1F85
 
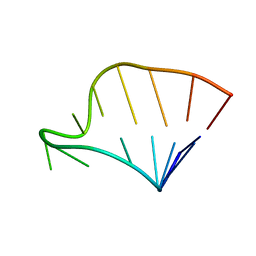 | | SOLUTION STRUCTURE OF HCV IRES RNA DOMAIN IIIE | | Descriptor: | HCV-1B IRES RNA DOMAIN IIIE | | Authors: | Lukavsky, P.J, Otto, G.A, Lancaster, A.M, Sarnow, P, Puglisi, J.D. | | Deposit date: | 2000-06-28 | | Release date: | 2000-11-17 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structures of two RNA domains essential for hepatitis C virus internal ribosome entry site function.
Nat.Struct.Biol., 7, 2000
|
|
1F84
 
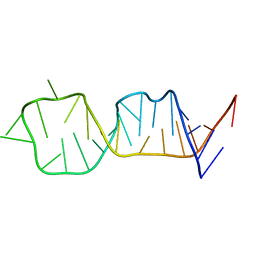 | | SOLUTION STRUCTURE OF HCV IRES RNA DOMAIN IIID | | Descriptor: | HCV-1B IRES RNA DOMAIN IIID | | Authors: | Lukavsky, P.J, Otto, G.A, Lancaster, A.M, Sarnow, P, Puglisi, J.D. | | Deposit date: | 2000-06-28 | | Release date: | 2000-11-17 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structures of two RNA domains essential for hepatitis C virus internal ribosome entry site function.
Nat.Struct.Biol., 7, 2000
|
|
1P5P
 
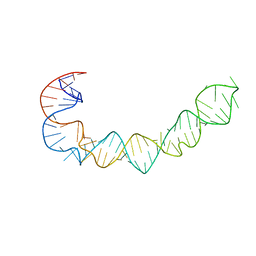 | |
1P5M
 
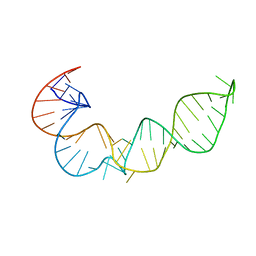 | |
1P5N
 
 | |
1P5O
 
 | |
4BS2
 
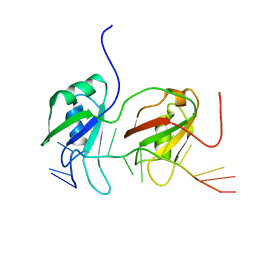 | | NMR structure of human TDP-43 tandem RRMs in complex with UG-rich RNA | | Descriptor: | 5'-R(*GP*UP*GP*UP*GP*AP*AP*UP*GP*AP*AP*UP)-3', TAR DNA-BINDING PROTEIN 43 | | Authors: | Lukavsky, P.J, Daujotyte, D, Tollervey, J.R, Ule, J, Stuani, C, Buratti, E, Baralle, F.E, Damberger, F.F, Allain, F.H.T. | | Deposit date: | 2013-06-06 | | Release date: | 2013-11-13 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | Molecular Basis of Ug-Rich RNA Recognition by the Human Splicing Factor Tdp-43
Nat.Struct.Mol.Biol., 20, 2013
|
|
2HUA
 
 | |
1CQ5
 
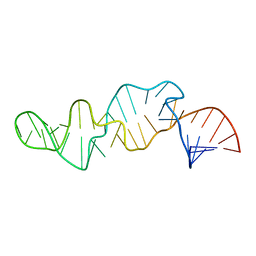 | | NMR STRUCTURE OF SRP RNA DOMAIN IV | | Descriptor: | SRP RNA DOMAIN IV | | Authors: | Schmitz, U, James, T.L, Behrens, S, Freymann, D.M, Lukavsky, P, Walter, P. | | Deposit date: | 1999-08-05 | | Release date: | 1999-08-23 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structure of the phylogenetically most conserved domain of SRP RNA.
RNA, 5, 1999
|
|
6SDY
 
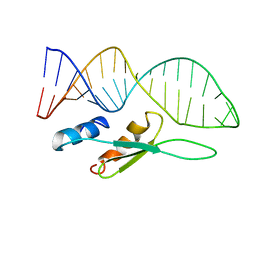 | |
6SDW
 
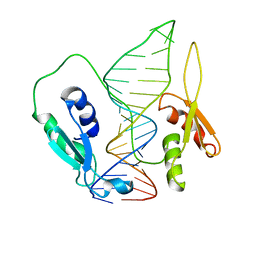 | |
4BL6
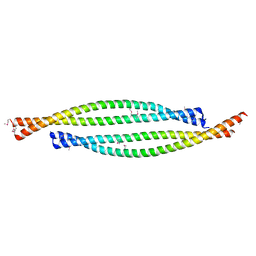 
 | | Bicaudal-D uses a parallel, homodimeric coiled coil with heterotypic registry to co-ordinate recruitment of cargos to dynein | | Descriptor: | ARGININE, PROTEIN BICAUDAL D | | Authors: | Liu, Y, Salter, H.K, Holding, A.N, Johnson, C.M, Stephens, E, Lukavsky, P.J, Walshaw, J, Bullock, S.L. | | Deposit date: | 2013-05-02 | | Release date: | 2013-06-12 | | Last modified: | 2017-07-12 | | Method: | X-RAY DIFFRACTION (2.18 Å) | | Cite: | Bicaudal-D Uses a Parallel, Homodimeric Coiled Coil with Heterotypic Registry to Coordinate Recruitment of Cargos to Dynein
Genes Dev., 27, 2013
|
|
3ZWL
 
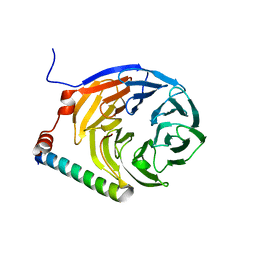 | |
28SR
 
 | |
28SP
 
 | |
2KRB
 
 | | Solution structure of EIF3B-RRM bound to EIF3J peptide | | Descriptor: | Eukaryotic translation initiation factor 3 subunit B, Eukaryotic translation initiation factor 3 subunit J | | Authors: | Elantak, L, Wagner, S, Herrmannova, A, Janoskova, M, Rutkai, E, Lukavsky, P.J, Valasek, L. | | Deposit date: | 2009-12-16 | | Release date: | 2010-01-05 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | The indispensable N-terminal half of eIF3j/HCR1 co-operates with
its structurally conserved binding partner eIF3b/PRT1-RRM and eIF1A in
stringent AUG selection
To be Published
|
|
2NLW
 
 | |
2KUW
 
 | |
2KUU
 
 | |
2KUR
 
 | |
2KUV
 
 | |
