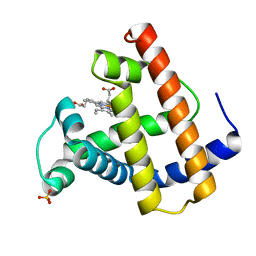
2MGB
 
 | |
2MGL
 
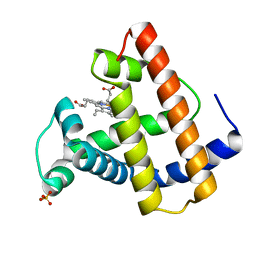 | |
1GOB
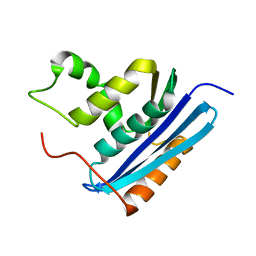 
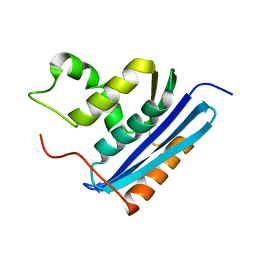
 | | COOPERATIVE STABILIZATION OF ESCHERICHIA COLI RIBONUCLEASE HI BY INSERTION OF GLY-80B AND GLY-77-> ALA SUBSTITUTION | | Descriptor: | RIBONUCLEASE H | | Authors: | Ishikawa, K, Kimura, S, Nakamura, H, Morikawa, K, Kanaya, S. | | Deposit date: | 1993-05-10 | | Release date: | 1994-01-31 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Cooperative stabilization of Escherichia coli ribonuclease HI by insertion of Gly-80b and Gly-77-->Ala substitution.
Biochemistry, 32, 1993
|
|
1GOC
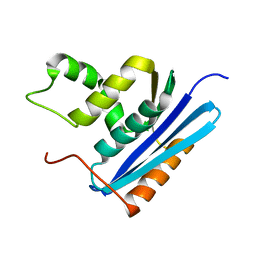 
 | | COOPERATIVE STABILIZATION OF ESCHERICHIA COLI RIBONUCLEASE HI BY INSERTION OF GLY-80B AND GLY-77-> ALA SUBSTITUTION | | Descriptor: | RIBONUCLEASE H | | Authors: | Ishikawa, K, Kimura, S, Nakamura, H, Morikawa, K, Kanaya, S. | | Deposit date: | 1993-05-10 | | Release date: | 1994-01-31 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Cooperative stabilization of Escherichia coli ribonuclease HI by insertion of Gly-80b and Gly-77-->Ala substitution.
Biochemistry, 32, 1993
|
|
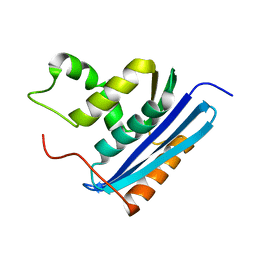
1LAW
 
 | |
1GOA
 
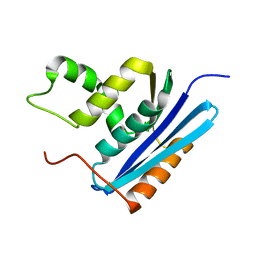 | | COOPERATIVE STABILIZATION OF ESCHERICHIA COLI RIBONUCLEASE HI BY INSERTION OF GLY-80B AND GLY-77-> ALA SUBSTITUTION | | Descriptor: | RIBONUCLEASE H | | Authors: | Ishikawa, K, Kimura, S, Nakamura, H, Morikawa, K, Kanaya, S. | | Deposit date: | 1993-05-10 | | Release date: | 1994-01-31 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Cooperative stabilization of Escherichia coli ribonuclease HI by insertion of Gly-80b and Gly-77-->Ala substitution.
Biochemistry, 32, 1993
|
|
1LAV
 
 | |
1ETA
 
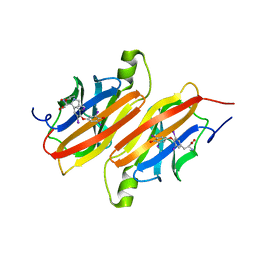 | | THE X-RAY CRYSTAL STRUCTURE REFINEMENTS OF NORMAL HUMAN TRANSTHYRETIN AND THE AMYLOIDOGENIC VAL 30-->MET VARIANT TO 1.7 ANGSTROMS RESOLUTION | | Descriptor: | 3,5,3',5'-TETRAIODO-L-THYRONINE, TRANSTHYRETIN | | Authors: | Braden, B.C, Steinrauf, L.K, Hamilton, J.A. | | Deposit date: | 1993-05-12 | | Release date: | 1995-01-26 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | The x-ray crystal structure refinements of normal human transthyretin and the amyloidogenic Val-30-->Met variant to 1.7-A resolution.
J.Biol.Chem., 268, 1993
|
|
1ETB
 
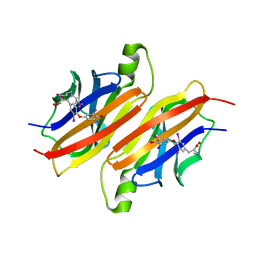 | | THE X-RAY CRYSTAL STRUCTURE REFINEMENTS OF NORMAL HUMAN TRANSTHYRETIN AND THE AMYLOIDOGENIC VAL 30-->MET VARIANT TO 1.7 ANGSTROMS RESOLUTION | | Descriptor: | 3,5,3',5'-TETRAIODO-L-THYRONINE, TRANSTHYRETIN | | Authors: | Braden, B.C, Steinrauf, L.K, Hamilton, J.A. | | Deposit date: | 1993-05-12 | | Release date: | 1995-01-26 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | The x-ray crystal structure refinements of normal human transthyretin and the amyloidogenic Val-30-->Met variant to 1.7-A resolution.
J.Biol.Chem., 268, 1993
|
|
1RGC
 
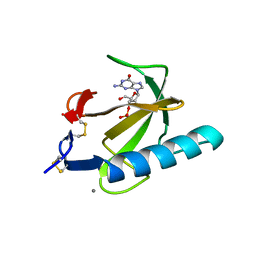 | | THE COMPLEX BETWEEN RIBONUCLEASE T1 AND 3'-GUANYLIC ACID SUGGESTS GEOMETRY OF ENZYMATIC REACTION PATH. AN X-RAY STUDY | | Descriptor: | CALCIUM ION, GUANOSINE-3'-MONOPHOSPHATE, RIBONUCLEASE T1 | | Authors: | Heydenreich, A, Koellner, G, Choe, H.W, Cordes, F, Kisker, C, Schindelin, H, Adamiak, R, Hahn, U, Saenger, W. | | Deposit date: | 1993-05-12 | | Release date: | 1994-01-31 | | Last modified: | 2017-11-29 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The complex between ribonuclease T1 and 3'GMP suggests geometry of enzymic reaction path. An X-ray study.
Eur.J.Biochem., 218, 1993
|
|
1RBL
 
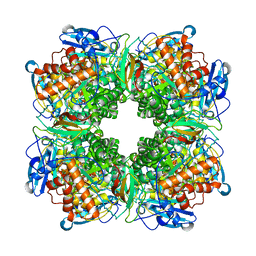 | | STRUCTURE DETERMINATION AND REFINEMENT OF RIBULOSE 1,5 BISPHOSPHATE CARBOXYLASE(SLASH)OXYGENASE FROM SYNECHOCOCCUS PCC6301 | | Descriptor: | 2-CARBOXYARABINITOL-1,5-DIPHOSPHATE, FORMIC ACID, MAGNESIUM ION, ... | | Authors: | Newman, J, Gutteridge, S, Branden, C.-I, Jones, T.A. | | Deposit date: | 1993-05-12 | | Release date: | 1994-06-22 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structure determination and refinement of ribulose 1,5-bisphosphate carboxylase/oxygenase from Synechococcus PCC6301.
Acta Crystallogr.,Sect.D, 49, 1993
|
|
1CSQ
 
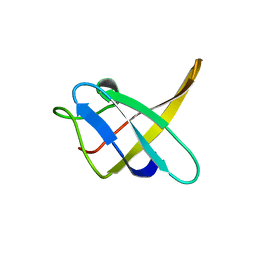 | |
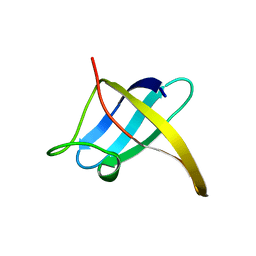
1CSP
 | |
1EDB
 
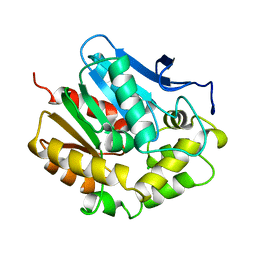 | |
1DYF
 
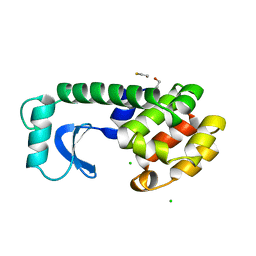 | |
1DYC
 
 | |
1DYD
 
 | |
1DYB
 
 | |
1EDD
 
 | |
1DYE
 
 | |
1DYA
 
 | |
1DYG
 
 | |
1GCA
 
 | |
1EDE
 
 | |
1RTP
 
 | | REFINED X-RAY STRUCTURE OF RAT PARVALBUMIN, A MAMMALIAN ALPHA-LINEAGE PARVALBUMIN, AT 2.0 A RESOLUTION | | Descriptor: | ALPHA-PARVALBUMIN, CALCIUM ION | | Authors: | Mcphalen, C.A, Sielecki, A.R, Santarsiero, B.D, James, M.N.G. | | Deposit date: | 1993-05-14 | | Release date: | 1994-01-31 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Refined crystal structure of rat parvalbumin, a mammalian alpha-lineage parvalbumin, at 2.0 A resolution.
J.Mol.Biol., 235, 1994
|
|
