2VQ0
 
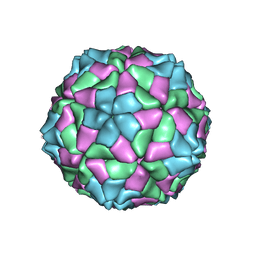 | | Capsid structure of Sesbania mosaic virus coat protein deletion mutant rCP(delta 48 to 59) | | Descriptor: | CALCIUM ION, COAT PROTEIN | | Authors: | Anju, P, Subashchandrabose, C, Satheshkumar, P.S, Savithri, H.S, Murthy, M.R.N. | | Deposit date: | 2008-03-10 | | Release date: | 2008-05-13 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (3.6 Å) | | Cite: | Structure of Recombinant Capsids Formed by the Beta-Annulus Deletion Mutant-Rcp (Delta 48-59) of Sesbania Mosaic Virus
Virology, 375, 2008
|
|
2W7J
 
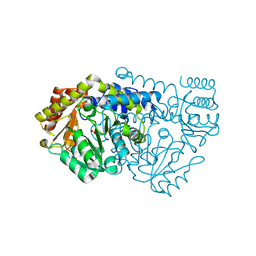 | | Crystal structure of Y61AbsSHMT Glycine external aldimine | | Descriptor: | GLYCINE, PYRIDOXAL-5'-PHOSPHATE, SERINE HYDROXYMETHYLTRANSFERASE | | Authors: | Rajaram, V, Bhavani, B.S, Bisht, S, Kaul, P, Prakash, V, Appaji Rao, N, Savithri, H.S, Murthy, M.R.N. | | Deposit date: | 2008-12-22 | | Release date: | 2010-08-18 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.68 Å) | | Cite: | Importance of Tyrosine Residues of Bacillus Stearothermophilus Serine Hydroxymethyltransferase in Cofactor Binding and L-Allo-Thr Cleavage.
FEBS J., 275, 2008
|
|
2W7D
 
 | | Crystal structure of Y51FbsSHMT internal aldimine | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, PHOSPHATE ION, PYRIDOXAL-5'-PHOSPHATE, ... | | Authors: | Rajaram, V, Bhavani, B.S, Bisht, S, Kaul, P, Prakash, V, Appaji Rao, N, Savithri, H.S, Murthy, M.R.N. | | Deposit date: | 2008-12-22 | | Release date: | 2010-08-18 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Importance of Tyrosine Residues of Bacillus Stearothermophilus Serine Hydroxymethyltransferase in Cofactor Binding and L-Allo-Thr Cleavage.
FEBS J., 275, 2008
|
|
2W7M
 
 | | Crystal structure of Y61AbsSHMT obtained in the presence of Glycine and 5-Formyl tetrahydrofolate | | Descriptor: | GLYCINE, PYRIDOXAL-5'-PHOSPHATE, SERINE HYDROXYMETHYLTRANSFERASE | | Authors: | Rajaram, V, Bhavani, B.S, Bisht, S, Kaul, P, Prakash, V, Appaji Rao, N, Savithri, H.S, Murthy, M.R.N. | | Deposit date: | 2008-12-22 | | Release date: | 2010-08-18 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Importance of Tyrosine Residues of Bacillus Stearothermophilus Serine Hydroxymethyltransferase in Cofactor Binding and L-Allo-Thr Cleavage.
FEBS J., 275, 2008
|
|
2W7F
 
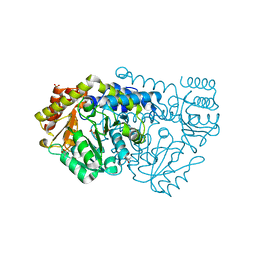 | | Crystal structure of Y51FbsSHMT L-Ser external aldimine | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, PHOSPHATE ION, PYRIDOXAL-5'-PHOSPHATE, ... | | Authors: | Rajaram, V, Bhavani, B.S, Bisht, S, Kaul, P, Prakash, V, Appaji Rao, N, Savithri, H.S, Murthy, M.R.N. | | Deposit date: | 2008-12-22 | | Release date: | 2010-08-18 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.67 Å) | | Cite: | Importance of Tyrosine Residues of Bacillus Stearothermophilus Serine Hydroxymethyltransferase in Cofactor Binding and L-Allo-Thr Cleavage.
FEBS J., 275, 2008
|
|
6LXF
 
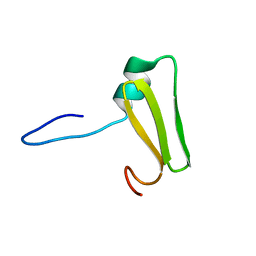 | | Aromatic interactions drive the coupled folding and binding of the intrinsically disordered Sesbania mosaic virus VPg protein. | | Descriptor: | Polyprotein | | Authors: | Dixit, K, Karanth, N.M, Nair, S, Kumari, K, Chakrabarti, K.S, Savithri, H.S, Sarma, S.P. | | Deposit date: | 2020-02-11 | | Release date: | 2021-01-20 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Aromatic Interactions Drive the Coupled Folding and Binding of the Intrinsically Disordered Sesbania mosaic Virus VPg Protein.
Biochemistry, 59, 2020
|
|
6M78
 
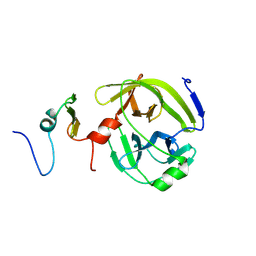 | | Aromatic interactions drive the coupled folding and binding of the intrinsically disordered Sesbania mosaic virus VPg protein | | Descriptor: | Polyprotein | | Authors: | Dixit, K, Karanth, N.M, Nair, S, Kumari, K, Chakraborti, K.S, Savithri, H.S, Sarma, S.P. | | Deposit date: | 2020-03-17 | | Release date: | 2021-01-27 | | Method: | SOLUTION NMR | | Cite: | Aromatic Interactions Drive the Coupled Folding and Binding of the Intrinsically Disordered Sesbania mosaic Virus VPg Protein.
Biochemistry, 59, 2020
|
|
1ZYO
 
 | | Crystal Structure of the Serine Protease Domain of Sesbania Mosaic Virus polyprotein | | Descriptor: | GLYCEROL, serine protease | | Authors: | Gayathri, P, Satheshkumar, P.S, Prasad, K, Savithri, H.S, Murthy, M.R.N. | | Deposit date: | 2005-06-10 | | Release date: | 2006-04-25 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of the serine protease domain of Sesbania mosaic virus polyprotein and mutational analysis of residues forming the S1-binding pocket
Virology, 346, 2006
|
|
3R0X
 
 | | Crystal structure of Selenomethionine incorporated apo D-serine deaminase from Salmonella tyhimurium | | Descriptor: | 1,2-ETHANEDIOL, D-serine dehydratase, SODIUM ION, ... | | Authors: | Bharath, S.R, Shveta, B, Savithri, H.S, Murthy, M.R.N. | | Deposit date: | 2011-03-09 | | Release date: | 2011-06-29 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | Crystal structures of open and closed forms of D-serine deaminase from Salmonella typhimurium - implications on substrate specificity and catalysis
Febs J., 2011
|
|
3R0Z
 
 | | Crystal structure of apo D-serine deaminase from Salmonella typhimurium | | Descriptor: | 1,2-ETHANEDIOL, D-serine dehydratase, SULFATE ION | | Authors: | Bharath, S.R, Shveta, B, Savithri, H.S, Murthy, M.R.N. | | Deposit date: | 2011-03-09 | | Release date: | 2011-06-29 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structures of open and closed forms of D-serine deaminase from Salmonella typhimurium - implications on substrate specificity and catalysis
Febs J., 2011
|
|
3O8J
 
 | |
5F0V
 
 | | X-ray crystal structure of a thiolase from Escherichia coli at 1.8 A resolution | | Descriptor: | 1,2-ETHANEDIOL, Acetyl-CoA acetyltransferase | | Authors: | Ithayaraja, M, Neelanjana, J, Wierenga, R, Savithri, H.S, Murthy, M.R.N. | | Deposit date: | 2015-11-28 | | Release date: | 2016-07-13 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of a thiolase from Escherichia coli at 1.8 angstrom resolution.
Acta Crystallogr.,Sect.F, 72, 2016
|
|
5F38
 
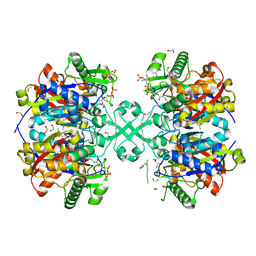 | | X-ray crystal structure of a thiolase from Escherichia coli at 1.8 A resolution | | Descriptor: | 1,2-ETHANEDIOL, Acetyl-CoA acetyltransferase, COENZYME A, ... | | Authors: | Ithayaraja, M, Neelanjana, J, Wierenga, R, Savithri, H.S, Murthy, M.R.N. | | Deposit date: | 2015-12-02 | | Release date: | 2016-07-13 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of a thiolase from Escherichia coli at 1.8 angstrom resolution.
Acta Crystallogr.,Sect.F, 72, 2016
|
|
2HTA
 
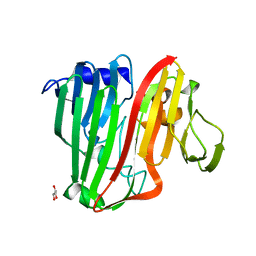 | | Crystal Structure of a putative mutarotase (YeaD) from Salmonella typhimurium in orthorhombic form | | Descriptor: | GLYCEROL, Putative enzyme related to aldose 1-epimerase, SULFATE ION | | Authors: | Chittori, S, Simanshu, D.K, Savithri, H.S, Murthy, M.R.N. | | Deposit date: | 2006-07-25 | | Release date: | 2007-01-23 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure of the putative mutarotase YeaD from Salmonella typhimurium: structural comparison with galactose mutarotases.
Acta Crystallogr.,Sect.D, 63, 2007
|
|
2HTB
 
 | | Crystal Structure of a putative mutarotase (YeaD) from Salmonella typhimurium in monoclinic form | | Descriptor: | Putative enzyme related to aldose 1-epimerase | | Authors: | Chittori, S, Simanshu, D.K, Savithri, H.S, Murthy, M.R.N. | | Deposit date: | 2006-07-25 | | Release date: | 2007-01-23 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structure of the putative mutarotase YeaD from Salmonella typhimurium: structural comparison with galactose mutarotases.
Acta Crystallogr.,Sect.D, 63, 2007
|
|
5YRX
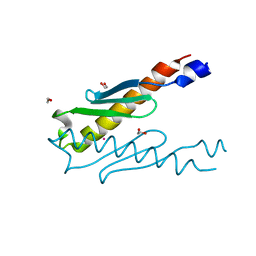 
 | | Crystal structure of a hypothetical protein Rv3716c from Mycobacterium tuberculosis | | Descriptor: | 1,2-ETHANEDIOL, CADMIUM ION, Nucleoid-associated protein Rv3716c | | Authors: | Deka, G, Gopalan, A, Prabhavathi, M, Savithri, H.S, Raja, A, Murthy, M.R.N. | | Deposit date: | 2017-11-10 | | Release date: | 2018-05-16 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural and biophysical characterization of Rv3716c, a hypothetical protein from Mycobacterium tuberculosis
Biochem. Biophys. Res. Commun., 495, 2018
|
|
2W7E
 
 | | Crystal structure of Y51FbsSHMT obtained in the presence of Glycine | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, GLYCINE, PHOSPHATE ION, ... | | Authors: | Rajaram, V, Bhavani, B.S, Bisht, S, Kaul, P, Prakash, V, Appaji Rao, N, Savithri, H.S, Murthy, M.R.N. | | Deposit date: | 2008-12-22 | | Release date: | 2010-08-18 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.69 Å) | | Cite: | Importance of Tyrosine Residues of Bacillus Stearothermophilus Serine Hydroxymethyltransferase in Cofactor Binding and L-Allo-Thr Cleavage.
FEBS J., 275, 2008
|
|
2W7H
 
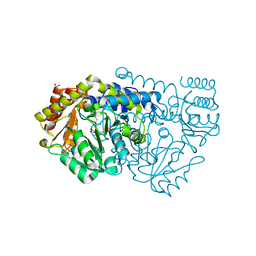 | | Crystal structure of Y51FbsSHMT obtained in the presence of Gly and 5- Formyl Tetrahydrofolate | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, GLYCINE, PHOSPHATE ION, ... | | Authors: | Rajaram, V, Bhavani, B.S, Bisht, S, Kaul, P, Prakash, V, Appaji Rao, N, Savithri, H.S, Murthy, M.R.N. | | Deposit date: | 2008-12-22 | | Release date: | 2010-08-18 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.67 Å) | | Cite: | Importance of Tyrosine Residues of Bacillus Stearothermophilus Serine Hydroxymethyltransferase in Cofactor Binding and L-Allo-Thr Cleavage.
FEBS J., 275, 2008
|
|
2WWS
 
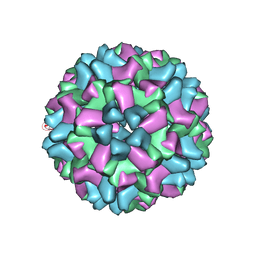 | |
5ZWB
 
 | | Crystal structure of Pyridoxal kinase (PdxK) from Salmonella typhimurium in complex with ADP, PL-linked to Lys233 via a Schiff base | | Descriptor: | 1,2-ETHANEDIOL, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, ADENOSINE-5'-DIPHOSPHATE, ... | | Authors: | Deka, G, Benazir, J.F, Kalyani, J.N, Savithri, H.S, Murthy, M.R.N. | | Deposit date: | 2018-05-14 | | Release date: | 2019-05-29 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural and functional studies on Salmonella typhimurium pyridoxal kinase: the first structural evidence for the formation of Schiff base with the substrate.
Febs J., 286, 2019
|
|
5ZW9
 
 | | Crystal structure of Pyridoxal kinase (PdxK) from Salmonella typhimurium | | Descriptor: | 1,2-ETHANEDIOL, 2-{2-[2-(2-{2-[2-(2-ETHOXY-ETHOXY)-ETHOXY]-ETHOXY}-ETHOXY)-ETHOXY]-ETHOXY}-ETHANOL, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Deka, G, Benazir, J.F, Kalyani, J.N, Savithri, H.S, Murthy, M.R.N. | | Deposit date: | 2018-05-14 | | Release date: | 2019-05-29 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structural and functional studies on Salmonella typhimurium pyridoxal kinase: the first structural evidence for the formation of Schiff base with the substrate.
Febs J., 286, 2019
|
|
5ZWA
 
 | | Crystal structure of Pyridoxal kinase (PdxK) from Salmonella typhimurium in complex with ADP, PL-linked to Lys233 via Schiff base in protomer A and the product (PLP) in protomer B | | Descriptor: | 1,2-ETHANEDIOL, ADENOSINE-5'-DIPHOSPHATE, GLYCEROL, ... | | Authors: | Deka, G, Benazir, J.F, Kalyani, J.N, Savithri, H.S, Murthy, M.R.N. | | Deposit date: | 2018-05-14 | | Release date: | 2019-05-29 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Structural and functional studies on Salmonella typhimurium pyridoxal kinase: the first structural evidence for the formation of Schiff base with the substrate.
Febs J., 286, 2019
|
|
2W7L
 
 | | Crystal structure of Y61AbsSHMT L-allo-Threonine external aldimine | | Descriptor: | ALLO-THREONINE, PYRIDOXAL-5'-PHOSPHATE, SERINE HYDROXYMETHYLTRANSFERASE | | Authors: | Rajaram, V, Bhavani, B.S, Bisht, S, Kaul, P, Prakash, V, Appaji Rao, N, Savithri, H.S, Murthy, M.R.N. | | Deposit date: | 2008-12-22 | | Release date: | 2010-08-18 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.41 Å) | | Cite: | Importance of Tyrosine Residues of Bacillus Stearothermophilus Serine Hydroxymethyltransferase in Cofactor Binding and L-Allo-Thr Cleavage.
FEBS J., 275, 2008
|
|
2W7K
 
 | | Crystal structure of Y61AbsSHMT L-Serine external aldimine | | Descriptor: | PYRIDOXAL-5'-PHOSPHATE, SERINE, SERINE HYDROXYMETHYLTRANSFERASE | | Authors: | Rajaram, V, Bhavani, B.S, Bisht, S, Kaul, P, Prakash, V, Appaji Rao, N, Savithri, H.S, Murthy, M.R.N. | | Deposit date: | 2008-12-22 | | Release date: | 2010-08-18 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.42 Å) | | Cite: | Importance of Tyrosine Residues of Bacillus Stearothermophilus Serine Hydroxymethyltransferase in Cofactor Binding and L-Allo-Thr Cleavage.
FEBS J., 275, 2008
|
|
2W7G
 
 | | Crystal structure of Y51FbsSHMT L-allo-Threonine extrnal aldimine | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, ALLO-THREONINE, PHOSPHATE ION, ... | | Authors: | Rajaram, V, Bhavani, B.S, Bisht, S, Kaul, P, Prakash, V, Appaji Rao, N, Savithri, H.S, Murthy, M.R.N. | | Deposit date: | 2008-12-22 | | Release date: | 2010-08-18 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.92 Å) | | Cite: | Importance of Tyrosine Residues of Bacillus Stearothermophilus Serine Hydroxymethyltransferase in Cofactor Binding and L-Allo-Thr Cleavage.
FEBS J., 275, 2008
|
|
