-Search query
-Search result
Showing 1 - 50 of 74 items for (author: haupt & m)

EMDB-18881: 
AL amyloid fibril from the FOR010 light chain
Method: helical / : Pfeiffer PB, Banerjee S, Schmidt M, Faendrich M

EMDB-19818: 
AL amyloid fibril from the FOR103 light chain
Method: helical / : Pfeiffer PB, Karimi-Farsijani S, Kupfer N, Schmidt M, Faendrich M

PDB-8r47: 
AL amyloid fibril from the FOR010 light chain
Method: helical / : Pfeiffer PB, Banerjee S, Schmidt M, Faendrich M

PDB-9eme: 
AL amyloid fibril from the FOR103 light chain
Method: helical / : Pfeiffer PB, Karimi-Farsijani S, Kupfer N, Schmidt M, Faendrich M

EMDB-17010: 
CryoEM Structure INO80core Hexasome complex Rvb core refinement state1
Method: single particle / : Zhang M, Jungblut A, Hoffmann T, Eustermann S

PDB-8ooc: 
CryoEM Structure INO80core Hexasome complex Rvb core refinement state1
Method: single particle / : Zhang M, Jungblut A, Hoffmann T, Eustermann S

EMDB-17006: 
CryoEM Structure INO80core Hexasome complex composite map state1
Method: single particle / : Zhang M, Jungblut A, Hoffmann T, Eustermann S

EMDB-17007: 
CryoEM Structure INO80core Hexasome complex ATPase-DNA refinement state1
Method: single particle / : Zhang M, Jungblut A, Hoffmann T, Eustermann S

EMDB-17008: 
CryoEM Structure INO80core Hexasome complex Hexasome refinement state1
Method: single particle / : Zhang M, Jungblut A, Hoffmann T, Eustermann S

EMDB-17012: 
CryoEM Structure INO80core Hexasome complex Arp5 Ies6 refinement state1
Method: single particle / : Zhang M, Jungblut A, Hoffmann T, Eustermann S

EMDB-17017: 
CryoEM Structure INO80core Hexasome complex Arp5 grappler refinement state1
Method: single particle / : Zhang M, Jungblut A, Hoffmann T, Eustermann S

EMDB-17025: 
INO80 core bound to hexasome composite map of state 2
Method: single particle / : Zhang M, Jungblut A, Hoffmann T, Eustermann S

EMDB-17026: 
CryoEM Structure INO80core Hexasome complex Rvb core refinement state2
Method: single particle / : Zhang M, Jungblut A, Hoffmann T, Eustermann S

EMDB-17027: 
CryoEM Structure INO80core Hexasome complex ATPase-hexasome refinement state 2
Method: single particle / : Zhang M, Jungblut A, Hoffmann T, Eustermann S

EMDB-17028: 
CryoEM Structure INO80core Hexasome complex Arp5 Ies6 refinement state2
Method: single particle / : Zhang M, Jungblut A, Hoffmann T, Eustermann S

EMDB-17676: 
INO80 core bound to hexasome focused refinement of Arp5 grappler
Method: single particle / : Zhang M, Jungblut A, Hoffmann T, Eustermann S

PDB-8oo7: 
CryoEM Structure INO80core Hexasome complex composite model state1
Method: single particle / : Zhang M, Jungblut A, Hoffmann T, Eustermann S

PDB-8oo9: 
CryoEM Structure INO80core Hexasome complex ATPase-DNA refinement state1
Method: single particle / : Zhang M, Jungblut A, Hoffmann T, Eustermann S

PDB-8ooa: 
CryoEM Structure INO80core Hexasome complex Hexasome refinement state1
Method: single particle / : Zhang M, Jungblut A, Hoffmann T, Eustermann S

PDB-8oof: 
CryoEM Structure INO80core Hexasome complex Arp5 Ies6 refinement state1
Method: single particle / : Zhang M, Jungblut A, Hoffmann T, Eustermann S

PDB-8ook: 
CryoEM Structure INO80core Hexasome complex Arp5 grappler refinement state1
Method: single particle / : Zhang M, Jungblut A, Hoffmann T, Eustermann S

PDB-8oop: 
CryoEM Structure INO80core Hexasome complex composite model state2
Method: single particle / : Zhang M, Jungblut A, Hoffmann T, Eustermann S

PDB-8oor: 
CryoEM Structure INO80core Hexasome complex Rvb core refinement state2
Method: single particle / : Zhang M, Jungblut A, Hoffmann T, Eustermann S

PDB-8oos: 
CryoEM Structure INO80core Hexasome complex ATPase-hexasome refinement state 2
Method: single particle / : Zhang M, Jungblut A, Hoffmann T, Eustermann S

PDB-8oot: 
CryoEM Structure INO80core Hexasome complex Arp5 Ies6 refinement state2
Method: single particle / : Zhang M, Jungblut A, Hoffmann T, Eustermann S

EMDB-17019: 
CryoEM Structure INO80core Hexasome complex overall refinement state1
Method: single particle / : Zhang M, Jungblut A, Hoffmann T, Eustermann S

EMDB-17023: 
CryoEM Structure INO80core Hexasome complex ATPase-hexasome refinement state 1
Method: single particle / : Zhang M, Jungblut A, Hoffmann T, Eustermann S

EMDB-17029: 
CryoEM Structure INO80core Hexasome complex overall refinement state2
Method: single particle / : Zhang M, Jungblut A, Hoffmann T, Eustermann S

EMDB-17032: 
CryoEM Structure INO80 hexasome complex Arp8 module bound to DNA
Method: single particle / : Zhang M, Jungblut A, Hoffmann T, Eustermann S

EMDB-16713: 
Local refinement map of TFIIIC TauB-DNA monomer
Method: single particle / : Wolfram SD, Mathias G, Luis H, Thomas H, Sebastian E, Christoph M
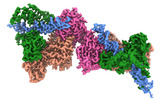
EMDB-16714: 
TFIIIC TauB-DNA dimer
Method: single particle / : Wolfram SD, Mathias G, Luis H, Thomas H, Sebastian E, Christoph M
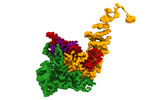
EMDB-16715: 
TFIIIC TauA complex map
Method: single particle / : Wolfram SD, Mathias G, Luis H, Thomas H, Sebastian E, Christoph M
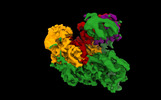
EMDB-16716: 
TFIIIC TauA complex map (sample without DNA)
Method: single particle / : Seifert-Davila W, Girbig M, Hauptmann L, Hoffmann T, Eustermann S, Mueller CW
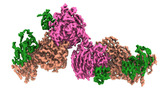
EMDB-16717: 
Structural insights into human TFIIIC promoter recognition
Method: single particle / : Wolfram SD, Mathias G, Luis H, Thomas H, Sebastian E, Christoph M

EMDB-17446: 
Consensus map of TauB-DNA dimer (Nu-refinement)
Method: single particle / : Seifert-Davila W, Girbig M, Hauptmann L, Hoffmann T, Eustermann S, Mueller CW

EMDB-17447: 
Local refinement map of TFIIIC TauB-DNA monomer 2
Method: single particle / : Seifert-Davila W, Girbig M, Hauptmann L, Hoffmann T, Eustermann S, Mueller CW

PDB-8cli: 
TFIIIC TauB-DNA monomer
Method: single particle / : Seifert-Davila W, Girbig M, Hauptmann L, Hoffmann T, Eustermann S, Mueller CW

PDB-8clj: 
TFIIIC TauB-DNA dimer
Method: single particle / : Seifert-Davila W, Girbig M, Hauptmann L, Hoffmann T, Eustermann S, Mueller CW

PDB-8clk: 
TFIIIC TauA complex
Method: single particle / : Seifert-Davila W, Girbig M, Hauptmann L, Hoffmann T, Eustermann S, Mueller CW

PDB-8cll: 
Structural insights into human TFIIIC promoter recognition
Method: single particle / : Seifert-Davila W, Girbig M, Hauptmann L, Hoffmann T, Eustermann S, Mueller CW

EMDB-27386: 
Cryo-EM structure of the zebrafish two pore domain K+ channel TREK1 (K2P2.1) in DDM detergent
Method: single particle / : Schmidpeter PAM, Nimigean CM, Riegelhaupt PM

EMDB-27387: 
Cryo-EM structure of the zebrafish two pore domain K+ channel TREK1 (K2P2.1) in DDM/POPA mixed micelles
Method: single particle / : Schmidpeter PAM, Nimigean CM, Riegelhaupt PM

EMDB-27388: 
Cryo-EM structure of the zebrafish two pore domain K+ channel TREK1 (K2P2.1) in DDM/POPE mixed micelles
Method: single particle / : Schmidpeter PAM, Nimigean CM, Riegelhaupt PM

PDB-8de7: 
Cryo-EM structure of the zebrafish two pore domain K+ channel TREK1 (K2P2.1) in DDM detergent
Method: single particle / : Schmidpeter PAM, Nimigean CM, Riegelhaupt PM

PDB-8de8: 
Cryo-EM structure of the zebrafish two pore domain K+ channel TREK1 (K2P2.1) in DDM/POPA mixed micelles
Method: single particle / : Schmidpeter PAM, Nimigean CM, Riegelhaupt PM

PDB-8de9: 
Cryo-EM structure of the zebrafish two pore domain K+ channel TREK1 (K2P2.1) in DDM/POPE mixed micelles
Method: single particle / : Schmidpeter PAM, Nimigean CM, Riegelhaupt PM

EMDB-14771: 
Amyloid fibril from human systemic AA amyloidosis (vascular variant)
Method: helical / : Banerjee S, Schmidt M, Faendrich M

PDB-7zky: 
Amyloid fibril from human systemic AA amyloidosis (vascular variant)
Method: helical / : Banerjee S, Schmidt M, Faendrich M

EMDB-25916: 
SthK closed state, cAMP-bound in the presence of POPA
Method: single particle / : Schmidpeter PA, Nimigean CM

EMDB-25917: 
SthK open state, cAMP-bound in the presence of POPA
Method: single particle / : Schmidpeter PA, Nimigean CM
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
