-Search query
-Search result
Showing all 32 items for (author: liang & sy)

EMDB-40825: 
10E8-GT10.2 immunogen in complex with human Fab 10E8 and mouse Fab W6-10
Method: single particle / : Huang J, Ozorowski G, Ward AB
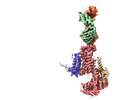
EMDB-37159: 
Cryo-EM structure of NADPH oxidase 2 in complex with p22phox and EROS
Method: single particle / : Liang SY, Liu AJ, Liu YZ, Ye RD

EMDB-40650: 
Phosphoinositide phosphate 3 kinase gamma
Method: single particle / : Chen CL, Tesmer JJG, Bandekar SJ, Cash J

EMDB-40651: 
Phosphoinositide phosphate 3 kinase gamma bound with ATP
Method: single particle / : Chen CL, Tesmer JJG, Bandekar SJ, Cash J

EMDB-40652: 
Phosphoinositide phosphate 3 kinase gamma bound with ADP
Method: single particle / : Chen CL, Tesmer JJG, Bandekar SJ, Cash J

EMDB-40653: 
Phosphoinositide phosphate 3 kinase gamma bound with ADP and Gbetagamma
Method: single particle / : Chen CL, Tesmer JJG, Bandekar SJ, Cash J

EMDB-40654: 
Phosphoinositide phosphate 3 kinase gamma bound with ADP and two Gbetagamma subunits in State 1
Method: single particle / : Chen CL, Tesmer JJG, Bandekar SJ, Cash J

EMDB-40655: 
Phosphoinositide phosphate 3 kinase gamma bound with ADP and two Gbetagamma subunits in State 2
Method: single particle / : Chen CL, Tesmer JJG, Bandekar SJ, Cash J

PDB-8so9: 
Phosphoinositide phosphate 3 kinase gamma
Method: single particle / : Chen CL, Tesmer JJG

PDB-8soa: 
Phosphoinositide phosphate 3 kinase gamma bound with ATP
Method: single particle / : Chen CL, Tesmer JJG

PDB-8sob: 
Phosphoinositide phosphate 3 kinase gamma bound with ADP
Method: single particle / : Chen CL, Tesmer JJG

PDB-8soc: 
Phosphoinositide phosphate 3 kinase gamma bound with ADP and Gbetagamma
Method: single particle / : Chen CL, Tesmer JJG

PDB-8sod: 
Phosphoinositide phosphate 3 kinase gamma bound with ADP and two Gbetagamma subunits in State 1
Method: single particle / : Chen CL, Tesmer JJG

PDB-8soe: 
Phosphoinositide phosphate 3 kinase gamma bound with ADP and two Gbetagamma subunits in State 2
Method: single particle / : Chen CL, Tesmer JJG

EMDB-38795: 
Cryo-EM structure of the [Pyr1]-apelin-13-bound human APLNR-Gi complex
Method: single particle / : Wang W, Ji S, Zhang Y

EMDB-38794: 
Cryo-EM structure of the WN561-bound human APLNR-Gi complex
Method: single particle / : Wang W, Ji S, Zhang Y

EMDB-38796: 
Cryo-EM structure of the MM07-bound human APLNR-Gi complex
Method: single particle / : Wang W, Ji S, Zhang Y

EMDB-38797: 
Cryo-EM structure of the CMF-019-bound human APLNR-Gi complex
Method: single particle / : Wang W, Ji S, Zhang Y

EMDB-38798: 
Cryo-EM structure of the WN353-bound human APLNR-Gi complex
Method: single particle / : Wang W, Ji S, Zhang Y

EMDB-33320: 
Cryo-EM map of hMCM-DH R195A/L209G mutant
Method: single particle / : Li J, Dong JQ, Dang SY, Zhai YL

EMDB-24077: 
CryoEM structure of neutralizing nanobody Nb30 in complex with SARS-CoV2 spike
Method: single particle / : Xu K, Kwong PD

EMDB-24078: 
CryoEM structure of neutralizing nanobody Nb12 in complex with SARS-CoV2 spike
Method: single particle / : Xu K, Kwong PD

PDB-7my2: 
CryoEM structure of neutralizing nanobody Nb30 in complex with SARS-CoV2 spike
Method: single particle / : Xu K, Kwong PD

PDB-7my3: 
CryoEM structure of neutralizing nanobody Nb12 in complex with SARS-CoV2 spike
Method: single particle / : Xu K, Kwong PD

EMDB-21992: 
GLP-1 peptide hormone bound to Glucagon-Like peptide-1 (GLP-1) Receptor
Method: single particle / : Belousoff MJ, Zhang X, Danev R

EMDB-21993: 
Non peptide agonist CHU-128, bound to Glucagon-Like peptide-1 (GLP-1) Receptor
Method: single particle / : Belousoff MJ, Zhang X, Danev R

EMDB-21994: 
Non peptide agonist PF-06882961, bound to Glucagon-Like peptide-1 (GLP-1) Receptor
Method: single particle / : Belousoff MJ, Zhang X, Danev R

EMDB-5698: 
Structure of NtrC1 ATPase in complex with Sigma-54 and promoter DNA
Method: single particle / : Chowdhury S, Sysoeva TA, Guo L, Nixon BT

PDB-3j0h: 
Fitting of the bacteriophage phiKZ gp29PR structure into the cryo-EM density map of the phiKZ extended tail sheath
Method: single particle / : Aksyuk AA, Fokine A, Kurochkina LP, Mesyanzhinov VV, Rossmann MG

PDB-3j0i: 
Fitting of the phiKZ gp29PR structure into the cryo-EM density map of the phiKZ polysheath
Method: single particle / : Aksyuk AA, Fokine A, Kurochkina LP, Mesyanzhinov VV, Rossmann MG

EMDB-5331: 
The cryoEM structure of the bacteriophage phiKZ polysheath
Method: single particle / : Aksyuk AA, Kurochkina LP, Fokine A, Mesyanzhinov VV, Rossmann MG

EMDB-5332: 
The cryoEM structure of the bacteriophage phiKZ polysheath
Method: single particle / : Aksyuk AA, Kurochkina LP, Fokine A, Mesyanzhinov VV, Rossmann MG
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
