5X49
 
 | | Crystal Structure of Human mitochondrial X-prolyl Aminopeptidase (XPNPEP3) | | 分子名称: | (2S,3R)-3-amino-2-hydroxy-4-phenylbutanoic acid, 1,2-ETHANEDIOL, DIMETHYL SULFOXIDE, ... | | 著者 | Singh, R, Kumar, A, Ghosh, B, Jamdar, S, Makde, R.D. | | 登録日 | 2017-02-10 | | 公開日 | 2017-05-17 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (1.65 Å) | | 主引用文献 | Structure of the human aminopeptidase XPNPEP3 and comparison of its in vitro activity with Icp55 orthologs: Insights into diverse cellular processes.
J. Biol. Chem., 292, 2017
|
|
4R60
 
 | | Crystal Structure of Xaa-Pro dipeptidase from Xanthomonas campestris | | 分子名称: | MANGANESE (II) ION, PHOSPHATE ION, Proline dipeptidase, ... | | 著者 | Kumar, A, Ghosh, B, Are, V.N, Jamdar, S.N, Makde, R.D, Sharma, S.M. | | 登録日 | 2014-08-22 | | 公開日 | 2014-09-17 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (1.83 Å) | | 主引用文献 | Crystal Structure of Xaa-Pro dipeptidase from Xanthomonas campestris
to be published
|
|
8J9C
 
 | | Crystal structure of M61 peptidase (apo-form) from Xanthomonas campestris | | 分子名称: | GLYCEROL, Putative glycyl aminopeptidase, SODIUM ION, ... | | 著者 | Yadav, P, Kumar, A, Jamdar, S.N, Makde, R.D. | | 登録日 | 2023-05-03 | | 公開日 | 2024-05-01 | | 最終更新日 | 2024-10-16 | | 実験手法 | X-RAY DIFFRACTION (2.1 Å) | | 主引用文献 | Crystal structure of a newly identified M61 family aminopeptidase with broad substrate specificity that is solely responsible for recycling acidic amino acids.
Febs J., 291, 2024
|
|
8J9D
 
 | | Crystal structure of M61 peptidase (bestatin-bound) from Xanthomonas campestris | | 分子名称: | 2-(3-AMINO-2-HYDROXY-4-PHENYL-BUTYRYLAMINO)-4-METHYL-PENTANOIC ACID, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, GLYCEROL, ... | | 著者 | Yadav, P, Kumar, A, Kulkarni, B.S, Jamdar, S.N, Makde, R.D. | | 登録日 | 2023-05-03 | | 公開日 | 2024-05-01 | | 最終更新日 | 2024-07-31 | | 実験手法 | X-RAY DIFFRACTION (1.9 Å) | | 主引用文献 | Crystal structure of a newly identified M61 family aminopeptidase with broad substrate specificity that is solely responsible for recycling acidic amino acids.
Febs J., 291, 2024
|
|
8JCR
 
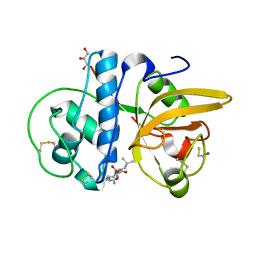 | | Crystal structure of Procerain from Calotropis gigantea (pH 6.0) | | 分子名称: | BETA-MERCAPTOETHANOL, GLYCEROL, N-[N-[1-HYDROXYCARBOXYETHYL-CARBONYL]LEUCYLAMINO-BUTYL]-GUANIDINE, ... | | 著者 | Kumar, A, Jamdar, S.N, Srivastava, G, Makde, R.D. | | 登録日 | 2023-05-11 | | 公開日 | 2024-05-22 | | 実験手法 | X-RAY DIFFRACTION (1.4 Å) | | 主引用文献 | Crystal structure of Procerain from Calotropis gigantea
To Be Published
|
|
8JCS
 
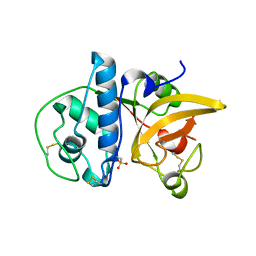 | |
8JCQ
 
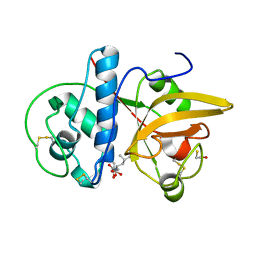 | |
5GIU
 
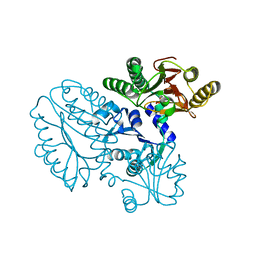 | | Crystal structure of Xaa-Pro peptidase from Deinococcus radiodurans, metal-free active site | | 分子名称: | PHOSPHATE ION, Proline dipeptidase, SODIUM ION | | 著者 | Are, V.N, Kumar, A, Singh, R, Ghosh, B, Jamdar, S.N, Makde, R.D. | | 登録日 | 2016-06-25 | | 公開日 | 2017-06-28 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (1.61 Å) | | 主引用文献 | Xaa-Pro peptidase from Deinococcus radiodurans, metal-free active site
To Be Published
|
|
5CDV
 
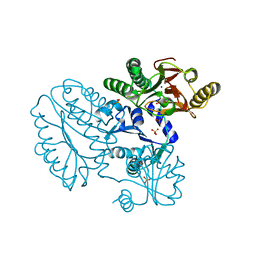 | | Proline dipeptidase from Deinococcus radiodurans R1 | | 分子名称: | MANGANESE (II) ION, PHOSPHATE ION, Proline dipeptidase, ... | | 著者 | Kumar, A, Are, V, Ghosh, B, Jamdar, S, Makde, R. | | 登録日 | 2015-07-05 | | 公開日 | 2016-08-10 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (1.45 Å) | | 主引用文献 | Proline dipeptidase from Deinococcus radiodurans R1 at 1.45 Angstrom resolution
To Be Published
|
|
5CDL
 
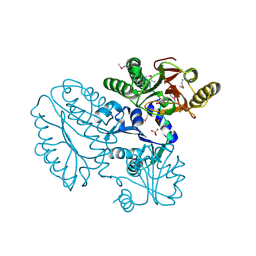 | | Proline dipeptidase from Deinococcus radiodurans (selenomethionine derivative) | | 分子名称: | MANGANESE (II) ION, PHOSPHATE ION, Proline dipeptidase | | 著者 | Kumar, A, Are, V, Ghosh, B, Jamdar, S, Makde, R. | | 登録日 | 2015-07-04 | | 公開日 | 2016-08-10 | | 最終更新日 | 2024-10-30 | | 実験手法 | X-RAY DIFFRACTION (1.8 Å) | | 主引用文献 | Proline dipeptidase from Deinococcus radiodurans (selenomethionine derivative)
To Be Published
|
|
5FCF
 
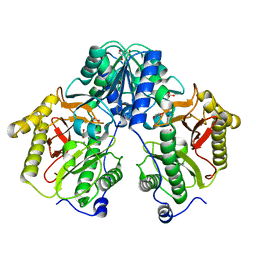 | | Crystal Structure of Xaa-Pro dipeptidase from Xanthomonas campestris, phosphate and Mn bound | | 分子名称: | DI(HYDROXYETHYL)ETHER, GLY-GLY-GLY, GLYCEROL, ... | | 著者 | Kumar, A, Are, V, Ghosh, B, Jamdar, S, Makde, R.D. | | 登録日 | 2015-12-15 | | 公開日 | 2016-12-07 | | 最終更新日 | 2024-03-20 | | 実験手法 | X-RAY DIFFRACTION (1.85 Å) | | 主引用文献 | Crystal structure and biochemical investigations reveal novel mode of substrate selectivity and illuminate substrate inhibition and allostericity in a subfamily of Xaa-Pro dipeptidases.
Biochim. Biophys. Acta, 1865, 2017
|
|
5FCH
 
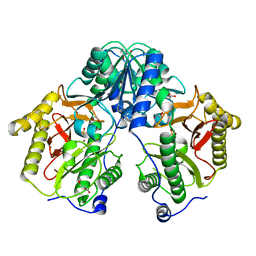 | | Crystal Structure of Xaa-Pro dipeptidase from Xanthomonas campestris, phosphate and Zn bound | | 分子名称: | DI(HYDROXYETHYL)ETHER, GLY-GLY-GLY, GLYCEROL, ... | | 著者 | Kumar, A, Are, V, Ghosh, B, Jamdar, S, Makde, R.D. | | 登録日 | 2015-12-15 | | 公開日 | 2016-12-07 | | 最終更新日 | 2024-03-20 | | 実験手法 | X-RAY DIFFRACTION (1.95 Å) | | 主引用文献 | Crystal structure and biochemical investigations reveal novel mode of substrate selectivity and illuminate substrate inhibition and allostericity in a subfamily of Xaa-Pro dipeptidases
Biochim. Biophys. Acta, 1865, 2017
|
|
5CNX
 
 | | Crystal structure of Xaa-Pro aminopeptidase from Escherichia coli K12 | | 分子名称: | Aminopeptidase YpdF, CACODYLATE ION, GLYCEROL, ... | | 著者 | Kumar, A, Are, V, Ghosh, B, Jamdar, S, Makde, R.D. | | 登録日 | 2015-07-18 | | 公開日 | 2016-07-20 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (2.6 Å) | | 主引用文献 | Structures and activities of widely conserved small prokaryotic aminopeptidases-P clarify classification of M24B peptidases
Proteins, 2018
|
|
5CIK
 
 | | Crystal Structure of Xaa-Pro dipeptidase from Xanthomonas campestris in citrate condition | | 分子名称: | CITRIC ACID, GLYCEROL, Proline dipeptidase | | 著者 | Kumar, A, Are, V, Ghosh, B, Agrawal, U, Jamdar, S, Makde, R.D. | | 登録日 | 2015-07-13 | | 公開日 | 2016-07-20 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (2.2 Å) | | 主引用文献 | Crystal Structure of Xaa-Pro dipeptidase from Xanthomonas campestris in citrate condition
To Be Published
|
|
5CDE
 
 | | R372A mutant of Xaa-Pro dipeptidase from Xanthomonas campestris | | 分子名称: | Proline dipeptidase, SULFATE ION, ZINC ION | | 著者 | Kumar, A, Are, V, Ghosh, B, Jamdar, S, Makde, R. | | 登録日 | 2015-07-03 | | 公開日 | 2016-09-14 | | 最終更新日 | 2024-03-20 | | 実験手法 | X-RAY DIFFRACTION (1.85 Å) | | 主引用文献 | R372A mutant of Xaa-Pro dipeptidase from Xanthomonas campestris at 1.85 Angstrom resolution
To Be Published
|
|
5YZN
 
 | | Crystal structure of S9 peptidase (active form) from Deinococcus radiodurans R1 | | 分子名称: | Acyl-peptide hydrolase, putative | | 著者 | Yadav, P, Jamdar, S.N, Kumar, A, Ghosh, B, Makde, R.D. | | 登録日 | 2017-12-15 | | 公開日 | 2018-11-14 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (2.3 Å) | | 主引用文献 | Carboxypeptidase in prolyl oligopeptidase family: Unique enzyme activation and substrate-screening mechanisms.
J.Biol.Chem., 294, 2019
|
|
5YZM
 
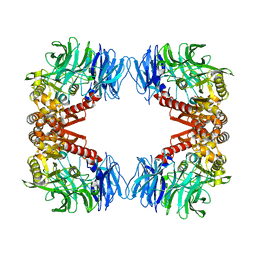 | | Crystal structure of S9 peptidase (inactive form) from Deinococcus radiodurans R1 | | 分子名称: | ACETATE ION, Acyl-peptide hydrolase, putative | | 著者 | Yadav, P, Jamdar, S.N, Kumar, A, Ghosh, B, Makde, R.D. | | 登録日 | 2017-12-15 | | 公開日 | 2018-11-14 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (2.3 Å) | | 主引用文献 | Carboxypeptidase in prolyl oligopeptidase family: Unique enzyme activation and substrate-screening mechanisms.
J.Biol.Chem., 294, 2019
|
|
5YZO
 
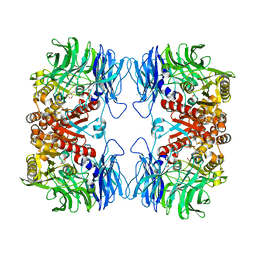 | | Crystal structure of S9 peptidase mutant (S514A) from Deinococcus radiodurans R1 | | 分子名称: | Acyl-peptide hydrolase, putative, DIMETHYL SULFOXIDE, ... | | 著者 | Yadav, P, Jamdar, S.N, Kumar, A, Ghosh, B, Makde, R.D. | | 登録日 | 2017-12-15 | | 公開日 | 2018-11-14 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (1.7 Å) | | 主引用文献 | Carboxypeptidase in prolyl oligopeptidase family: Unique enzyme activation and substrate-screening mechanisms.
J.Biol.Chem., 294, 2019
|
|
5GIQ
 
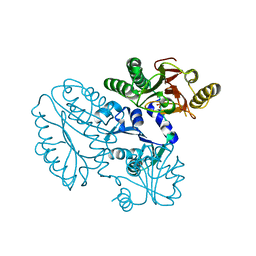 | | Xaa-Pro peptidase from Deinococcus radiodurans, Zinc bound | | 分子名称: | PHOSPHATE ION, Proline dipeptidase, ZINC ION | | 著者 | Are, V.N, Singh, R, Kumar, A, Ghosh, B, Jamdar, S.N, Makde, R.D. | | 登録日 | 2016-06-24 | | 公開日 | 2017-06-28 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (1.8 Å) | | 主引用文献 | Structures and activities of widely conserved small prokaryotic aminopeptidases-P clarify classification of M24B peptidases.
Proteins, 2018
|
|
5GIV
 
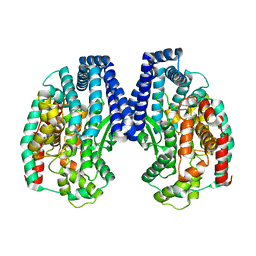 | | Crystal structure of M32 carboxypeptidase from Deinococcus radiodurans R1 | | 分子名称: | ACETATE ION, Carboxypeptidase 1, ZINC ION | | 著者 | Sharma, B, Singh, R, Yadav, P, Ghosh, B, Kumar, A, Jamdar, S.N, Makde, R.D. | | 登録日 | 2016-06-25 | | 公開日 | 2017-07-12 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (2.4 Å) | | 主引用文献 | Active site gate of M32 carboxypeptidases illuminated by crystal structure and molecular dynamics simulations
Biochim. Biophys. Acta, 1865, 2017
|
|
