1UEK
 
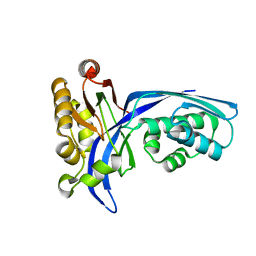 | | Crystal structure of 4-(cytidine 5'-diphospho)-2C-methyl-D-erythritol kinase | | Descriptor: | 4-(cytidine 5'-diphospho)-2C-methyl-D-erythritol kinase | | Authors: | Wada, T, Kuramitsu, S, Yokoyama, S, Tame, J.R.H, Park, S.Y, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2003-05-17 | | Release date: | 2003-06-17 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystal Structure of 4-(Cytidine 5'-diphospho)-2-C-methyl-D-erythritol kinase, an Enzyme in the Non-mevalonate Pathway of Isoprenoid Synthesis.
J.Biol.Chem., 278, 2003
|
|
3VUS
 
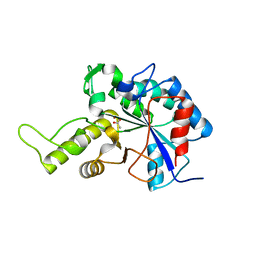 | | Escherichia coli PgaB N-terminal domain | | Descriptor: | ACETATE ION, MERCURY (II) ION, Poly-beta-1,6-N-acetyl-D-glucosamine N-deacetylase, ... | | Authors: | Nishiyama, T, Noguchi, H, Yoshida, H, Park, S.-Y, Tame, J.R.H. | | Deposit date: | 2012-07-05 | | Release date: | 2012-11-14 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | The structure of the deacetylase domain of Escherichia coli PgaB, an enzyme required for biofilm formation: a circularly permuted member of the carbohydrate esterase 4 family
Acta Crystallogr.,Sect.D, 69, 2013
|
|
1B3H
 
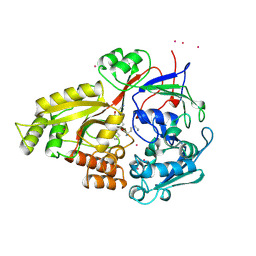 | |
1B2H
 
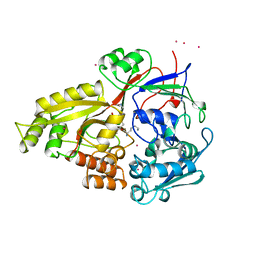 | |
1B1H
 
 | | OLIGO-PEPTIDE BINDING PROTEIN/TRIPEPTIDE (LYS HPE LYS) COMPLEX | | Descriptor: | PROTEIN (LYS HPE LYS), PROTEIN (OLIGO-PEPTIDE BINDING PROTEIN), URANYL (VI) ION | | Authors: | Davies, T.G, Tame, J.R.H. | | Deposit date: | 1998-11-10 | | Release date: | 1998-11-18 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Relating structure to thermodynamics: the crystal structures and binding affinity of eight OppA-peptide complexes.
Protein Sci., 8, 1999
|
|
1B5H
 
 | |
1B4H
 
 | |
1B6H
 
 | |
3X1Y
 
 | | Ras-related protein Rap1B(L9V) with GppNHp | | Descriptor: | CADMIUM ION, GLYCEROL, MAGNESIUM ION, ... | | Authors: | Noguchi, H, Ikegami, T, Park, S.Y, Tame, J.R.H, Unzai, S. | | Deposit date: | 2014-12-02 | | Release date: | 2015-06-03 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.168 Å) | | Cite: | The structure and conformational switching of Rap1B
Biochem.Biophys.Res.Commun., 462, 2015
|
|
3X1W
 | | Ras-related protein Rap1B with GDP | | Descriptor: | CADMIUM ION, GUANOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, ... | | Authors: | Noguchi, H, Ikegami, T, Akashi, S, Park, S.Y, Tame, J.R.H, Unzai, S. | | Deposit date: | 2014-12-02 | | Release date: | 2015-06-03 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | The structure and conformational switching of Rap1B
Biochem.Biophys.Res.Commun., 462, 2015
|
|
3X1X
 
 | | Ras-related protein Rap1B with GppNHp | | Descriptor: | CADMIUM ION, MAGNESIUM ION, PHOSPHOAMINOPHOSPHONIC ACID-GUANYLATE ESTER, ... | | Authors: | Noguchi, H, Ikegami, T, Park, S.Y, Tame, J.R.H, Unzai, S. | | Deposit date: | 2014-12-02 | | Release date: | 2015-06-03 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1 Å) | | Cite: | The structure and conformational switching of Rap1B
Biochem.Biophys.Res.Commun., 462, 2015
|
|
3X1Z
 
 | | Ras-related protein Rap1B(T65A) with GppNHp | | Descriptor: | GLYCEROL, MAGNESIUM ION, PHOSPHOAMINOPHOSPHONIC ACID-GUANYLATE ESTER, ... | | Authors: | Noguchi, H, Ikegami, T, Park, S.Y, Tame, J.R.H, Unzai, S. | | Deposit date: | 2014-12-02 | | Release date: | 2015-06-03 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.25 Å) | | Cite: | The structure and conformational switching of Rap1B
Biochem.Biophys.Res.Commun., 462, 2015
|
|
1B7H
 
 | |
1IV4
 
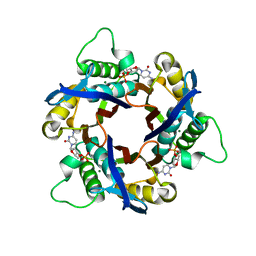 | | Structure of 2C-Methyl-D-erythritol-2,4-cyclodiphosphate Synthase (bound form Substrate) | | Descriptor: | 2-C-methyl-D-erythritol 2,4-cyclodiphosphate synthase, CYTIDINE-5'-MONOPHOSPHATE, MAGNESIUM ION | | Authors: | Kishida, H, Wada, T, Unzai, S, Kuzuyama, T, Terada, T, Sirouzu, M, Yokoyama, S, Tame, J.R.H, Park, S.-Y, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2002-03-11 | | Release date: | 2002-09-11 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Structure and catalytic mechanism of 2-C-methyl-D-erythritol 2,4-cyclodiphosphate (MECDP) synthase, an enzyme in the non-mevalonate pathway of isoprenoid synthesis.
Acta Crystallogr.,Sect.D, 59, 2003
|
|
1IV1
 
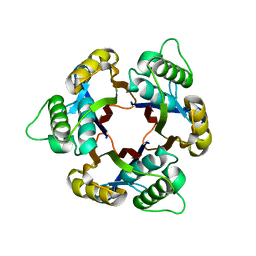 | | Structure of 2C-Methyl-D-erythritol-2,4-cyclodiphosphate Synthase | | Descriptor: | 2-C-methyl-D-erythritol 2,4-cyclodiphosphate synthase | | Authors: | Kishida, H, Wada, T, Unzai, S, Kuzuyama, T, Terada, T, Sirouzu, M, Yokoyama, S, Tame, J.R.H, Park, S.-Y, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2002-03-11 | | Release date: | 2002-09-11 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Structure and catalytic mechanism of 2-C-methyl-D-erythritol 2,4-cyclodiphosphate (MECDP) synthase, an enzyme in the non-mevalonate pathway of isoprenoid synthesis.
Acta Crystallogr.,Sect.D, 59, 2003
|
|
1IV2
 
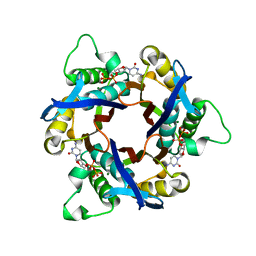 | | Structure of 2C-Methyl-D-erythritol-2,4-cyclodiphosphate Synthase (bound form CDP) | | Descriptor: | 2-C-methyl-D-erythritol 2,4-cyclodiphosphate synthase, CYTIDINE-5'-DIPHOSPHATE, MAGNESIUM ION | | Authors: | Kishida, H, Wada, T, Unzai, S, Kuzuyama, T, Terada, T, Sirouzu, M, Yokoyama, S, Tame, J.R.H, Park, S.-Y, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2002-03-11 | | Release date: | 2002-09-11 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Structure and catalytic mechanism of 2-C-methyl-D-erythritol 2,4-cyclodiphosphate (MECDP) synthase, an enzyme in the non-mevalonate pathway of isoprenoid synthesis.
Acta Crystallogr.,Sect.D, 59, 2003
|
|
1IV3
 
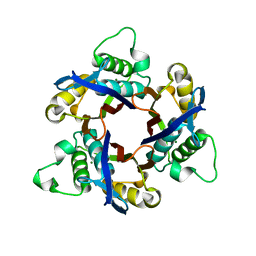 | | Structure of 2C-Methyl-D-erythritol-2,4-cyclodiphosphate Synthase (bound form MG atoms) | | Descriptor: | 2-C-methyl-D-erythritol 2,4-cyclodiphosphate synthase, MAGNESIUM ION | | Authors: | Kishida, H, Wada, T, Unzai, S, Kuzuyama, T, Terada, T, Sirouzu, M, Yokoyama, S, Tame, J.R.H, Park, S.-Y, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2002-03-11 | | Release date: | 2002-09-11 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.52 Å) | | Cite: | Structure and catalytic mechanism of 2-C-methyl-D-erythritol 2,4-cyclodiphosphate (MECDP) synthase, an enzyme in the non-mevalonate pathway of isoprenoid synthesis.
Acta Crystallogr.,Sect.D, 59, 2003
|
|
1J3M
 
 | | Crystal structure of the conserved hypothetical protein TT1751 from Thermus thermophilus HB8 | | Descriptor: | SULFITE ION, the conserved hypothetical protein TT1751 | | Authors: | Kishishita, S, Terada, T, Shirouzu, M, Kuramitsu, S, Park, S.-Y, Tame, R.H, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2003-02-05 | | Release date: | 2004-05-18 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of a conserved hypothetical protein TT1751 from Thermus thermophilus HB8
Proteins, 57, 2004
|
|
1UDX
 
 | | Crystal structure of the conserved protein TT1381 from Thermus thermophilus HB8 | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, ACETATE ION, the GTP-binding protein Obg | | Authors: | Kukimoto-Niino, M, Shirouzu, M, Murayama, K, Inoue, M, Kuramitsu, S, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2003-05-07 | | Release date: | 2004-03-16 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.07 Å) | | Cite: | Crystal structure of the GTP-binding protein Obg from Thermus thermophilus HB8.
J.Mol.Biol., 337, 2004
|
|
3HPE
 
 | |
1B0H
 
 | |
1V4W
 | | Crystal structure of bluefin tuna hemoglobin deoxy form at pH7.5 | | Descriptor: | PROTOPORPHYRIN IX CONTAINING FE, hemoglobin alpha chain, hemoglobin beta chain | | Authors: | Yokoyama, T, Chong, K.T, Miyazaki, Y, Nakatsukasa, T, Unzai, S, Miyazaki, G, Morimoto, H, Jeremy, R.H.T, Park, S.Y. | | Deposit date: | 2003-11-19 | | Release date: | 2004-07-06 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Novel Mechanisms of pH Sensitivity in Tuna Hemoglobin: A STRUCTURAL EXPLANATION OF THE ROOT EFFECT
J.Biol.Chem., 279, 2004
|
|
1V4U
 
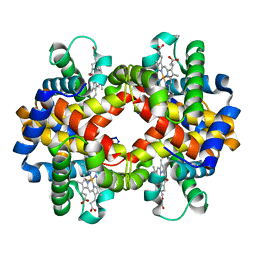 | | Crystal structure of bluefin tuna carbonmonoxy-hemoglobin | | Descriptor: | CARBON MONOXIDE, PROTOPORPHYRIN IX CONTAINING FE, hemoglobin alpha chain, ... | | Authors: | Yokoyama, T, Chong, K.T, Miyazaki, Y, Nakatsukasa, T, Unzai, S, Miyazaki, G, Morimoto, H, Jeremy, R.H.T, Park, S.Y. | | Deposit date: | 2003-11-19 | | Release date: | 2004-07-06 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Novel Mechanisms of pH Sensitivity in Tuna Hemoglobin: A STRUCTURAL EXPLANATION OF THE ROOT EFFECT
J.Biol.Chem., 279, 2004
|
|
1V4X
 
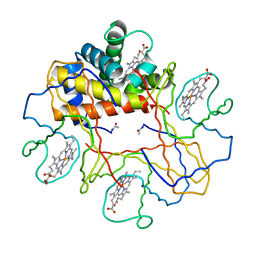 | | Crystal structure of bluefin tuna hemoglobin deoxy form at pH5.0 | | Descriptor: | PROTOPORPHYRIN IX CONTAINING FE, hemoglobin alpha chain, hemoglobin beta chain | | Authors: | Yokoyama, T, Chong, K.T, Miyazaki, Y, Nakatsukasa, T, Unzai, S, Miyazaki, G, Morimoto, H, Jeremy, R.H.T, Park, S.Y. | | Deposit date: | 2003-11-19 | | Release date: | 2004-07-06 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Novel Mechanisms of pH Sensitivity in Tuna Hemoglobin: A STRUCTURAL EXPLANATION OF THE ROOT EFFECT
J.Biol.Chem., 279, 2004
|
|
3V7A
 
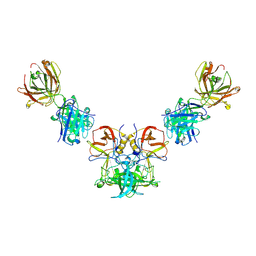 | |
