3KI0
 
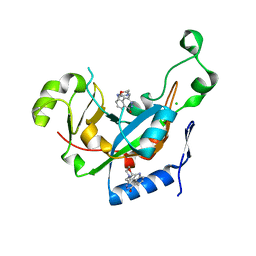 | | Catalytic fragment of Cholix toxin from Vibrio Cholerae in complex with inhibitor GP-D | | Descriptor: | 3-(morpholin-4-ylmethyl)-1,5-dihydro-6H-[1,2]diazepino[4,5,6-cd]indol-6-one, CHLORIDE ION, Cholix toxin | | Authors: | Jorgensen, R, Edwards, P.R, Merrill, A.R. | | Deposit date: | 2009-10-31 | | Release date: | 2010-10-27 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.29 Å) | | Cite: | Structure function analysis of soluble inhibitors of cholix toxin from Vibrio cholerae
To be Published
|
|
3KI7
 
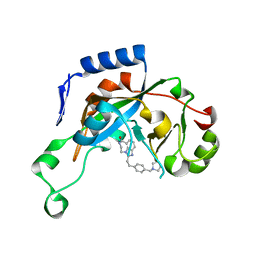 | | Catalytic fragment of Cholix toxin from Vibrio Cholerae in complex with inhibitor GP-I | | Descriptor: | 2-{2-[4-(pyrrolidin-1-ylmethyl)phenyl]ethyl}-5,6-dihydroimidazo[4,5,1-jk][1,4]benzodiazepin-7(4H)-one, Cholix toxin | | Authors: | Jorgensen, R, Edwards, P.R, Merrill, A.R. | | Deposit date: | 2009-10-31 | | Release date: | 2010-10-27 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.32 Å) | | Cite: | Structure function analysis of soluble inhibitors of cholix toxin from Vibrio cholerae
To be Published
|
|
3KI2
 
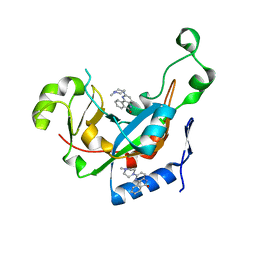 | |
3KI4
 
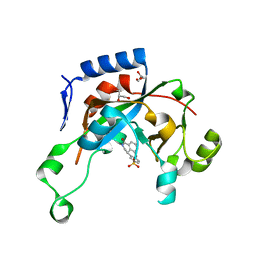 | | Catalytic fragment of Cholix toxin from Vibrio Cholerae in complex with inhibitor GP-P | | Descriptor: | (11bR)-3-oxo-1,2,3,11b-tetrahydrochromeno[4,3,2-de]isoquinoline-10-sulfonic acid, Cholix toxin, GLYCEROL | | Authors: | Jorgensen, R, Edwards, P.R, Merrill, A.R. | | Deposit date: | 2009-10-31 | | Release date: | 2010-10-27 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Structure function analysis of soluble inhibitors of cholix toxin from Vibrio cholerae
To be Published
|
|
3KI6
 
 | | Catalytic fragment of Cholix toxin from Vibrio Cholerae in complex with inhibitor GP-L | | Descriptor: | 8-fluoro-2-(3-piperidin-1-ylpropanoyl)-1,3,4,5-tetrahydrobenzo[c][1,6]naphthyridin-6(2H)-one, Cholix toxin | | Authors: | Jorgensen, R, Edwards, P.R, Merrill, A.R. | | Deposit date: | 2009-10-31 | | Release date: | 2010-10-27 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.54 Å) | | Cite: | Structure function analysis of soluble inhibitors of cholix toxin from Vibrio cholerae
To be Published
|
|
3KI3
 
 | | Catalytic fragment of Cholix toxin from Vibrio Cholerae in complex with inhibitor GP-H | | Descriptor: | 2-[(dimethylamino)methyl]-5,6-dihydroimidazo[4,5,1-jk][1,4]benzodiazepin-7(4H)-one, Cholix toxin | | Authors: | Jorgensen, R, Edwards, P.R, Merrill, A.R. | | Deposit date: | 2009-10-31 | | Release date: | 2010-10-27 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.27 Å) | | Cite: | Structure function analysis of soluble inhibitors of cholix toxin from Vibrio cholerae
To be Published
|
|
3KI5
 
 | |
3KI1
 
 | | Catalytic fragment of Cholix toxin from Vibrio Cholerae in complex with inhibitor GP-F | | Descriptor: | 2-{4-[3-(dimethylamino)propoxy]phenyl}-5,6-dihydroimidazo[4,5,1-jk][1,4]benzodiazepin-7(4H)-one, Cholix toxin | | Authors: | Jorgensen, R, Edwards, P.R, Merrill, A.R. | | Deposit date: | 2009-10-31 | | Release date: | 2010-10-27 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.43 Å) | | Cite: | Structure function analysis of soluble inhibitors of cholix toxin from Vibrio cholerae
To be Published
|
|
7N6M
 
 | | Crystal structure of the substrate-binding domain of E. coli DnaK in complex with the peptide RQKPLLGLSR | | Descriptor: | Alkaline phosphatase peptide, Chaperone protein DnaK, SULFATE ION | | Authors: | Jansen, R.M, Ozden, C, Gierasch, L.M, Garman, S.C. | | Deposit date: | 2021-06-08 | | Release date: | 2021-06-23 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.82 Å) | | Cite: | Selective promiscuity in the binding of E. coli Hsp70 to an unfolded protein.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
7N6K
 
 | | Crystal structure of the substrate-binding domain of E. coli DnaK in complex with the peptide RALALLPLSR | | Descriptor: | Alkaline phosphatase peptide, Chaperone protein DnaK, SULFATE ION | | Authors: | Jansen, R.M, Ozden, C, Gierasch, L.M, Garman, S.C. | | Deposit date: | 2021-06-08 | | Release date: | 2021-06-23 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Selective promiscuity in the binding of E. coli Hsp70 to an unfolded protein.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
7N6J
 
 | | Crystal structure of the substrate-binding domain of E. coli DnaK in complex with the peptide RKQSTIALALLPLLFTPRR | | Descriptor: | Alkaline phosphatase peptide, Chaperone protein DnaK, SULFATE ION | | Authors: | Jansen, R.M, Ozden, C, Gierasch, L.M, Garman, S.C. | | Deposit date: | 2021-06-08 | | Release date: | 2021-06-23 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Selective promiscuity in the binding of E. coli Hsp70 to an unfolded protein.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
7N6L
 
 | | Crystal structure of the substrate-binding domain of E. coli DnaK in complex with the peptide EANQQKPLLGLFADG | | Descriptor: | Alkaline phosphatase peptide, Chaperone protein DnaK, GLYCEROL, ... | | Authors: | Jansen, R.M, Ozden, C, Gierasch, L.M, Garman, S.C. | | Deposit date: | 2021-06-08 | | Release date: | 2021-06-23 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Selective promiscuity in the binding of E. coli Hsp70 to an unfolded protein.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
5JWQ
 
 | | Crystal structure of KaiC S431E in complex with foldswitch-stabilized KaiB from Thermosynechococcus elongatus | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, Circadian clock protein KaiB, Circadian clock protein kinase KaiC | | Authors: | Tseng, R, Goularte, N.F, Chavan, A, Luu, J, Chang, Y, Heilser, J, Tripathi, S, LiWang, A, Partch, C.L. | | Deposit date: | 2016-05-12 | | Release date: | 2017-03-29 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (3.871 Å) | | Cite: | Structural basis of the day-night transition in a bacterial circadian clock.
Science, 355, 2017
|
|
3LE4
 
 | |
7JNE
 
 | | Crystal structure of the substrate-binding domain of E. coli DnaK in complex with the peptide RGSQLRIASR | | Descriptor: | Alkaline phosphatase peptide, Chaperone protein DnaK, SULFATE ION | | Authors: | Jansen, R.M, Ozden, C, Gierasch, L.M, Garman, S.C. | | Deposit date: | 2020-08-04 | | Release date: | 2020-08-26 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.54 Å) | | Cite: | Selective promiscuity in the binding of E. coli Hsp70 to an unfolded protein.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
7JMM
 
 | | Crystal structure of the substrate-binding domain of E. coli DnaK in complex with the peptide RAKNIILLSR | | Descriptor: | Alkaline phosphatase, Chaperone protein DnaK, SULFATE ION | | Authors: | Jansen, R.M, Ozden, C, Gierasch, L.M, Garman, S.C. | | Deposit date: | 2020-08-02 | | Release date: | 2020-08-26 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.56 Å) | | Cite: | Selective promiscuity in the binding of E. coli Hsp70 to an unfolded protein.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
7JN8
 
 | | Crystal structure of the substrate-binding domain of E. coli DnaK in complex with the peptide RGNTLVIVSR | | Descriptor: | Alkaline phosphatase peptide, Chaperone protein DnaK, SULFATE ION | | Authors: | Jansen, R.M, Ozden, C, Gierasch, L.M, Garman, S.C. | | Deposit date: | 2020-08-04 | | Release date: | 2020-08-26 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (3.09 Å) | | Cite: | Selective promiscuity in the binding of E. coli Hsp70 to an unfolded protein.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
7JN9
 
 | | Crystal structure of the substrate-binding domain of E. coli DnaK in complex with the peptide QEHTGSQLRIAAYGP | | Descriptor: | Alkaline phosphatase peptide, Chaperone protein DnaK, SULFATE ION | | Authors: | Jansen, R.M, Ozden, C, Gierasch, L.M, Garman, S.C. | | Deposit date: | 2020-08-04 | | Release date: | 2020-08-26 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Selective promiscuity in the binding of E. coli Hsp70 to an unfolded protein.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
3NQS
 
 | | Crystal Structure of Inducible Nitric Oxide Synthase with N-Nitrosated-pterin | | Descriptor: | 1,2-ETHANEDIOL, 5,6,7,8-TETRAHYDROBIOPTERIN, ETHYL 4-[(4-METHYLPYRIDIN-2-YL)AMINO]PIPERIDINE-1-CARBOXYLATE, ... | | Authors: | Rosenfeld, R.J, Getzoff, E.D, Tainer, J.A. | | Deposit date: | 2010-06-29 | | Release date: | 2010-07-21 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Nitric-oxide synthase forms N-NO-pterin and S-NO-cys: implications for activity, allostery, and regulation.
J.Biol.Chem., 285, 2010
|
|
1U2R
 
 | | Crystal Structure of ADP-ribosylated Ribosomal Translocase from Saccharomyces cerevisiae | | Descriptor: | ADENOSINE-5-DIPHOSPHORIBOSE, Elongation factor 2, GUANOSINE-5'-DIPHOSPHATE, ... | | Authors: | Jorgensen, R, Yates, S.P, Nilsson, J, Prentice, G.A, Teal, D.J, Merrill, A.R, Andersen, G.R. | | Deposit date: | 2004-07-20 | | Release date: | 2004-09-14 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Crystal Structure of ADP-ribosylated Ribosomal Translocase from Saccharomyces cerevisiae
J.Biol.Chem., 279, 2004
|
|
2WV9
 
 | | Crystal Structure of the NS3 protease-helicase from Murray Valley encephalitis virus | | Descriptor: | FLAVIVIRIN PROTEASE NS2B REGULATORY SUBUNIT, FLAVIVIRIN PROTEASE NS3 CATALYTIC SUBUNIT | | Authors: | Assenberg, R, Mastrangelo, E, Walter, T.S, Verma, A, Milani, M, Owens, R.J, Stuart, D.I, Grimes, J.M, Mancini, E.J. | | Deposit date: | 2009-10-15 | | Release date: | 2009-12-01 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Crystal Structure of a Novel Conformational State of the Flavivirus Ns3 Protein: Implications for Polyprotein Processing and Viral Replication.
J.Virol., 83, 2009
|
|
2WZL
 
 | | The Structure of the N-RNA Binding Domain of the Mokola virus Phosphoprotein | | Descriptor: | GLYCEROL, PHOSPHOPROTEIN | | Authors: | Assenberg, R, Delmas, O, Ren, J, Vidalain, P, Verma, A, Larrous, F, Graham, S, Tangy, F, Grimes, J, Bourhy, H. | | Deposit date: | 2009-11-30 | | Release date: | 2009-12-15 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The Structure of the N-RNA Binding Domain of the Mokola Virus Phosphoprotein
J.Virol., 84, 2010
|
|
1ZM3
 
 | | Structure of the apo eEF2-ETA complex | | Descriptor: | Elongation factor 2, exotoxin A | | Authors: | Joergensen, R, Merrill, A.R, Yates, S.P, Marquez, V.E, Schwan, A.L, Boesen, T, Andersen, G.R. | | Deposit date: | 2005-05-10 | | Release date: | 2005-05-24 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (3.07 Å) | | Cite: | Exotoxin A-eEF2 complex structure indicates ADP ribosylation by ribosome mimicry.
Nature, 436, 2005
|
|
1ZM9
 
 | | Structure of eEF2-ETA in complex with PJ34 | | Descriptor: | Elongation factor 2, N~2~,N~2~-DIMETHYL-N~1~-(6-OXO-5,6-DIHYDROPHENANTHRIDIN-2-YL)GLYCINAMIDE, exotoxin A | | Authors: | Joergensen, R, Merrill, A.R, Yates, S.P, Marquez, V.E, Schwan, A.L, Boesen, T, Andersen, G.R. | | Deposit date: | 2005-05-10 | | Release date: | 2005-05-24 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Exotoxin A-eEF2 complex structure indicates ADP ribosylation by ribosome mimicry.
Nature, 436, 2005
|
|
1ZM4
 
 | | Structure of the eEF2-ETA-bTAD complex | | Descriptor: | BETA-METHYLENE-THIAZOLE-4-CARBOXYAMIDE-ADENINE DINUCLEOTIDE, Elongation factor 2, exotoxin A | | Authors: | Joergensen, R, Merrill, A.R, Yates, S.P, Marquez, V.E, Schwan, A.L, Boesen, T, Andersen, G.R. | | Deposit date: | 2005-05-10 | | Release date: | 2005-05-24 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Exotoxin A-eEF2 complex structure indicates ADP ribosylation by ribosome mimicry.
Nature, 436, 2005
|
|
