5R92
 
 | |
5R9I
 
 | |
5R9W
 
 | |
5R90
 
 | |
5R9J
 
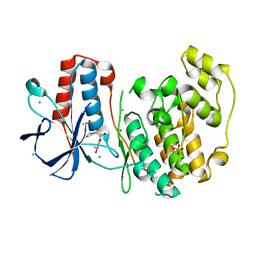 | | PanDDA analysis group deposition Form1 MAP kinase p38-alpha -- Fragment N14231a in complex with MAP kinase p38-alpha | | Descriptor: | 3-ethyl-1-[(1~{R},8~{S},9~{S},10~{S})-10-oxidanyl-11-oxatricyclo[6.2.1.0^{2,7}]undeca-2(7),3,5-trien-9-yl]imidazolidine-2,4-dione, CHLORIDE ION, DIMETHYL SULFOXIDE, ... | | Authors: | De Nicola, G.F, Nichols, C.E. | | Deposit date: | 2020-03-04 | | Release date: | 2020-07-22 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.52 Å) | | Cite: | Mining the PDB for Tractable Cases Where X-ray Crystallography Combined with Fragment Screens Can Be Used to Systematically Design Protein-Protein Inhibitors: Two Test Cases Illustrated by IL1 beta-IL1R and p38 alpha-TAB1 Complexes.
J.Med.Chem., 63, 2020
|
|
5RA4
 
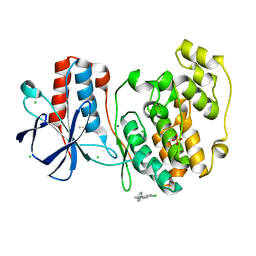 | |
5R85
 
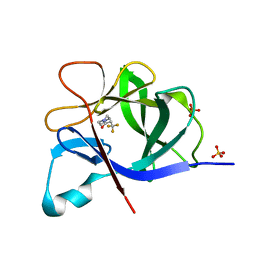 | |
5R8G
 
 | |
5R8X
 
 | |
5R9D
 
 | |
5R9T
 
 | | PanDDA analysis group deposition Form1 MAP kinase p38-alpha -- Fragment N13477a in complex with MAP kinase p38-alpha | | Descriptor: | 6-methyl-5-[(4-propan-2-ylphenyl)amino]-2~{H}-1,2,4-triazin-3-one, CHLORIDE ION, MAGNESIUM ION, ... | | Authors: | De Nicola, G.F, Nichols, C.E. | | Deposit date: | 2020-03-04 | | Release date: | 2020-07-22 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Mining the PDB for Tractable Cases Where X-ray Crystallography Combined with Fragment Screens Can Be Used to Systematically Design Protein-Protein Inhibitors: Two Test Cases Illustrated by IL1 beta-IL1R and p38 alpha-TAB1 Complexes.
J.Med.Chem., 63, 2020
|
|
5RA9
 
 | |
5R8E
 
 | | PanDDA analysis group deposition INTERLEUKIN-1 BETA -- Fragment Z57475877 in complex with INTERLEUKIN-1 BETA | | Descriptor: | Interleukin-1 beta, SULFATE ION, ~{N}-(2-ethyl-1,2,3,4-tetrazol-5-yl)butanamide | | Authors: | De Nicola, G.F, Nichols, C.E. | | Deposit date: | 2020-03-03 | | Release date: | 2020-04-22 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Mining the PDB for Tractable Cases Where X-ray Crystallography Combined with Fragment Screens Can Be Used to Systematically Design Protein-Protein Inhibitors: Two Test Cases Illustrated by IL1 beta-IL1R and p38 alpha-TAB1 Complexes.
J.Med.Chem., 63, 2020
|
|
5R8Q
 
 | |
5R98
 
 | |
5R9N
 
 | |
5R9Z
 
 | |
5R8C
 
 | |
5R8U
 
 | |
5R9B
 
 | |
5R9S
 
 | |
5R8I
 
 | |
5R91
 
 | |
5R9M
 
 | |
5RA1
 
 | |
