peterson, F.C, Center for Eukaryotic Structural Genomics (CESG) | | Deposit date: | 2006-04-14 | | Release date: | 2006-04-25 | | Last modified: | 2022-03-09 | | Method: | SOLUTION NMR | | Cite: | Solution structure of a membrane-anchored ubiquitin-fold (MUB) protein from Homo sapiens.
Protein Sci., 16, 2007
|
|
2HDM
 
 | |
2I9Y
 
 | | Solution structure of Arabidopsis thaliana protein At1g70830, a member of the major latex protein family | | Descriptor: | major latex protein-like protein 28 or MLP-like protein 28 | | Authors: | Volkman, B.F, de la Cruz, N.B, Lytle, B.L, Peterson, F.C, Center for Eukaryotic Structural Genomics (CESG) | | Deposit date: | 2006-09-06 | | Release date: | 2006-09-19 | | Last modified: | 2022-03-09 | | Method: | SOLUTION NMR | | Cite: | Structures of two Arabidopsis thaliana major latex proteins represent novel helix-grip folds.
Proteins, 76, 2009
|
|
2K04
 
 | |
2K01
 
 | |
2K05
 
 | |
2K03
 
 | |
2JQV
 
 | |
2JP1
 
 | |
2KOJ
 
 | |
2KEC
 
 | |
2KOM
 
 | | Solution structure of humar Par-3b PDZ2 (residues 451-549) | | Descriptor: | Partitioning defective 3 homolog | | Authors: | Volkman, B.F, Tyler, R.C, Peterson, F.C, Center for Eukaryotic Structural Genomics (CESG) | | Deposit date: | 2009-09-24 | | Release date: | 2009-11-10 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | Rapid, robotic, small-scale protein production for NMR screening and structure determination.
Protein Sci., 19, 2010
|
|
2KVB
 
 | |
2KEE
 
 | |
2KVA
 
 | |
2KOH
 
 | |
