3MHF
 
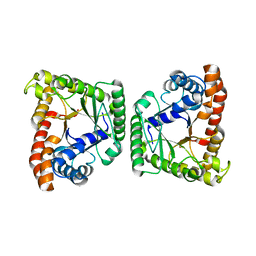 | | Tagatose-1,6-bisphosphate aldolase from Streptococcus pyogenes | | Descriptor: | CALCIUM ION, Tagatose 1,6-diphosphate aldolase 2 | | Authors: | LowKam, C. | | Deposit date: | 2010-04-07 | | Release date: | 2010-04-28 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.87 Å) | | Cite: | Structure of a class I tagatose-1,6-bisphosphate aldolase: investigation into an apparent loss of stereospecificity.
J.Biol.Chem., 285, 2010
|
|
3MHG
 
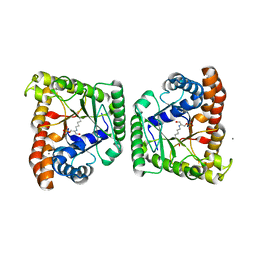 | | Dihydroxyacetone phosphate carbanion intermediate in tagatose-1,6-bisphosphate aldolase from Streptococcus pyogenes | | Descriptor: | 1,3-DIHYDROXYACETONEPHOSPHATE, CALCIUM ION, Tagatose 1,6-diphosphate aldolase 2 | | Authors: | LowKam, C, Liotard, B. | | Deposit date: | 2010-04-07 | | Release date: | 2010-04-28 | | Last modified: | 2019-07-17 | | Method: | X-RAY DIFFRACTION (1.92 Å) | | Cite: | Structure of a class I tagatose-1,6-bisphosphate aldolase: investigation into an apparent loss of stereospecificity.
J.Biol.Chem., 285, 2010
|
|
5F2M
 
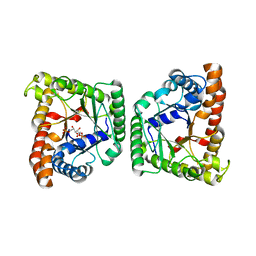 | |
5F2I
 
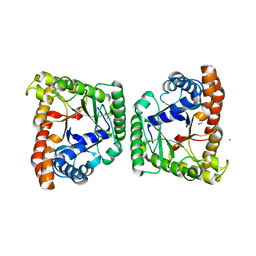 | |
5F2L
 
 | |
5F4S
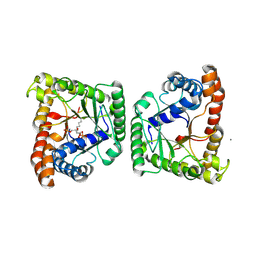 
 | |
5F4X
 
 | |
5F2G
 
 | |
5F4W
 
 | |
4TPJ
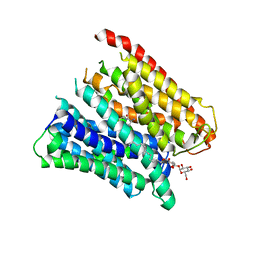 
 | | Selectivity mechanism of a bacterial homologue of the human drug peptide transporters PepT1 and PepT2 | | Descriptor: | ALA-ALA-ALA, DODECYL-BETA-D-MALTOSIDE, Proton:oligopeptide symporter POT family, ... | | Authors: | Guettou, F, Quistgaard, E, Raba, M, Moberg, P, Low, C, Nordlund, P. | | Deposit date: | 2014-06-07 | | Release date: | 2014-07-09 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (3.201 Å) | | Cite: | Selectivity mechanism of a bacterial homolog of the human drug-peptide transporters PepT1 and PepT2.
Nat.Struct.Mol.Biol., 21, 2014
|
|
4TPH
 
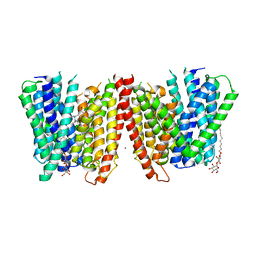 | | Selectivity mechanism of a bacterial homologue of the human drug peptide transporters PepT1 and PepT2 | | Descriptor: | 3,5 DIBROMOTYROSINE, ALANINE, DODECYL-BETA-D-MALTOSIDE, ... | | Authors: | Guettou, F, Quistgaard, E, Raba, M, Moberg, P, Low, C, Nordlund, P. | | Deposit date: | 2014-06-07 | | Release date: | 2014-07-09 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (3.155 Å) | | Cite: | Selectivity mechanism of a bacterial homolog of the human drug-peptide transporters PepT1 and PepT2.
Nat.Struct.Mol.Biol., 21, 2014
|
|
4TPG
 
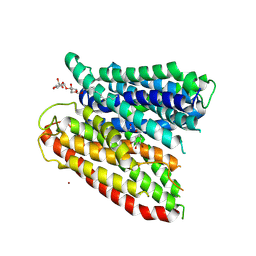 | | Selectivity mechanism of a bacterial homologue of the human drug peptide transporters PepT1 and PepT2 | | Descriptor: | Ala-L-3-Br-Tyr-Ala, DODECYL-BETA-D-MALTOSIDE, Proton:oligopeptide symporter POT family, ... | | Authors: | Guettou, F, Quistgaard, E.M, Raba, M, Moberg, P, Low, C, Nordlund, P. | | Deposit date: | 2014-06-07 | | Release date: | 2014-07-09 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (3.91 Å) | | Cite: | Selectivity mechanism of a bacterial homolog of the human drug-peptide transporters PepT1 and PepT2.
Nat.Struct.Mol.Biol., 21, 2014
|
|
4LEP
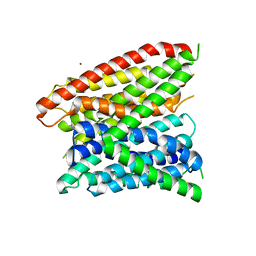 
 | | Structural insights into substrate recognition in proton dependent oligopeptide transporters | | Descriptor: | N-[(1R)-1-phosphonoethyl]-L-alaninamide, Proton:oligopeptide symporter POT family, ZINC ION | | Authors: | Guettou, F, Quistgaard, E.M, Tresaugues, L, Moberg, P, Jegerschold, C, Zhu, L, Jong, A.J, Nordlund, P, Low, C. | | Deposit date: | 2013-06-26 | | Release date: | 2013-07-10 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Structural insights into substrate recognition in proton-dependent oligopeptide transporters.
Embo Rep., 14, 2013
|
|
4JZL
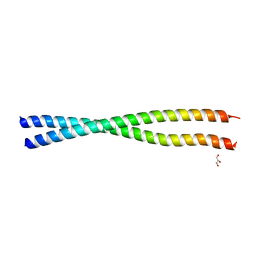 
 | | Crystal structure of BAP31 vDED at alkaline pH | | Descriptor: | B-cell receptor-associated protein 31, GLYCEROL | | Authors: | Quistgaard, E.M, Low, C, Moberg, P, Guettou, F, Maddi, K, Nordlund, P. | | Deposit date: | 2013-04-03 | | Release date: | 2013-09-25 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural and Biophysical Characterization of the Cytoplasmic Domains of Human BAP29 and BAP31.
Plos One, 8, 2013
|
|
4JZP
 
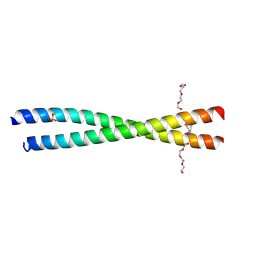 | | Crystal structure of BAP31 vDED at acidic pH | | Descriptor: | B-cell receptor-associated protein 31, SULFATE ION, TETRAETHYLENE GLYCOL | | Authors: | Quistgaard, E.M, Low, C, Moberg, P, Guettou, F, Maddi, K, Nordlund, P. | | Deposit date: | 2013-04-03 | | Release date: | 2013-09-25 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural and Biophysical Characterization of the Cytoplasmic Domains of Human BAP29 and BAP31.
Plos One, 8, 2013
|
|
3TCH
 
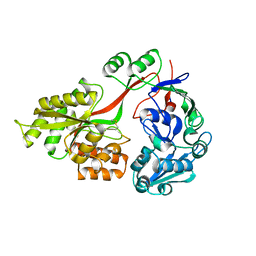 | | Crystal structure of E. coli OppA in an open conformation | | Descriptor: | Periplasmic oligopeptide-binding protein | | Authors: | Klepsch, M.M, Kovermann, M, Low, C, Balbach, J, de Gier, J.W, Slotboom, D.J, Berntsson, R.P.-A. | | Deposit date: | 2011-08-09 | | Release date: | 2011-10-12 | | Last modified: | 2012-01-11 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | Escherichia coli peptide binding protein OppA has a preference for positively charged peptides.
J.Mol.Biol., 414, 2011
|
|
3TCG
 
 | | Crystal structure of E. coli OppA complexed with the tripeptide KGE | | Descriptor: | KGE Peptide, Periplasmic oligopeptide-binding protein | | Authors: | Klepsch, M.M, Kovermann, M, Low, C, Balbach, J, de Gier, J.W, Slotboom, D.J, Berntsson, R.P.-A. | | Deposit date: | 2011-08-09 | | Release date: | 2011-10-12 | | Last modified: | 2017-10-25 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Escherichia coli peptide binding protein OppA has a preference for positively charged peptides.
J.Mol.Biol., 414, 2011
|
|
3TCF
 
 | | Crystal structure of E. coli OppA complexed with endogenous ligands | | Descriptor: | Endogenous peptide, Periplasmic oligopeptide-binding protein | | Authors: | Klepsch, M.M, Kovermann, M, Low, C, Balbach, J, de Gier, J.W, Slotboom, D.J, Berntsson, R.P.-A. | | Deposit date: | 2011-08-09 | | Release date: | 2011-10-12 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Escherichia coli peptide binding protein OppA has a preference for positively charged peptides.
J.Mol.Biol., 414, 2011
|
|
4JA4
 
 | | Inward open conformation of the xylose transporter XylE from E. coli | | Descriptor: | CADMIUM ION, D-xylose-proton symporter | | Authors: | Quistgaard, E.M, Low, C, Moberg, P, Tresaugues, L, Nordlund, P. | | Deposit date: | 2013-02-18 | | Release date: | 2013-05-01 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (4.2 Å) | | Cite: | Structural basis for substrate transport in the GLUT-homology family of monosaccharide transporters.
Nat.Struct.Mol.Biol., 20, 2013
|
|
4JA3
 
 | | Partially occluded inward open conformation of the xylose transporter XylE from E. coli | | Descriptor: | CADMIUM ION, D-xylose-proton symporter, LUTETIUM (III) ION | | Authors: | Quistgaard, E.M, Low, C, Moberg, P, Tresaugues, L, Nordlund, P. | | Deposit date: | 2013-02-18 | | Release date: | 2013-05-01 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (3.8 Å) | | Cite: | Structural basis for substrate transport in the GLUT-homology family of monosaccharide transporters.
Nat.Struct.Mol.Biol., 20, 2013
|
|
4MEW
 
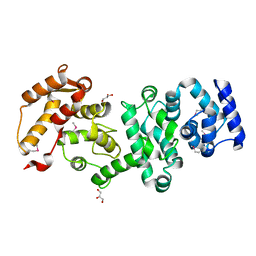 | | Structure of the core fragment of human PR70 | | Descriptor: | CALCIUM ION, GLYCEROL, Serine/threonine-protein phosphatase 2A regulatory subunit B'' subunit beta | | Authors: | Dovega, R.B, Quistgaard, E.M, Tsutakawa, S, Anandapadamanaban, M, Low, C, Nordlund, P. | | Deposit date: | 2013-08-27 | | Release date: | 2014-08-13 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.993 Å) | | Cite: | Structural and Biochemical Characterization of Human PR70 in Isolation and in Complex with the Scaffolding Subunit of Protein Phosphatase 2A.
Plos One, 9, 2014
|
|
4ODO
 
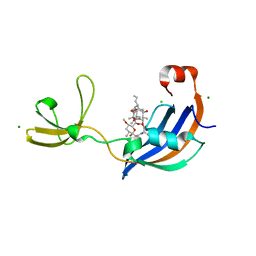 | | Structure of SlyD from Thermus thermophilus in complex with FK506 | | Descriptor: | 2-[BIS-(2-HYDROXY-ETHYL)-AMINO]-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, 8-DEETHYL-8-[BUT-3-ENYL]-ASCOMYCIN, CHLORIDE ION, ... | | Authors: | Quistgaard, E.M, Low, C, Nordlund, P. | | Deposit date: | 2014-01-10 | | Release date: | 2015-01-14 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.599 Å) | | Cite: | Molecular insights into substrate recognition and catalytic mechanism of the chaperone and FKBP peptidyl-prolyl isomerase SlyD.
BMC Biol., 14, 2016
|
|
4ODP
 
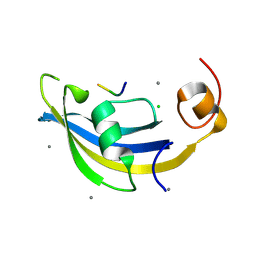 | | Structure of SlyD delta-IF from Thermus thermophilus in complex with S2-W23A peptide | | Descriptor: | 30S ribosomal protein S2, CALCIUM ION, CHLORIDE ION, ... | | Authors: | Quistgaard, E.M, Low, C, Nordlund, P. | | Deposit date: | 2014-01-10 | | Release date: | 2015-01-14 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.747 Å) | | Cite: | Molecular insights into substrate recognition and catalytic mechanism of the chaperone and FKBP peptidyl-prolyl isomerase SlyD.
BMC Biol., 14, 2016
|
|
4ODM
 
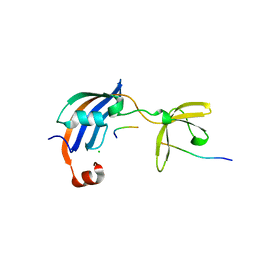 | | Structure of SlyD from Thermus thermophilus in complex with S2-W23A peptide | | Descriptor: | 30S ribosomal protein S2, ACETATE ION, CHLORIDE ION, ... | | Authors: | Quistgaard, E.M, Low, C, Nordlund, P. | | Deposit date: | 2014-01-10 | | Release date: | 2015-01-14 | | Last modified: | 2017-10-04 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Molecular insights into substrate recognition and catalytic mechanism of the chaperone and FKBP peptidyl-prolyl isomerase SlyD.
BMC Biol., 14, 2016
|
|
4ODN
 
 | | Structure of SlyD from Thermus thermophilus in complex with S2-plus peptide | | Descriptor: | 30S ribosomal protein S2, CHLORIDE ION, GLYCEROL, ... | | Authors: | Quistgaard, E.M, Low, C, Nordlund, P. | | Deposit date: | 2014-01-10 | | Release date: | 2015-01-14 | | Last modified: | 2017-10-04 | | Method: | X-RAY DIFFRACTION (1.598 Å) | | Cite: | Molecular insights into substrate recognition and catalytic mechanism of the chaperone and FKBP peptidyl-prolyl isomerase SlyD.
BMC Biol., 14, 2016
|
|
