2M8W
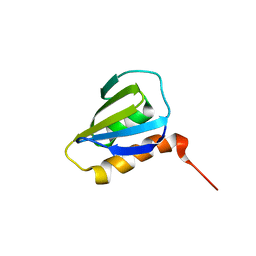 
 | | Restrained CS-Rosetta Solution NMR Structure of Staphylococcus aureus protein SAV1430. Northeast Structural Genomics Target ZR18. Structure determination | | Descriptor: | Uncharacterized protein | | Authors: | Mao, B, Tejero, R.T, Aramini, J.M, Snyder, D.A, Montelione, G.T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2013-05-29 | | Release date: | 2013-08-21 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | PDBStat: a universal restraint converter and restraint analysis software package for protein NMR.
J.Biomol.Nmr, 56, 2013
|
|
2M8X
 
 | | Restrained CS-Rosetta Solution NMR structure of the CARDB domain of PF1109 from Pyrococcus furiosus. Northeast Structural Genomics Consortium target PfR193A | | Descriptor: | Uncharacterized protein | | Authors: | Mao, B, Tejero, R.T, Aramini, J.M, Snyder, D.A, Montelione, G.T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2013-05-29 | | Release date: | 2013-08-21 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | PDBStat: a universal restraint converter and restraint analysis software package for protein NMR.
J.Biomol.Nmr, 56, 2013
|
|
2M6Q
 
 | | Refined Solution NMR Structure of Staphylococcus aureus protein SAV1430. Northeast Strucutral Genomics Consortium Target ZR18 | | Descriptor: | SAV1430 | | Authors: | Baran, M.C, Aramini, J.M, Huang, Y.J, Xiao, R, Acton, T.B, Shih, L, Montelione, G.T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2013-04-08 | | Release date: | 2013-05-01 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | PDBStat: a universal restraint converter and restraint analysis software package for protein NMR.
J.Biomol.Nmr, 56, 2013
|
|
3VUV
 
 | | Crystal structure of the merozoite surface protein MSPDBL2 from P. falciparum bound to zinc | | Descriptor: | Erythrocyte membrane protein, putative, ZINC ION | | Authors: | Czabotar, P.E, Hodder, A.N, Clarke, O.B, Lin, C.S, Smith, B.J, Cowman, A.F. | | Deposit date: | 2012-07-09 | | Release date: | 2012-08-08 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.114 Å) | | Cite: | Insights into Duffy binding-like domains through the crystal structure and function of the merozoite surface protein MSPDBL2 from Plasmodium falciparum
J.Biol.Chem., 287, 2012
|
|
3VUU
 
 | | Crystal structure of the merozoite surface protein MSPDBL2 from P. falciparum | | Descriptor: | CHLORIDE ION, Erythrocyte membrane protein, putative | | Authors: | Czabotar, P.E, Hodder, A.N, Clarke, O.B, Lin, C.S, Smith, B.J, Cowman, A.F. | | Deposit date: | 2012-07-09 | | Release date: | 2012-08-08 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.093 Å) | | Cite: | Insights into Duffy binding-like domains through the crystal structure and function of the merozoite surface protein MSPDBL2 from Plasmodium falciparum
J.Biol.Chem., 287, 2012
|
|
5M1Y
 
 | |
4B6Z
 
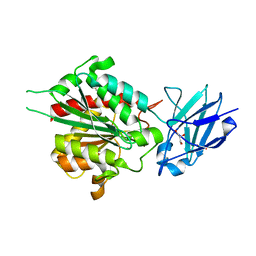 | | Crystal structure of metallo-carboxypeptidase from Burkholderia cenocepacia | | Descriptor: | 1,2-ETHANEDIOL, ACETATE ION, CHLORIDE ION, ... | | Authors: | Rimsa, V, Eadsforth, T.C, Joosten, R.P, Hunter, W.N. | | Deposit date: | 2012-08-16 | | Release date: | 2013-03-27 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | High-Resolution Structure of the M14-Type Cytosolic Carboxypeptidase from Burkholderia Cenocepacia Refined Exploiting Pdb_Redo Strategies.
Acta Crystallogr.,Sect.D, 70, 2014
|
|
1S3O
 
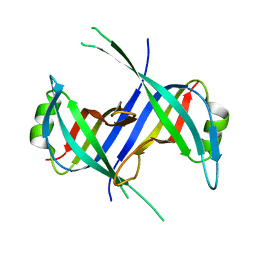 | |
5R8D
 
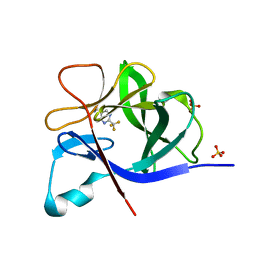 | |
5R87
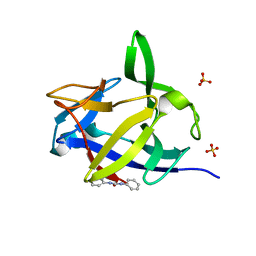 
 | |
5R8L
 
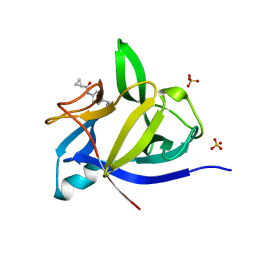 | |
5R8I
 
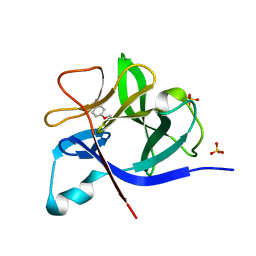 | |
5R8B
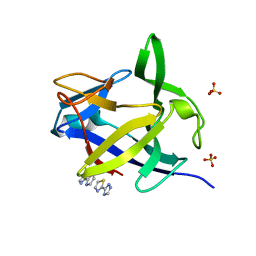 
 | | PanDDA analysis group deposition INTERLEUKIN-1 BETA -- Fragment Z2027049478 in complex with INTERLEUKIN-1 BETA | | Descriptor: | 5-(1,3-thiazol-2-yl)-1H-1,2,4-triazole, Interleukin-1 beta, SULFATE ION | | Authors: | De Nicola, G.F, Nichols, C.E. | | Deposit date: | 2020-03-03 | | Release date: | 2020-04-22 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.49 Å) | | Cite: | Mining the PDB for Tractable Cases Where X-ray Crystallography Combined with Fragment Screens Can Be Used to Systematically Design Protein-Protein Inhibitors: Two Test Cases Illustrated by IL1 beta-IL1R and p38 alpha-TAB1 Complexes.
J.Med.Chem., 63, 2020
|
|
5R88
 
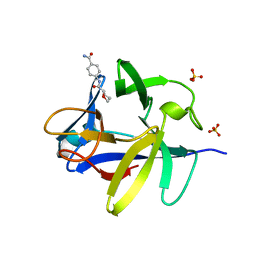 | |
5R8P
 
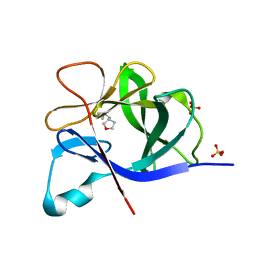 | |
5R8J
 
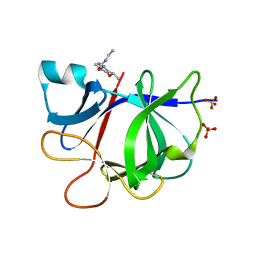 | |
5R8F
 
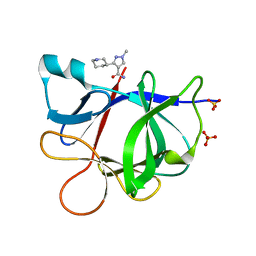 | |
5R8A
 
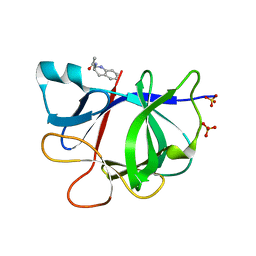 | | PanDDA analysis group deposition INTERLEUKIN-1 BETA -- Fragment Z1492796719 in complex with INTERLEUKIN-1 BETA | | Descriptor: | Interleukin-1 beta, SULFATE ION, ~{N}-[(3~{R})-1,2,3,4-tetrahydroquinolin-3-yl]ethanamide | | Authors: | De Nicola, G.F, Nichols, C.E. | | Deposit date: | 2020-03-03 | | Release date: | 2020-04-22 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.47 Å) | | Cite: | Mining the PDB for Tractable Cases Where X-ray Crystallography Combined with Fragment Screens Can Be Used to Systematically Design Protein-Protein Inhibitors: Two Test Cases Illustrated by IL1 beta-IL1R and p38 alpha-TAB1 Complexes.
J.Med.Chem., 63, 2020
|
|
5R8O
 
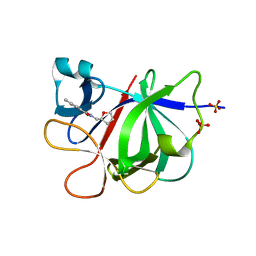 | |
5R8M
 
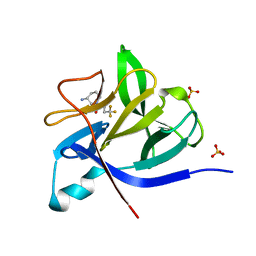 | |
5R89
 
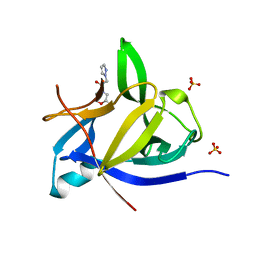 | |
5R8N
 
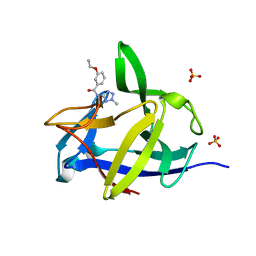 | | PanDDA analysis group deposition INTERLEUKIN-1 BETA -- Fragment Z57292400 in complex with INTERLEUKIN-1 BETA | | Descriptor: | 3-ethoxy-~{N}-(2-methyl-1,2,3,4-tetrazol-5-yl)benzamide, Interleukin-1 beta, SULFATE ION | | Authors: | De Nicola, G.F, Nichols, C.E. | | Deposit date: | 2020-03-03 | | Release date: | 2020-04-22 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.48 Å) | | Cite: | Mining the PDB for Tractable Cases Where X-ray Crystallography Combined with Fragment Screens Can Be Used to Systematically Design Protein-Protein Inhibitors: Two Test Cases Illustrated by IL1 beta-IL1R and p38 alpha-TAB1 Complexes.
J.Med.Chem., 63, 2020
|
|
5R86
 
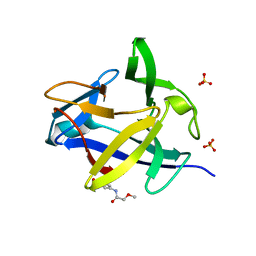 | |
5R8H
 
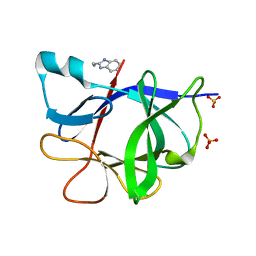 | |
5R8K
 
 | |
