2DV6
 
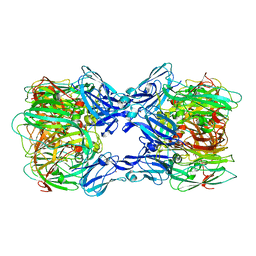 | | Crystal structure of nitrite reductase from Hyphomicrobium denitrificans | | Descriptor: | COPPER (II) ION, Nitrite reductase, POTASSIUM ION | | Authors: | Nojiri, M, Xie, Y, Yamamoto, T, Inoue, T, Suzuki, S, Kai, Y. | | Deposit date: | 2006-07-28 | | Release date: | 2007-02-20 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structure and function of a hexameric copper-containing nitrite reductase.
Proc.Natl.Acad.Sci.USA, 104, 2007
|
|
2ZON
 
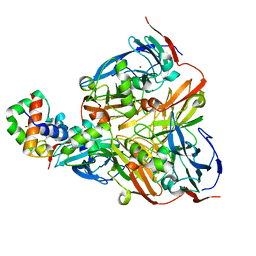 | | Crystal structure of electron transfer complex of nitrite reductase with cytochrome c | | Descriptor: | COPPER (II) ION, Dissimilatory copper-containing nitrite reductase, PROTOPORPHYRIN IX CONTAINING FE, ... | | Authors: | Nojiri, M, Koteishi, H, Yamaguchi, K, Suzuki, S. | | Deposit date: | 2008-05-27 | | Release date: | 2009-06-09 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural basis of inter-protein electron transfer for nitrite reduction in denitrification
Nature, 462, 2009
|
|
2ZOO
 
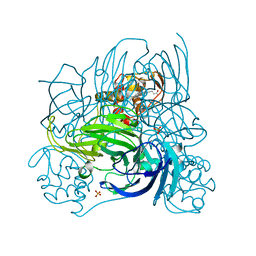 | | Crystal structure of nitrite reductase from Pseudoalteromonas haloplanktis TAC125 | | Descriptor: | COPPER (II) ION, PROTOPORPHYRIN IX CONTAINING FE, Probable nitrite reductase, ... | | Authors: | Nojiri, M, Tsuda, A, Yamaguchi, K, Suzuki, S. | | Deposit date: | 2008-05-27 | | Release date: | 2009-06-09 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Electron transfer processes within and between proteins containing the HEME C and blue Cu
To be Published
|
|
5B1K
 
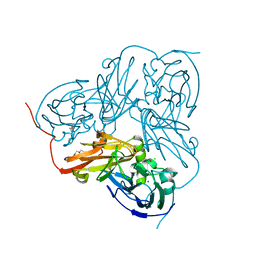 | |
5B1J
 
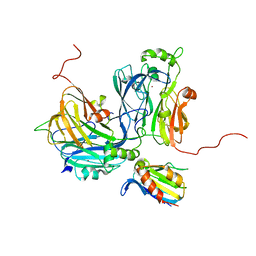 | | Crystal structure of the electron-transfer complex of copper nitrite reductase with a cupredoxin | | Descriptor: | Blue copper protein, COPPER (II) ION, Copper-containing nitrite reductase | | Authors: | Nojiri, M, Koteishi, H, Yoneda, R, Hira, D. | | Deposit date: | 2015-12-04 | | Release date: | 2016-11-23 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structure and Function of Copper Nitrite Reductase
Metalloenzymes in denitrification: Applications and Environmental impacts, 2016
|
|
2CZ6
 
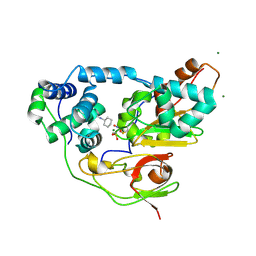 | | Complex of Inactive Fe-type NHase with Cyclohexyl isocyanide | | Descriptor: | CYCLOHEXYL ISOCYANIDE, FE (III) ION, MAGNESIUM ION, ... | | Authors: | Nojiri, M, Kawano, Y, Hashimoto, K, Kamiya, N, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2005-07-11 | | Release date: | 2006-01-11 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | x-ray snap shots of inhibitor binding process in photo-reactive nitrile hydratase
To be Published
|
|
2D0W
 
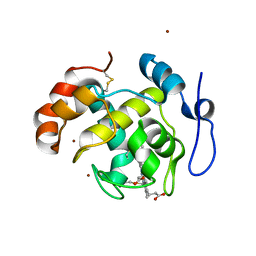 | | Crystal structure of cytochrome cL from Hyphomicrobium denitrificans | | Descriptor: | PROTOPORPHYRIN IX CONTAINING FE, ZINC ION, cytochrome cL | | Authors: | Nojiri, M, Hira, D, Yamaguchi, K, Suzuki, S. | | Deposit date: | 2005-08-10 | | Release date: | 2006-08-10 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | Crystal structures of cytochrome c(L) and methanol dehydrogenase from Hyphomicrobium denitrificans: structural and mechanistic insights into interactions between the two proteins
Biochemistry, 45, 2006
|
|
2D0V
 
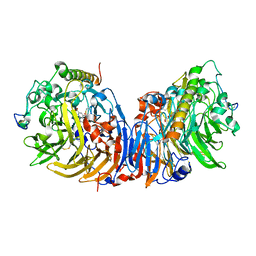 | | Crystal structure of methanol dehydrogenase from Hyphomicrobium denitrificans | | Descriptor: | CALCIUM ION, PYRROLOQUINOLINE QUINONE, methanol dehydrogenase large subunit, ... | | Authors: | Nojiri, M, Hira, D, Yamaguchi, K, Suzuki, S. | | Deposit date: | 2005-08-09 | | Release date: | 2006-08-09 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.49 Å) | | Cite: | Crystal structures of cytochrome c(L) and methanol dehydrogenase from Hyphomicrobium denitrificans: structural and mechanistic insights into interactions between the two proteins
Biochemistry, 45, 2006
|
|
2D0Q
 
 | | Complex of Fe-type NHase with Cyclohexyl isocyanide, photo-activated for 1hr at 277K | | Descriptor: | CYCLOHEXYL ISOCYANIDE, FE (III) ION, MAGNESIUM ION, ... | | Authors: | Nojiri, M, Kawano, Y, Hashimoto, K, Kamiya, N, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2005-08-05 | | Release date: | 2006-02-05 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | X-ray snap shots of Inhibitor Binding Process in Photo-reactive Nitrile Hydratase
To be Published
|
|
2CZ7
 
 | | Fe-type NHase photo-activated for 75min at 105K | | Descriptor: | FE (III) ION, MAGNESIUM ION, NITRIC OXIDE, ... | | Authors: | Nojiri, M, Kawano, Y, Hashimoto, K, Kamiya, N, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2005-07-11 | | Release date: | 2006-01-11 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Active Center Structures of Photo-reactive Nitrile Hydratase during its Photo-activation Process
To be Published
|
|
5GXG
 
 | | High-resolution crystal structure of the electron transfer complex of cytochrome p450cam with putidaredoxin | | Descriptor: | 2,3-DIHYDROXY-1,4-DITHIOBUTANE, Camphor 5-monooxygenase, FE2/S2 (INORGANIC) CLUSTER, ... | | Authors: | Kikui, Y, Hiruma, Y, Ubbink, M, Nojiri, M. | | Deposit date: | 2016-09-17 | | Release date: | 2017-01-18 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Identification of productive and futile encounters in an electron transfer protein complex
Proc. Natl. Acad. Sci. U.S.A., 114, 2017
|
|
3EF4
 
 | | Crystal structure of native pseudoazurin from Hyphomicrobium denitrificans | | Descriptor: | Blue copper protein, COPPER (II) ION, PHOSPHATE ION | | Authors: | Hira, D, Nojiri, M, Suzuki, S. | | Deposit date: | 2008-09-08 | | Release date: | 2008-12-30 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.18 Å) | | Cite: | Atomic resolution structure of pseudoazurin from the methylotrophic denitrifying bacterium Hyphomicrobium denitrificans: structural insights into its spectroscopic properties
Acta Crystallogr.,Sect.D, 65, 2009
|
|
3WIA
 
 | |
3W9C
 
 | | Crystal structure of the electron transfer complex of cytochrome p450cam with putidaredoxin | | Descriptor: | Camphor 5-monooxygenase, FE2/S2 (INORGANIC) CLUSTER, GLYCEROL, ... | | Authors: | Kikui, Y, Hiruma, Y, Hass, M.A, Koteishi, H, Ubbink, M, Nojiri, M. | | Deposit date: | 2013-04-03 | | Release date: | 2013-08-21 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | The structure of the cytochrome p450cam-putidaredoxin complex determined by paramagnetic NMR spectroscopy and crystallography.
J.Mol.Biol., 425, 2013
|
|
3WI9
 
 | | Crystal structure of copper nitrite reductase from Geobacillus kaustophilus | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, ACETIC ACID, COPPER (II) ION, ... | | Authors: | Fukuda, Y, Nojiri, M. | | Deposit date: | 2013-09-09 | | Release date: | 2014-07-23 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Structural and functional characterization of the Geobacillus copper nitrite reductase: involvement of the unique N-terminal region in the interprotein electron transfer with its redox partner
Biochim.Biophys.Acta, 1837, 2014
|
|
