6IF6
 
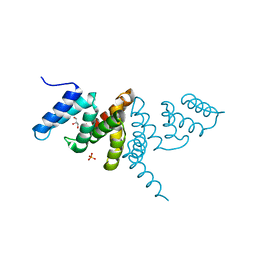 | | Structure of the periplasmic domain of SflA | | Descriptor: | GLYCEROL, PHOSPHATE ION, Protein SflA | | Authors: | Nishikawa, S, Sakuma, M, Kojima, S, Homma, M, Imada, K. | | Deposit date: | 2018-09-18 | | Release date: | 2019-05-01 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure of the periplasmic domain of SflA involved in spatial regulation of the flagellar biogenesis of Vibrio reveals a TPR/SLR-like fold.
J.Biochem., 166, 2019
|
|
2D4U
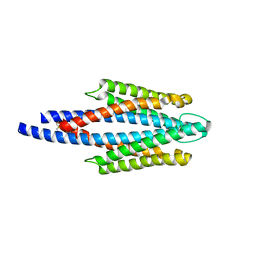 
 | | Crystal Structure of the ligand binding domain of the bacterial serine chemoreceptor Tsr | | Descriptor: | Methyl-accepting chemotaxis protein I | | Authors: | Imada, K, Tajima, H, Namba, K, Sakuma, M, Homma, M, Kawagishi, I. | | Deposit date: | 2005-10-24 | | Release date: | 2006-11-14 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Ligand specificity determined by differentially arranged common ligand-binding residues in the bacterial amino acid chemoreceptors Tsr and Tar.
J.Biol.Chem., 2011
|
|
3ATP
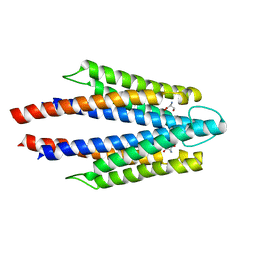 
 | | Structure of the ligand binding domain of the bacterial serine chemoreceptor Tsr with ligand | | Descriptor: | Methyl-accepting chemotaxis protein I, SERINE | | Authors: | Tajima, H, Sakuma, M, Homma, K, Kawagishi, I, Imada, K. | | Deposit date: | 2011-01-07 | | Release date: | 2011-10-19 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Ligand specificity determined by differentially arranged common ligand-binding residues in bacterial amino acid chemoreceptors Tsr and Tar.
J.Biol.Chem., 286, 2011
|
|
5Y40
 
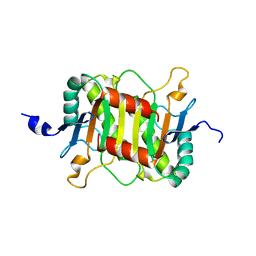 | | Structure of the periplasmic domain of the MotB L119P mutant from Salmonella (crystal form 2) | | Descriptor: | Motility protein B | | Authors: | Takao, M, Kojima, S, Sakuma, M, Homma, M, Imada, K. | | Deposit date: | 2017-07-31 | | Release date: | 2018-04-11 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | The Helix Rearrangement in the Periplasmic Domain of the Flagellar Stator B Subunit Activates Peptidoglycan Binding and Ion Influx.
Structure, 26, 2018
|
|
6AHP
 
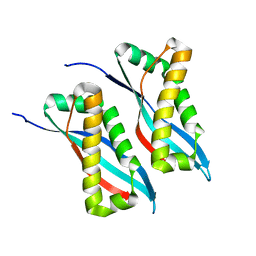 | | Structure of the 58-167 fragment of FliL | | Descriptor: | Flagellar protein FliL | | Authors: | Takekawa, N, Isumi, M, Sakuma, M, Kojima, S, Homma, M, Imada, K. | | Deposit date: | 2018-08-20 | | Release date: | 2019-03-06 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structure ofVibrioFliL, a New Stomatin-like Protein That Assists the Bacterial Flagellar Motor Function.
MBio, 10, 2019
|
|
5Y3Z
 
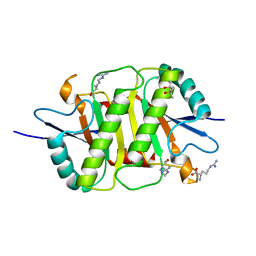 | | Structure of the periplasmic domain of the MotB L119P mutant from Salmonella (crystal form 1) | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, ARGININE, IMIDAZOLE, ... | | Authors: | Takao, M, Kojima, S, Sakuma, M, Homma, M, Imada, K. | | Deposit date: | 2017-07-31 | | Release date: | 2018-04-11 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The Helix Rearrangement in the Periplasmic Domain of the Flagellar Stator B Subunit Activates Peptidoglycan Binding and Ion Influx.
Structure, 26, 2018
|
|
6AHQ
 
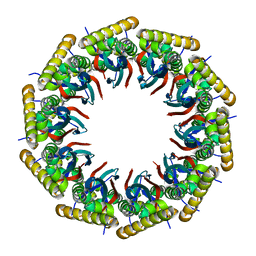 | | Structure of the 40-167 fragment of FliL | | Descriptor: | Flagellar protein FliL | | Authors: | Takekawa, N, Isumi, M, Sakuma, M, Kojima, S, Homma, M, Imada, K. | | Deposit date: | 2018-08-20 | | Release date: | 2019-03-06 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (3.398 Å) | | Cite: | Structure ofVibrioFliL, a New Stomatin-like Protein That Assists the Bacterial Flagellar Motor Function.
MBio, 10, 2019
|
|
3W1E
 
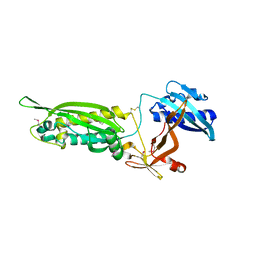 | | Structure of FlgT, a flagellar basal body component protein | | Descriptor: | Flagella basal-body protein | | Authors: | Terashima, H, Sakuma, M, Homma, M, Imada, K. | | Deposit date: | 2012-11-14 | | Release date: | 2013-04-03 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Insight into the assembly mechanism in the supramolecular rings of the sodium-driven Vibrio flagellar motor from the structure of FlgT
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
3WPW
 
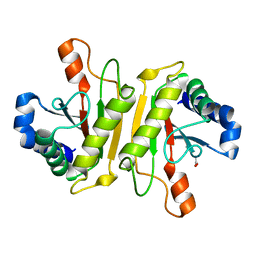 | | Structure of PomBc5, a periplasmic fragment of PomB from Vibrio | | Descriptor: | ACETATE ION, PomB | | Authors: | Takao, M, Sakuma, M, Zhu, S, Homma, M, Kojima, S, Imada, K. | | Deposit date: | 2014-01-17 | | Release date: | 2014-09-10 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Conformational change in the periplasmic region of the flagellar stator coupled with the assembly around the rotor
Proc. Natl. Acad. Sci. U.S.A., 111, 2014
|
|
3WPX
 
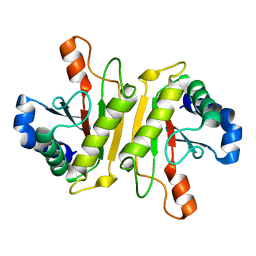 | | Structure of PomBc4, a periplasmic fragment of PomB from Vibrio alginolyticus | | Descriptor: | PomB | | Authors: | Takao, M, Sakuma, M, Zhu, S, Homma, M, Kojima, S, Imada, K. | | Deposit date: | 2014-01-17 | | Release date: | 2014-09-10 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Conformational change in the periplasmic region of the flagellar stator coupled with the assembly around the rotor
Proc. Natl. Acad. Sci. U.S.A., 111, 2014
|
|
3W0D
 
 | | Structure of elastase inhibitor AFUEI (cyrstal form I) | | Descriptor: | Elastase inhibitor AFUEI | | Authors: | Imada, K, Sakuma, M, Okumura, Y, Ogawa, K, Nikai, T, Homma, M. | | Deposit date: | 2012-10-29 | | Release date: | 2013-05-15 | | Last modified: | 2013-07-03 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | X-ray Structure Analysis and Characterization of AFUEI, an Elastase Inhibitor from Aspergillus fumigatus
J.Biol.Chem., 288, 2013
|
|
3W0E
 
 | | Structure of elastase inhibitor AFUEI (crystal form II) | | Descriptor: | Elastase inhibitor AFUEI | | Authors: | Imada, K, Sakuma, M, Okumura, Y, Ogawa, K, Nikai, T, Homma, M. | | Deposit date: | 2012-10-29 | | Release date: | 2013-05-15 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | X-ray Structure Analysis and Characterization of AFUEI, an Elastase Inhibitor from Aspergillus fumigatus
J.Biol.Chem., 288, 2013
|
|
7CIK
 
 | | Structure of the 58-213 fragment of FliF | | Descriptor: | (4R)-2-METHYLPENTANE-2,4-DIOL, Flagellar M-ring protein | | Authors: | Takekawa, T, Sakuma, M, Kojima, S, Homma, M, Imada, K. | | Deposit date: | 2020-07-07 | | Release date: | 2021-02-17 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.29 Å) | | Cite: | Two Distinct Conformations in 34 FliF Subunits Generate Three Different Symmetries within the Flagellar MS-Ring.
Mbio, 12, 2021
|
|
