3DX7
 
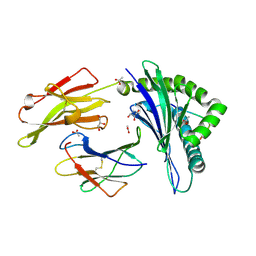 | | Crystal Structure of HLA-B*4403 presenting 10mer EBV antigen | | Descriptor: | ACETATE ION, Beta-2-microglobulin, EBV decapeptide epitope, ... | | Authors: | Archbold, J.K, Ely, L.K, Rossjohn, J. | | Deposit date: | 2008-07-23 | | Release date: | 2009-01-27 | | Last modified: | 2021-10-20 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Natural micropolymorphism in human leukocyte antigens provides a basis for genetic control of antigen recognition.
J.Exp.Med., 206, 2009
|
|
3DX6
 
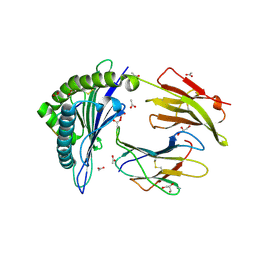 | | Crystal Structure of B*4402 presenting a 10mer EBV epitope | | Descriptor: | ACETATE ION, Beta-2-microglobulin, EBV decapeptide epitope, ... | | Authors: | Archbold, J.K, Ely, L.K, Rossjohn, J. | | Deposit date: | 2008-07-23 | | Release date: | 2009-01-27 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.701 Å) | | Cite: | Natural micropolymorphism in human leukocyte antigens provides a basis for genetic control of antigen recognition.
J.Exp.Med., 206, 2009
|
|
5BZE
 
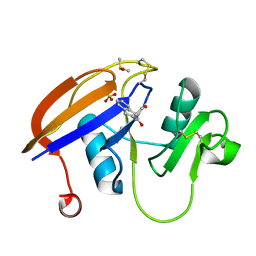 | |
5BZM
 
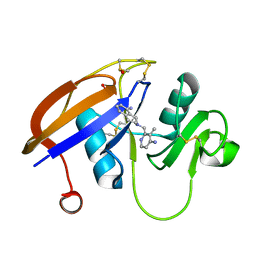 | |
5BZT
 
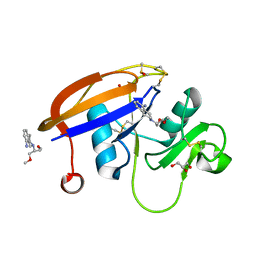 | | Crystal structure of the murine cd44 hyaluronan binding domain complex with a small molecule | | Descriptor: | 2-(1,3-dimethoxypropan-2-yl)-1,2,3,4-tetrahydroisoquinolin-8-amine, CD44 antigen, DIMETHYL SULFOXIDE, ... | | Authors: | Liu, L.K, Finzel, B.C. | | Deposit date: | 2015-06-11 | | Release date: | 2016-06-29 | | Method: | X-RAY DIFFRACTION (1.25 Å) | | Cite: | Crystal structure of the murine cd44 hyaluronan binding domain complex with a small molecule
To Be Published
|
|
5BZF
 
 | |
5BZK
 
 | |
5BZR
 
 | |
5BZH
 
 | |
5BZP
 
 | |
5BZC
 
 | |
5BZI
 
 | |
5BZO
 
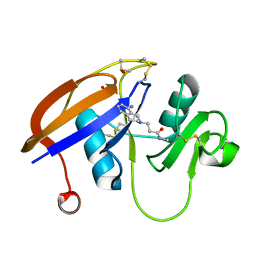 | |
3GB8
 
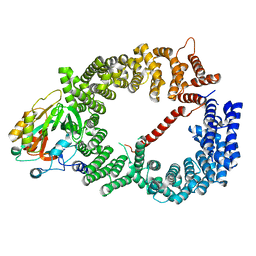 | | Crystal structure of CRM1/Snurportin-1 complex | | Descriptor: | Exportin-1, Snurportin-1 | | Authors: | Dong, X, Biswas, A, Suel, K.E, Jackson, L.K, Martinez, R, Gu, H, Chook, Y.M. | | Deposit date: | 2009-02-19 | | Release date: | 2009-03-31 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Structural basis for leucine-rich nuclear export signal recognition by CRM1.
Nature, 458, 2009
|
|
5BZS
 
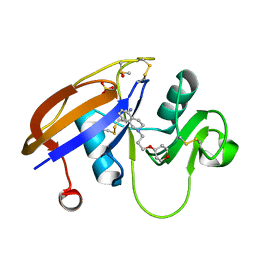 | |
3DFC
 
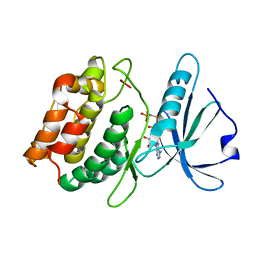 | |
3GU4
 
 | |
3GU6
 
 | |
3GUB
 
 | | Crystal structure of DAPKL93G complexed with N6-(2-Phenylethyl)adenosine | | Descriptor: | 9-alpha-L-lyxofuranosyl-N-(2-phenylethyl)-9H-purin-6-amine, Death-associated protein kinase 1, SULFATE ION | | Authors: | McNamara, L.K, Schumacher, A.M, Schavocky, J.S, Watterson, D.M, Brunzelle, J.S. | | Deposit date: | 2009-03-29 | | Release date: | 2010-03-09 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.71 Å) | | Cite: | Crystal structures of the DAPK gatekeeper mutant complexed with N6-modified adenosine analogs.
To be Published
|
|
3GU7
 
 | |
3GU5
 
 | |
3GU8
 
 | | Crystal structure of DAPKL93G with N6-cyclopentyladenosine | | Descriptor: | Death-associated protein kinase 1, N6-cyclopentyladenosine | | Authors: | McNamara, L.K, Schumacher, A.M, Schavocky, J.S, Watterson, D.M, Brunzelle, J.S. | | Deposit date: | 2009-03-28 | | Release date: | 2010-03-09 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystal structures of the DAPK gatekeeper mutant with N6-modified adenosine analogs.
To be Published
|
|
