3GIW
 
 | |
3G7P
 
 | |
3GB0
 
 | |
3HWU
 
 | |
3I09
 
 | |
3HTV
 
 | |
3HOI
 
 | |
3HLZ
 
 | |
3HM4
 
 | |
3HZP
 
 | |
3I0Y
 
 | |
3HMZ
 
 | |
3HN5
 
 | |
3HWR
 
 | |
3HVY
 
 | |
3HX8
 
 | |
3HTY
 
 | |
3I4G
 
 | |
3HN0
 
 | |
3HOS
 
 | |
3IEE
 | |
3ISY
 
 | |
8EM4
 
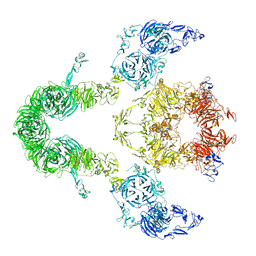 | | Cryo-EM structure of LRP2 at pH 7.5 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-galactopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Beenken, A, Cerutti, G, Brasch, J, Fitzpatrick, A.W, Barasch, J, Shapiro, L. | | Deposit date: | 2022-09-26 | | Release date: | 2023-02-08 | | Last modified: | 2023-03-08 | | Method: | ELECTRON MICROSCOPY (2.83 Å) | | Cite: | Structures of LRP2 reveal a molecular machine for endocytosis.
Cell, 186, 2023
|
|
3HBZ
 
 | |
3H3H
 
 | |
