7P0P
 
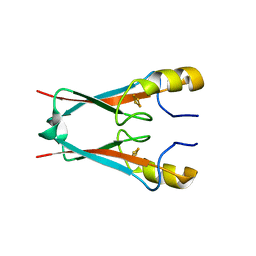 | | NAF-1 bound to M1 molecule | | Descriptor: | 2-benzamido-4-[(2~{R})-1,2,3,4-tetrahydronaphthalen-2-yl]thiophene-3-carboxylic acid, CDGSH iron-sulfur domain-containing protein 2, FE2/S2 (INORGANIC) CLUSTER | | Authors: | Livnah, O, Eisenberg-Domovich, Y, Marjault, H.B, Nechushtai, R. | | Deposit date: | 2021-06-30 | | Release date: | 2022-05-25 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.74 Å) | | Cite: | An anti-diabetic drug targets NEET (CISD) proteins through destabilization of their [2Fe-2S] clusters.
Commun Biol, 5, 2022
|
|
8ASU
 
 | |
8AVP
 
 | |
8AMH
 
 | |
8AN6
 
 | |
8ASR
 
 | |
8AST
 
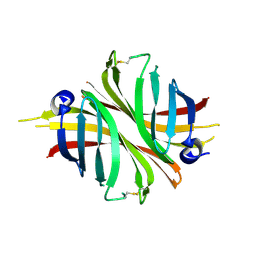 | |
8ASS
 
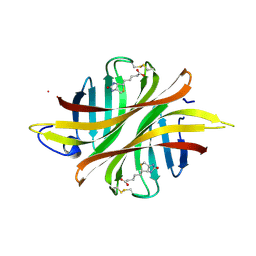 | |
8AVJ
 
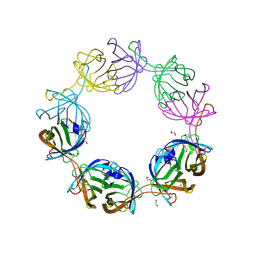 | |
6RTQ
 
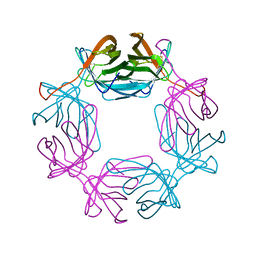 | |
3P63
 
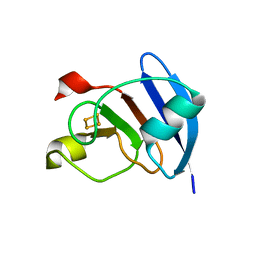 | | Structure of M. laminosus Ferredoxin with a shorter L1,2 loop | | Descriptor: | FE2/S2 (INORGANIC) CLUSTER, Ferredoxin | | Authors: | Livnah, O, Nechushtai, R, Eisenberg-Domovich, Y, Michaeli, D. | | Deposit date: | 2010-10-11 | | Release date: | 2011-02-09 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Allostery in the ferredoxin protein motif does not involve a conformational switch.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
3S2R
 
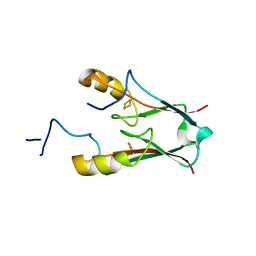 | | ATChloroNEET (H87C mutant) | | Descriptor: | AT5g51720/MIO24_14, FE2/S2 (INORGANIC) CLUSTER | | Authors: | livnah, O, Eisenberg-Domovich, Y, nechushtai, R. | | Deposit date: | 2011-05-17 | | Release date: | 2012-05-23 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.14 Å) | | Cite: | Arabidopsis thaliana ChloroNEET, a Member of the New NEET Family of Human Proteins, is Involved in Development, Senescence and Iron Metabolism.
To be Published
|
|
3S2Q
 
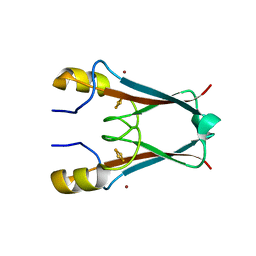 | | The crystal structure of AT5g51720 (AT-NEET) | | Descriptor: | AT5g51720/MIO24_14, FE2/S2 (INORGANIC) CLUSTER, ZINC ION | | Authors: | Livnah, O, Eisenberg-Domovich, Y, Nechushtai, R. | | Deposit date: | 2011-05-17 | | Release date: | 2012-05-23 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Arabidopsis thaliana ChloroNEET, a Member of the New NEET Family of Human Proteins, is Involved in Development, Senescence and Iron Metabolism.
To be Published
|
|
3T2X
 
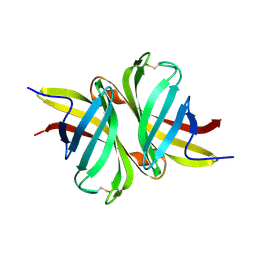 | |
3SZJ
 
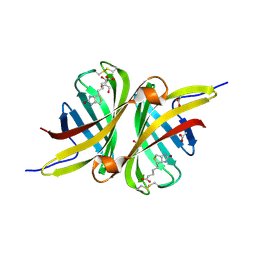 | | Structure of the shwanavidin-biotin complex | | Descriptor: | ACETIC ACID, Avidin/streptavidin, BIOTIN, ... | | Authors: | Livnah, O, Meir, A. | | Deposit date: | 2011-07-19 | | Release date: | 2012-04-11 | | Last modified: | 2012-06-13 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Structural Adaptation of a Thermostable Biotin-binding Protein in a Psychrophilic Environment.
J.Biol.Chem., 287, 2012
|
|
3SZI
 
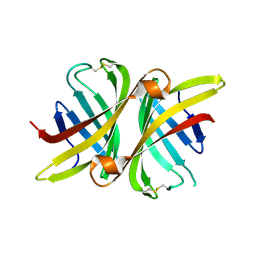 | | Structure of apo shwanavidin (P21 form) | | Descriptor: | Avidin/streptavidin, FORMIC ACID | | Authors: | Livnah, O, Meir, A. | | Deposit date: | 2011-07-19 | | Release date: | 2012-04-25 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Structural Adaptation of a Thermostable Biotin-binding Protein in a Psychrophilic Environment.
J.Biol.Chem., 287, 2012
|
|
4S2Z
 
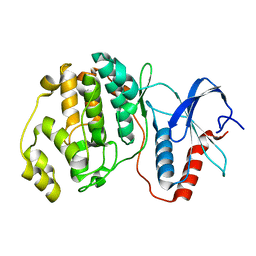 | |
4S34
 
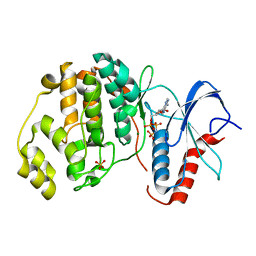 | | ERK2 (I84A) in complex with AMP-PNP | | Descriptor: | MAGNESIUM ION, Mitogen-activated protein kinase 1, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER, ... | | Authors: | Livnah, O, Karamansha, Y, Tzarum, N. | | Deposit date: | 2015-01-26 | | Release date: | 2016-01-27 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Multiple mechanisms render Erk proteins MEK-independent
To be Published
|
|
4S32
 
 | |
4S30
 
 | |
4S31
 
 | |
4S33
 
 | | ERK2 R65S mutant complexed with AMP-PNP | | Descriptor: | MAGNESIUM ION, Mitogen-activated protein kinase 1, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER, ... | | Authors: | Livnah, O, Karamansha, Y, Tzarum, N. | | Deposit date: | 2015-01-26 | | Release date: | 2016-01-27 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.48 Å) | | Cite: | Multiple mechanisms render Erk proteins MEK-independent
To be Published
|
|
3T2W
 
 | |
3SZH
 
 | | Crystal structure of apo shwanavidin (P1 form) | | Descriptor: | Avidin/streptavidin | | Authors: | Livnah, O, Meir, A. | | Deposit date: | 2011-07-19 | | Release date: | 2012-04-11 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.07 Å) | | Cite: | Structural Adaptation of a Thermostable Biotin-binding Protein in a Psychrophilic Environment.
J.Biol.Chem., 287, 2012
|
|
2OF9
 
 | |
