3PAY
 
 | |
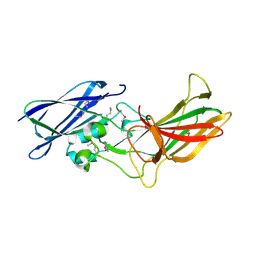
3UP6
 
 | |
4K4K
 
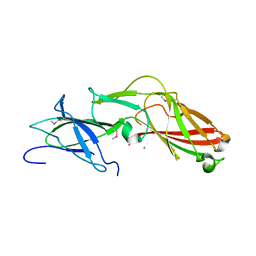 | |
5CAG
 | |
4JG5
 
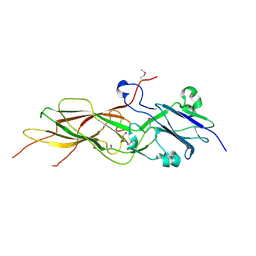 | |
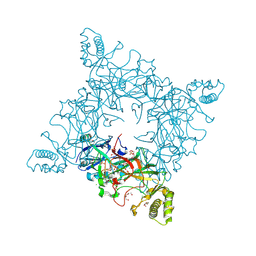
4JRF
 
 | |
4Q98
 
 | |
4QB7
 
 | |
4RDB
 
 | |
4H40
 
 | |
4GPV
 
 | |
4QDG
 
 | |
7QYI
 
 | |
3SY6
 
 | |
3T2L
 | |
3R4R
 
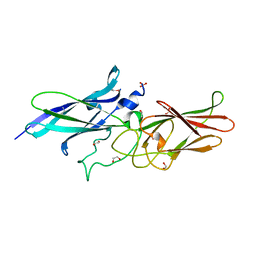 | |
8FK0
 
 | |
6IXZ
 | |
8DFT
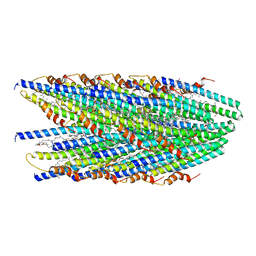 
 | | Cryo-EM structure of conjugative pili from Pyrobaculum calidifontis | | Descriptor: | Pilin protein, [(2~{S},7~{S},11~{S},15~{S},19~{R},22~{R},26~{S},30~{R},34~{R},38~{S},43~{S},47~{S},51~{S},55~{R},58~{R},62~{S},66~{R},70~{R})-38-(hydroxymethyl)-7,11,15,19,22,26,30,34,43,47,51,55,58,62,66,70-hexadecamethyl-1,4,37,40-tetraoxacyclodoheptacont-2-yl]methanol | | Authors: | Beltran, L.C, Egelman, E.H. | | Deposit date: | 2022-06-22 | | Release date: | 2023-03-22 | | Last modified: | 2024-06-12 | | Method: | ELECTRON MICROSCOPY (4.1 Å) | | Cite: | Archaeal DNA-import apparatus is homologous to bacterial conjugation machinery.
Nat Commun, 14, 2023
|
|
2WTS
 
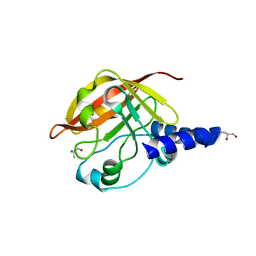 | | Crystal structure of sortase C-1 (SrtC-1) mutant H131D from S. pneumoniae | | Descriptor: | ALANINE, GLYCEROL, PUTATIVE SORTASE | | Authors: | Manzano, C, Izore, T, Job, V, DiGuilmi, A.M, Dessen, A. | | Deposit date: | 2009-09-21 | | Release date: | 2010-09-01 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Sortase Activity is Controlled by a Flexible Lid in the Pilus Biogenesis Mechanism of Gram-Positive Pathogens.
Biochemistry, 48, 2009
|
|
4EPS
 
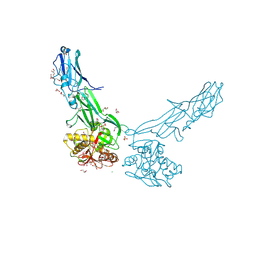 | |
8FJ5
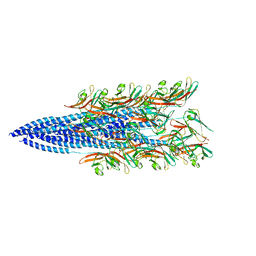 
 | | Structure of the Haloferax volcanii archaeal type IV pilus | | Descriptor: | Pilin_N domain-containing protein | | Authors: | Wang, F, Kreutzberger, M.A, Baquero, D.P, Krupovic, M, Egelman, E.H. | | Deposit date: | 2022-12-19 | | Release date: | 2023-06-28 | | Last modified: | 2024-06-19 | | Method: | ELECTRON MICROSCOPY (2.9 Å) | | Cite: | The evolution of archaeal flagellar filaments.
Proc.Natl.Acad.Sci.USA, 120, 2023
|
|
8FJS
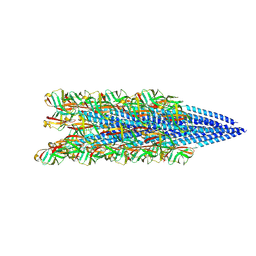 
 | |
4DGU
 
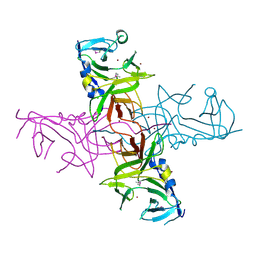 | |
6NM5
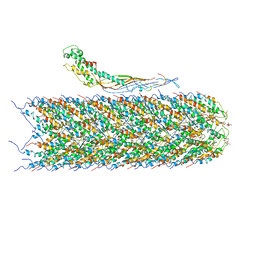 
 | | F-pilus/MS2 Maturation protein complex | | Descriptor: | (2R)-2,3-dihydroxypropyl ethyl hydrogen (S)-phosphate, Maturation protein, Type IV conjugative transfer system pilin TraA | | Authors: | Meng, R, Chang, J, Zhang, J. | | Deposit date: | 2019-01-10 | | Release date: | 2019-07-24 | | Last modified: | 2024-03-20 | | Method: | ELECTRON MICROSCOPY (6.2 Å) | | Cite: | Structural basis for the adsorption of a single-stranded RNA bacteriophage.
Nat Commun, 10, 2019
|
|
