3UEX
 
 | |
3UEU
 
 | |
6QI7
 
 | | Engineered beta-lactoglobulin: variant L39Y in complex with endogenous ligand | | Descriptor: | 1,2-ETHANEDIOL, Beta-lactoglobulin, PALMITIC ACID | | Authors: | Loch, J.I, Siuda, M.K, Lewinski, K. | | Deposit date: | 2019-01-17 | | Release date: | 2019-01-30 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structure-based design approach to rational site-directed mutagenesis of beta-lactoglobulin.
J.Struct.Biol., 210, 2020
|
|
6ZSX
 
 | |
6ZSW
 | |
6RWP
 | |
6RWQ
 
 | | Engineered beta-lactoglobulin: variant F105L in complex with myristic acid | | Descriptor: | 1,2-ETHANEDIOL, Beta-lactoglobulin, MYRISTIC ACID | | Authors: | Loch, J.I, Gotkowski, M, Lewinski, K. | | Deposit date: | 2019-06-05 | | Release date: | 2019-06-19 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Structure-based design approach to rational site-directed mutagenesis of beta-lactoglobulin.
J.Struct.Biol., 210, 2020
|
|
6RWR
 
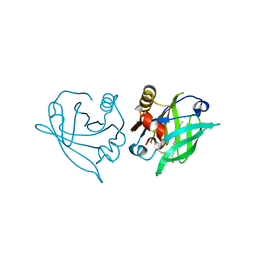 | |
8A0D
 
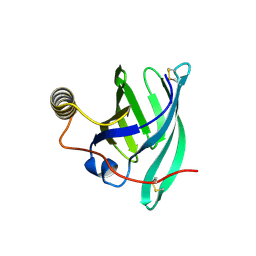 | | Crystal structure of the major guinea pig allergen Cav p 1.0101 part of the lipocalin family | | Descriptor: | Allergen lipocalin Cav p 1 isoform 1 | | Authors: | Herman, R, Charlier, P, Janssen-Weets, B, Hilger, C, Swiontek, K. | | Deposit date: | 2022-05-27 | | Release date: | 2022-08-03 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (3.685 Å) | | Cite: | Mammalian derived lipocalin and secretoglobin respiratory allergens strongly bind ligands with potentially immune modulating properties.
Front Allergy, 3, 2022
|
|
6S2S
 
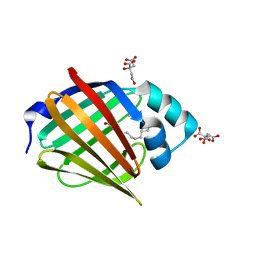 | |
3RZY
 | |
3UEV
 
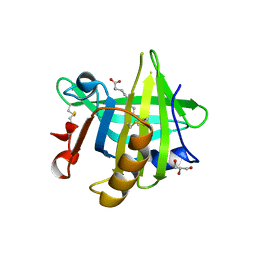 | |
3U9P
 
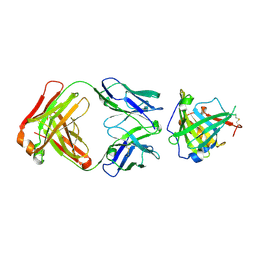 | | Crystal Structure of Murine Siderocalin in Complex with an Fab Fragment | | Descriptor: | Monoclonal Fab Fragment Heavy Chain, Monoclonal Fab Fragment Light Chain, Neutrophil gelatinase-associated lipocalin | | Authors: | Correnti, C, Strong, R.K. | | Deposit date: | 2011-10-19 | | Release date: | 2013-05-01 | | Last modified: | 2018-01-31 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Siderocalin/Lcn2/NGAL/24p3 does not drive apoptosis through gentisic acid mediated iron withdrawal in hematopoietic cell lines.
Plos One, 7, 2012
|
|
1TOW
 | | Crystal structure of human adipocyte fatty acid binding protein in complex with a carboxylic acid ligand | | Descriptor: | 4-(9H-CARBAZOL-9-YL)BUTANOIC ACID, Fatty acid-binding protein, adipocyte | | Authors: | Lehmann, F, Haile, S, Axen, E, Medina, C, Uppenberg, J, Svensson, S, Lundback, T, Rondahl, L, Barf, T. | | Deposit date: | 2004-06-15 | | Release date: | 2004-08-24 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Discovery of inhibitors of human adipocyte fatty acid-binding protein, a potential type 2 diabetes target.
Bioorg.Med.Chem.Lett., 14, 2004
|
|
1S44
 
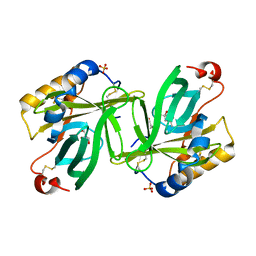 | | The structure and refinement of apocrustacyanin C2 to 1.6A resolution and the search for differences between this protein and the homologous apoproteins A1 and C1. | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, Crustacyanin A1 subunit, GLYCEROL, ... | | Authors: | Habash, J, Helliwell, J.R, Raftery, J, Cianci, M, Rizkallah, P.J, Chayen, N.E, Nneji, G.A, Zagalsky, P.F. | | Deposit date: | 2004-01-15 | | Release date: | 2004-04-27 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | The structure and refinement of apocrustacyanin C2 to 1.3 A resolution and the search for differences between this protein and the homologous apoproteins A1 and C1.
Acta Crystallogr.,Sect.D, 60, 2004
|
|
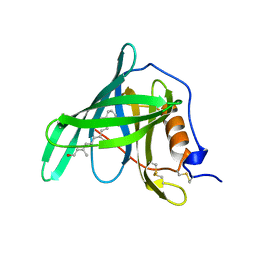
1RBP
 
 | |
1TW4
 | | Crystal Structure of Chicken Liver Basic Fatty Acid Binding Protein (Bile Acid Binding Protein) Complexed With Cholic Acid | | Descriptor: | CHOLIC ACID, Fatty acid-binding protein | | Authors: | Nichesola, D, Perduca, M, Capaldi, S, Carrizo, M.E, Righetti, P.G, Monaco, H.L. | | Deposit date: | 2004-06-30 | | Release date: | 2004-11-23 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of chicken liver basic Fatty Acid-binding protein complexed with cholic acid
Biochemistry, 43, 2004
|
|
1TOU
 | | Crystal structure of human adipocyte fatty acid binding protein in complex with a non-covalent ligand | | Descriptor: | 2-[(2-OXO-2-PIPERIDIN-1-YLETHYL)SULFANYL]-6-(TRIFLUOROMETHYL)PYRIMIDIN-4-OL, Fatty acid-binding protein, adipocyte | | Authors: | Ringom, R, Axen, E, Uppenberg, J, Lundback, T, Rondahl, L, Barf, T. | | Deposit date: | 2004-06-15 | | Release date: | 2004-08-24 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Substituted benzylamino-6-(trifluoromethyl)pyrimidin-4(1H)-ones: a novel class of selective human A-FABP inhibitors.
Bioorg.Med.Chem.Lett., 14, 2004
|
|
1TVQ
 | | Crystal Structure of Apo Chicken Liver Basic Fatty Acid Binding Protein (or Bile Acid Binding Protein) | | Descriptor: | Fatty acid-binding protein | | Authors: | Nichesola, D, Perduca, M, Capaldi, S, Carrizo, M.E, Righetti, P.G, Monaco, H.L. | | Deposit date: | 2004-06-30 | | Release date: | 2004-11-23 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of chicken liver basic Fatty Acid-binding protein complexed with cholic acid
Biochemistry, 43, 2004
|
|
1SA8
 
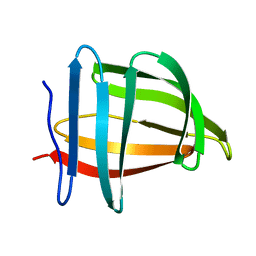 | |
1S2P
 
 | | The structure and refinement of apocrustacyanin C2 to 1.3A resolution and the search for differences between this protein and the homologous apoproteins A1 and C1 | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, Crustacyanin C2 subunit, SULFATE ION | | Authors: | Habash, J, Helliwell, J.R, Raftery, J, Cianci, M, Rizkallah, P.J, Chayen, N.E, NNeji, G.A, Zakalsky, P.F. | | Deposit date: | 2004-01-09 | | Release date: | 2004-03-02 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | The structure and refinement of apocrustacyanin C2 to 1.3 A resolution and the search for differences between this protein and the homologous apoproteins A1 and C1.
Acta Crystallogr.,Sect.D, 60, 2004
|
|
1T8V
 
 | |
1T0V
 
 | | NMR Solution Structure of the Engineered Lipocalin FluA(R95K) Northeast Structural Genomics Target OR17 | | Descriptor: | BILIN-BINDING PROTEIN | | Authors: | Mills, J.L, Liu, G, Skerra, A, Szyperski, T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2004-04-13 | | Release date: | 2005-06-14 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | NMR structure and dynamics of the engineered fluorescein-binding lipocalin FluA reveal rigidification of beta-barrel and variable loops upon enthalpy-driven ligand binding.
Biochemistry, 48, 2009
|
|
1URE
 | |
1UZ2
 
 | | The Cys121Ser Mutant of Beta-Lactoglobulin | | Descriptor: | BETA-LACTOGLOBULIN | | Authors: | McNae, I, Jayat, D, Haertle, T, Holt, C, Sawyer, L. | | Deposit date: | 2004-03-03 | | Release date: | 2004-05-20 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | A recombinant C121S mutant of bovine beta-lactoglobulin is more susceptible to peptic digestion and to denaturation by reducing agents and heating.
Biochemistry, 43, 2004
|
|
