1FEC
 | |
1FEE
 
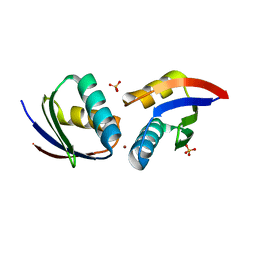 | | CRYSTAL STRUCTURE OF COPPER-HAH1 | | Descriptor: | COPPER (I) ION, COPPER TRANSPORT PROTEIN ATOX1, SULFATE ION, ... | | Authors: | Wernimont, A.K, Huffman, D.L, Lamb, A.L, O'Halloran, T.V, Rosenzweig, A.C. | | Deposit date: | 2000-07-21 | | Release date: | 2001-01-21 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural basis for copper transfer by the metallochaperone for the Menkes/Wilson disease proteins.
Nat.Struct.Biol., 7, 2000
|
|
1FEH
 
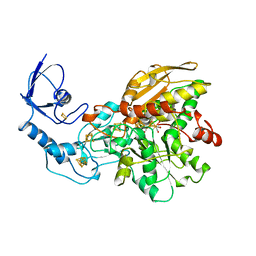 | | FE-ONLY HYDROGENASE FROM CLOSTRIDIUM PASTEURIANUM | | Descriptor: | 2 IRON/2 SULFUR/5 CARBONYL/2 WATER INORGANIC CLUSTER, FE2/S2 (INORGANIC) CLUSTER, IRON/SULFUR CLUSTER, ... | | Authors: | Peters, J.W, Lanzilotta, W.N, Lemon, B.J, Seefeldt, L.C. | | Deposit date: | 1998-10-28 | | Release date: | 1999-01-13 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | X-ray crystal structure of the Fe-only hydrogenase (CpI) from Clostridium pasteurianum to 1.8 angstrom resolution.
Science, 282, 1998
|
|
1FEJ
 
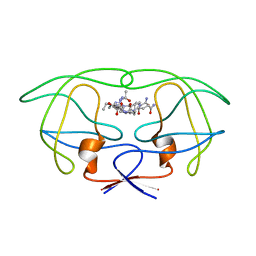 | | STRUCTURAL IMPLICATIONS OF DRUG RESISTANT MUTANTS OF HIV-1 PROTEASE: HIGH RESOLUTION CRYSTAL STRUCTURES OF THE MUTANT PROTEASE/SUBSTRATE ANALOG COMPLEXES | | Descriptor: | N-{(2S)-2-[(N-acetyl-L-threonyl-L-isoleucyl)amino]hexyl}-L-norleucyl-L-glutaminyl-N~5~-[amino(iminio)methyl]-L-ornithinamide, PROTEASE RETROPEPSIN | | Authors: | Mahalingam, B, Louis, J.M, Harrison, R.W, Weber, I.T. | | Deposit date: | 2000-07-21 | | Release date: | 2001-06-01 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.78 Å) | | Cite: | Structural implications of drug-resistant mutants of HIV-1 protease: high-resolution crystal structures of the mutant protease/substrate analogue complexes.
Proteins, 43, 2001
|
|
1FEL
 | | CRYSTALLOGRAPHIC STUDIES ON COMPLEXES BETWEEN RETINOIDS AND PLASMA RETINOL-BINDING PROTEIN | | Descriptor: | N-(4-HYDROXYPHENYL)ALL-TRANS RETINAMIDE, RETINOL BINDING PROTEIN | | Authors: | Zanotti, G, Marcello, M, Malpeli, G, Sartori, G, Berni, R. | | Deposit date: | 1994-08-29 | | Release date: | 1994-11-01 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystallographic studies on complexes between retinoids and plasma retinol-binding protein.
J.Biol.Chem., 269, 1994
|
|
1FEM
 
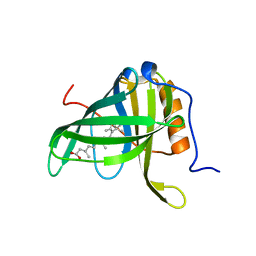 | | CRYSTALLOGRAPHIC STUDIES ON COMPLEXES BETWEEN RETINOIDS AND PLASMA RETINOL-BINDING PROTEIN | | Descriptor: | RETINOIC ACID, RETINOL BINDING PROTEIN | | Authors: | Zanotti, G, Marcello, M, Malpeli, G, Sartori, G, Berni, R. | | Deposit date: | 1994-08-29 | | Release date: | 1994-11-01 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystallographic studies on complexes between retinoids and plasma retinol-binding protein.
J.Biol.Chem., 269, 1994
|
|
1FEN
 
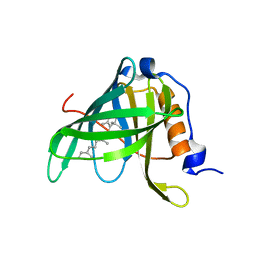 | | CRYSTALLOGRAPHIC STUDIES ON COMPLEXES BETWEEN RETINOIDS AND PLASMA RETINOL-BINDING PROTEIN | | Descriptor: | ALL-TRANS AXEROPHTHENE, RETINOL BINDING PROTEIN | | Authors: | Zanotti, G, Marcello, M, Malpeli, G, Sartori, G, Berni, R. | | Deposit date: | 1994-08-29 | | Release date: | 1994-11-01 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystallographic studies on complexes between retinoids and plasma retinol-binding protein.
J.Biol.Chem., 269, 1994
|
|
1FEO
 | |
1FEP
 
 | | FERRIC ENTEROBACTIN RECEPTOR | | Descriptor: | FERRIC ENTEROBACTIN RECEPTOR | | Authors: | Buchanan, S.K, Smith, B.S, Ventatramani, L, Xia, D, Esser, L, Palnitkar, M, Chakraborty, R, Van Der Helm, D, Deisenhofer, J. | | Deposit date: | 1998-11-24 | | Release date: | 1999-01-13 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of the outer membrane active transporter FepA from Escherichia coli.
Nat.Struct.Biol., 6, 1999
|
|
1FEQ
 
 | | NMR SOLUTION STRUCTURE OF THE ANTICODON OF TRNA(LYS3) WITH T6A MODIFICATION AT POSITION 37 | | Descriptor: | 5'-R(*GP*CP*AP*GP*AP*CP*UP*UP*UP*UP*(T6A)P*AP*UP*CP*UP*GP*C)-3' | | Authors: | Stuart, J.W, Gdaniec, Z, Guenther, R.H, Marszalek, M, Sochacka, E, Malkiewicz, A, Agris, P.F. | | Deposit date: | 2000-07-21 | | Release date: | 2000-11-22 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Functional anticodon architecture of human tRNALys3 includes disruption of intraloop hydrogen bonding by the naturally occurring amino acid modification, t6A.
Biochemistry, 39, 2000
|
|
1FER
 
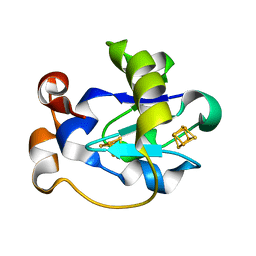 | | STRUCTURE AT PH 6.5 OF FERREDOXIN I FROM AZOTOBACTER VINELANDII AT 2.3 ANGSTROMS RESOLUTION | | Descriptor: | FE3-S4 CLUSTER, FERREDOXIN I, IRON/SULFUR CLUSTER | | Authors: | Merritt, E.A, Stout, G.H, Turley, S, Sieker, L.C, Jensen, L.H, Orme-Johnson, W.H. | | Deposit date: | 1992-09-02 | | Release date: | 1993-04-15 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure at pH 6.5 of ferredoxin I from Azotobacter vinelandii at 2.3 A resolution.
Acta Crystallogr.,Sect.D, 49, 1993
|
|
1FES
 
 | | SOLUTION STRUCTURE OF THE APO FORM OF THE YEAST METALLOCHAPERONE, ATX1 | | Descriptor: | ATX1 COPPER CHAPERONE | | Authors: | Arnesano, F, Banci, L, Bertini, I, Huffman, D.L, O'Halloran, T.V. | | Deposit date: | 2000-07-22 | | Release date: | 2001-03-14 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the Cu(I) and apo forms of the yeast metallochaperone, Atx1.
Biochemistry, 40, 2001
|
|
1FEU
 
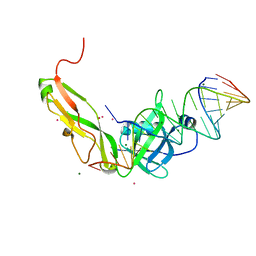 | | CRYSTAL STRUCTURE OF RIBOSOMAL PROTEIN TL5, ONE OF THE CTC FAMILY PROTEINS, COMPLEXED WITH A FRAGMENT OF 5S RRNA. | | Descriptor: | 19 NT FRAGMENT OF 5S RRNA, 21 NT FRAGMENT OF 5S RRNA, 50S RIBOSOMAL PROTEIN L25, ... | | Authors: | Fedorov, R.V, Meshcheryakov, V.A, Gongadze, G.M, Fomenkova, N.P, Nevskaya, N.A, Selmer, M, Laurberg, M, Kristensen, O, Al-Karadaghi, S, Liljas, A, Garber, M.B, Nikonov, S.V. | | Deposit date: | 2000-07-23 | | Release date: | 2001-06-25 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure of ribosomal protein TL5 complexed with RNA provides new insights into the CTC family of stress proteins.
Acta Crystallogr.,Sect.D, 57, 2001
|
|
1FEV
 
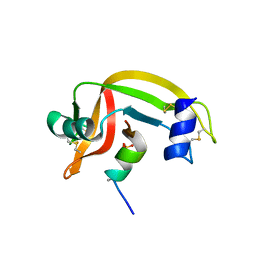 | |
1FEW
 
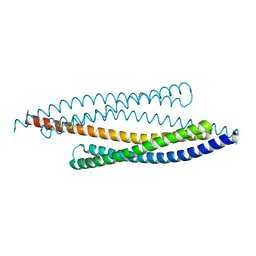 | | CRYSTAL STRUCTURE OF SMAC/DIABLO | | Descriptor: | SECOND MITOCHONDRIA-DERIVED ACTIVATOR OF CASPASES | | Authors: | Chai, J, Shi, Y. | | Deposit date: | 2000-07-23 | | Release date: | 2000-09-13 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural and biochemical basis of apoptotic activation by Smac/DIABLO.
Nature, 406, 2000
|
|
1FEX
 
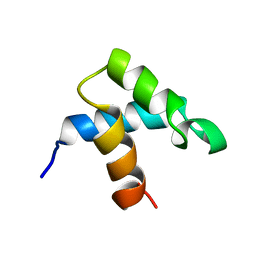 | |
1FEZ
 
 | | THE CRYSTAL STRUCTURE OF BACILLUS CEREUS PHOSPHONOACETALDEHYDE HYDROLASE COMPLEXED WITH TUNGSTATE, A PRODUCT ANALOG | | Descriptor: | MAGNESIUM ION, PHOSPHONOACETALDEHYDE HYDROLASE, TUNGSTATE(VI)ION | | Authors: | Morais, M.C, Zhang, W, Baker, A.S, Zhang, G, Dunaway-Mariano, D, Allen, K.N. | | Deposit date: | 2000-07-24 | | Release date: | 2000-10-04 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | The crystal structure of bacillus cereus phosphonoacetaldehyde hydrolase: insight into catalysis of phosphorus bond cleavage and catalytic diversification within the HAD enzyme superfamily.
Biochemistry, 39, 2000
|
|
1FF0
 
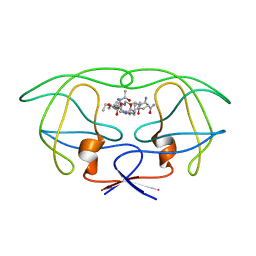 | | STRUCTURAL IMPLICATIONS OF DRUG RESISTANT MUTANTS OF HIV-1 PROTEASE: HIGH RESOLUTION CRYSTAL STRUCTURES OF THE MUTANT PROTEASE/SUBSTRATE ANALOG COMPLEXES. | | Descriptor: | N-{(2S)-2-[(N-acetyl-L-threonyl-L-isoleucyl)amino]hexyl}-L-norleucyl-L-glutaminyl-N~5~-[amino(iminio)methyl]-L-ornithinamide, PROTEASE RETROPEPSIN | | Authors: | Mahalingam, B, Louis, J.M, Harrison, R.W, Weber, I.T. | | Deposit date: | 2000-07-24 | | Release date: | 2001-06-01 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Structural implications of drug-resistant mutants of HIV-1 protease: high-resolution crystal structures of the mutant protease/substrate analogue complexes.
Proteins, 43, 2001
|
|
1FF1
 
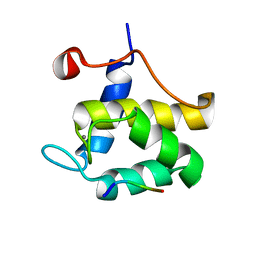 | | STRUCTURE OF THE SECOND EPS15 HOMOLOGY DOMAIN OF HUMAN EPS15 IN COMPLEX WITH PTGSSSTNPFL | | Descriptor: | CALCIUM ION, EPIDERMAL GROWTH FACTOR RECEPTOR SUBSTRATE 15, PTGSSSTNPFL PEPTIDE | | Authors: | De Beer, T, Hoofnagle, A.N, Enmon, J.L, Bowers, R.C, Yamabhai, M, Kay, B.K, Overduin, M. | | Deposit date: | 2000-07-24 | | Release date: | 2000-11-01 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Molecular mechanism of NPF recognition by EH domains.
Nat.Struct.Biol., 7, 2000
|
|
1FF2
 
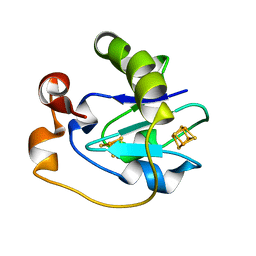 | |
1FF3
 
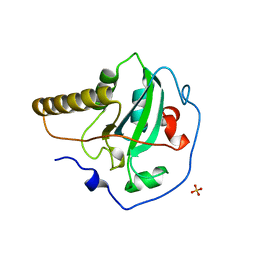 | | STRUCTURE OF THE PEPTIDE METHIONINE SULFOXIDE REDUCTASE FROM ESCHERICHIA COLI | | Descriptor: | PEPTIDE METHIONINE SULFOXIDE REDUCTASE, SULFATE ION | | Authors: | Tete-Favier, F, Cobessi, D, Boschi-Muller, S, Azza, S, Branlant, G, Aubry, A. | | Deposit date: | 2000-07-25 | | Release date: | 2000-12-06 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of the Escherichia coli peptide methionine sulphoxide reductase at 1.9 A resolution.
Structure Fold.Des., 8, 2000
|
|
1FF4
 
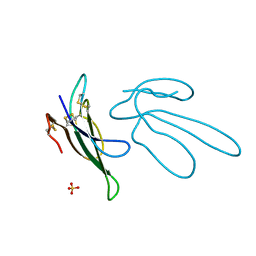 | |
1FF5
 | | STRUCTURE OF E-CADHERIN DOUBLE DOMAIN | | Descriptor: | CALCIUM ION, EPITHELIAL CADHERIN | | Authors: | Pertz, O, Bozic, D, Koch, A.W, Fauser, C, Brancaccio, A, Engel, J. | | Deposit date: | 2000-07-25 | | Release date: | 2000-08-23 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.93 Å) | | Cite: | A new crystal structure, Ca2+ dependence and mutational analysis reveal molecular details of E-cadherin homoassociation.
EMBO J., 18, 1999
|
|
1FF7
 
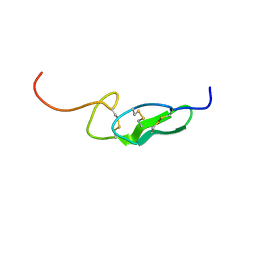 | | THE FIRST EGF-LIKE DOMAIN FROM HUMAN BLOOD COAGULATION FVII (FUCOSYLATED AT SER-60), NMR, 20 STRUCTURES | | Descriptor: | PROTEIN (Blood Coagulation Factor VII), alpha-L-fucopyranose | | Authors: | Kao, Y.-H, Lee, G.F, Wang, Y, Starovasnik, M.A, Kelley, R.F, Spellman, M.W, Lerner, L. | | Deposit date: | 1999-02-19 | | Release date: | 1999-06-16 | | Last modified: | 2024-10-16 | | Method: | SOLUTION NMR | | Cite: | The effect of O-fucosylation on the first EGF-like domain from human blood coagulation factor VII.
Biochemistry, 38, 1999
|
|
1FF9
 
 | | APO SACCHAROPINE REDUCTASE | | Descriptor: | SACCHAROPINE REDUCTASE, SULFATE ION | | Authors: | Johansson, E, Steffens, J.J, Lindqvist, Y, Schneider, G. | | Deposit date: | 2000-07-25 | | Release date: | 2000-11-08 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of saccharopine reductase from Magnaporthe grisea, an enzyme of the alpha-aminoadipate pathway of lysine biosynthesis.
Structure Fold.Des., 8, 2000
|
|
