2IIJ
 
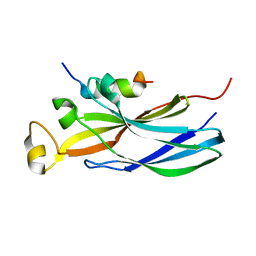 | | Structure of human Asf1a in complex with histone H3 | | Descriptor: | ASF1A protein, Histone H3 | | Authors: | Agez, M, Guerois, R, van Heijenoort, C, Mann, C, Ochsenbein, F. | | Deposit date: | 2006-09-28 | | Release date: | 2007-02-13 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Structure of the histone chaperone asf1 bound to the histone h3 C-terminal helix and functional insights.
Structure, 15, 2007
|
|
2IH6
 
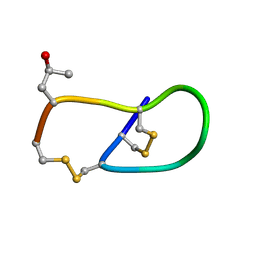 | | Pro6 variant of CMrVIA conotoxin | | Descriptor: | Lambda-conotoxin CMrVIA | | Authors: | Kini, R.M, Kang, T.S. | | Deposit date: | 2006-09-26 | | Release date: | 2007-08-14 | | Last modified: | 2020-06-24 | | Method: | SOLUTION NMR | | Cite: | Protein folding determinants: structural features determining alternative disulfide pairing in alpha- and chi/lambda-conotoxins
Biochemistry, 46, 2007
|
|
2IHA
 
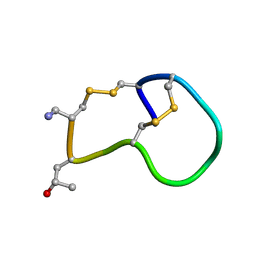 | | Amidated variant of CMrVIA conotoxin | | Descriptor: | Lambda-conotoxin CMrVIA | | Authors: | Kini, R.M, Kang, T.S. | | Deposit date: | 2006-09-26 | | Release date: | 2007-08-14 | | Last modified: | 2020-06-24 | | Method: | SOLUTION NMR | | Cite: | Protein folding determinants: structural features determining alternative disulfide pairing in alpha- and chi/lambda-conotoxins
Biochemistry, 46, 2007
|
|
2IFI
 
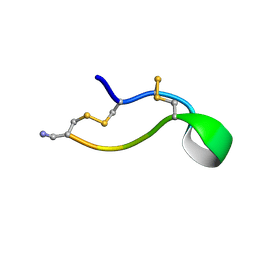 | | Ala6 Variant of ImI Conotoxin | | Descriptor: | Alpha-conotoxin ImI | | Authors: | Kini, R.M, Kang, T.S. | | Deposit date: | 2006-09-21 | | Release date: | 2007-08-14 | | Last modified: | 2020-06-24 | | Method: | SOLUTION NMR | | Cite: | Protein folding determinants: structural features determining alternative disulfide pairing in alpha- and chi/lambda-conotoxins
Biochemistry, 46, 2007
|
|
2IFZ
 
 | | Lys6 Variant of ImI Conotoxin | | Descriptor: | Alpha-conotoxin ImI | | Authors: | Kini, R.M, Kang, T.S. | | Deposit date: | 2006-09-22 | | Release date: | 2007-08-14 | | Last modified: | 2020-06-24 | | Method: | SOLUTION NMR | | Cite: | Protein folding determinants: structural features determining alternative disulfide pairing in alpha- and chi/lambda-conotoxins
Biochemistry, 46, 2007
|
|
2IT8
 | |
2I9O
 
 | |
2IGU
 
 | | Deamidated analogue of ImI Conotoxin | | Descriptor: | Alpha-conotoxin ImI | | Authors: | Kini, R.M, Kang, T.S. | | Deposit date: | 2006-09-25 | | Release date: | 2007-08-14 | | Last modified: | 2022-03-09 | | Method: | SOLUTION NMR | | Cite: | Protein folding determinants: structural features determining alternative disulfide pairing in alpha- and chi/lambda-conotoxins
Biochemistry, 46, 2007
|
|
2IH7
 
 | | Amidated Pro6 Analogue of CMrVIA conotoxin | | Descriptor: | Lambda-conotoxin CMrVIA | | Authors: | Kini, R.M, Kang, T.S. | | Deposit date: | 2006-09-26 | | Release date: | 2007-08-14 | | Last modified: | 2020-06-24 | | Method: | SOLUTION NMR | | Cite: | Protein folding determinants: structural features determining alternative disulfide pairing in alpha- and chi/lambda-conotoxins
Biochemistry, 46, 2007
|
|
2JX2
 
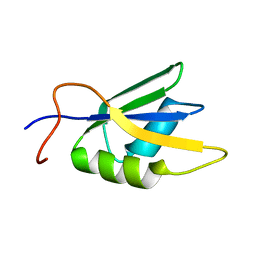 | | Solution conformation of RNA-bound NELF-E RRM | | Descriptor: | Negative elongation factor E | | Authors: | Jampani, N, Schweimer, K, Wenzel, S, Woehrl, B.M, Roesch, P. | | Deposit date: | 2007-11-02 | | Release date: | 2008-04-08 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | NELF-E RRM Undergoes Major Structural Changes in Flexible Protein Regions on Target RNA Binding
Biochemistry, 47, 2008
|
|
2K6A
 
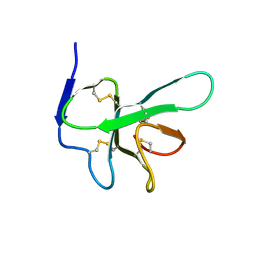 | | Solution structure of EAS D15 truncation mutant | | Descriptor: | Hydrophobin | | Authors: | Kwan, A.H. | | Deposit date: | 2008-07-07 | | Release date: | 2008-08-19 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | The Cys3-Cys4 loop of the hydrophobin EAS is not required for rodlet formation and surface activity.
J.Mol.Biol., 382, 2008
|
|
2K95
 
 | | Solution structure of the wild-type P2B-P3 pseudoknot of human telomerase RNA | | Descriptor: | Telomerase RNA P2b-P3 pseudoknot | | Authors: | Kim, N.-K, Zhang, Q, Zhou, J, Theimer, C.A, Peterson, R.D, Feigon, J. | | Deposit date: | 2008-09-29 | | Release date: | 2008-11-25 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution Structure and Dynamics of the Wild-type Pseudoknot of Human Telomerase RNA.
J.Mol.Biol., 384, 2008
|
|
2I9M
 
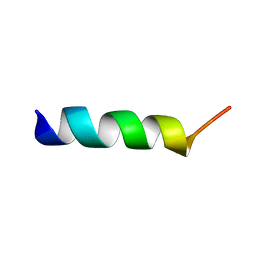 | |
2HI3
 
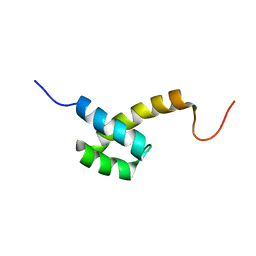 | | Solution structure of the homeodomain-only protein HOP | | Descriptor: | Homeodomain-only protein | | Authors: | Mackay, J.P, Kook, H, Epstein, J.A, Simpson, R.J, Yung, W.W. | | Deposit date: | 2006-06-28 | | Release date: | 2007-01-02 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Analysis of the structure and function of the transcriptional coregulator HOP
Biochemistry, 45, 2006
|
|
2KJG
 
 | |
2KUK
 
 | |
2KN7
 
 | | Structure of the XPF-single strand DNA complex | | Descriptor: | DNA (5'-D(*CP*AP*GP*TP*GP*GP*CP*TP*GP*A)-3'), DNA repair endonuclease XPF | | Authors: | Das, D, Folkers, G.E, van Dijk, M, Jaspers, N.G.J, Hoeijmakers, J.H.J, Kaptein, R, Boelens, R. | | Deposit date: | 2009-08-16 | | Release date: | 2010-08-04 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | The structure of the XPF-ssDNA complex underscores the distinct roles of the XPF and ERCC1 helix- hairpin-helix domains in ss/ds DNA recognition
Structure, 20, 2012
|
|
2KTD
 
 | | Solution structure of mouse lipocalin-type prostaglandin D synthase / substrate analog (U-46619) complex | | Descriptor: | (5Z)-7-{(1R,4S,5S,6R)-6-[(1E,3S)-3-hydroxyoct-1-en-1-yl]-2-oxabicyclo[2.2.1]hept-5-yl}hept-5-enoic acid, Prostaglandin-H2 D-isomerase | | Authors: | Shimamoto, S, Maruo, H, Yoshida, T, Kato, N, Ohkubo, T. | | Deposit date: | 2010-01-27 | | Release date: | 2011-02-02 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of Lipocalin-type Prostaglandin D synthase / Substrate analog complex reveals Open-Closed Conformational Change required for Substrate Recognition
To be Published
|
|
2L7H
 
 | |
2LL4
 
 | |
2L6S
 | | Efficacy of an HIV-1 entry inhibitor targeting the GP41 fusion peptide | | Descriptor: | VIR-576 | | Authors: | Forssmann, W, The, Y, Stoll, M, Adermann, K, Albrecht, U, Barlos, K, Busmann, A, Canales-Mayordomo, A, Gimenez-Gallego, G, Hirsch, J, Jimenez-Barbero, J, Meyer-Olson, D, Muench, J, Perez-Castells, J, Standker, L, Kirchhoff, F, Schmidt, R.E. | | Deposit date: | 2010-11-24 | | Release date: | 2011-01-19 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Short-term monotherapy in HIV-infected patients with a virus entry inhibitor against the gp41 fusion peptide.
Sci Transl Med, 2, 2010
|
|
2L1V
 
 | |
2LCL
 
 | |
2IB1
 
 | |
2I9N
 
 | |
