7A0J
 
 | |
7A0K
 
 | |
7Z67
 
 | |
7Z66
 
 | |
1RJ4
 
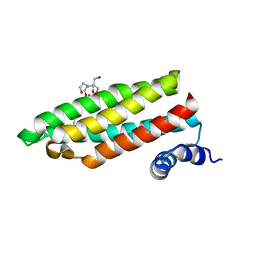 | | Structure of a Cell Wall Invertase Inhibitor from Tobacco in Complex with Cd2+ | | Descriptor: | 2-[BIS-(2-HYDROXY-ETHYL)-AMINO]-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, CADMIUM ION, invertase inhibitor | | Authors: | Hothorn, M, D'Angelo, I, Marquez, J.A, Greiner, S, Scheffzek, K. | | Deposit date: | 2003-11-18 | | Release date: | 2004-02-03 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The invertase inhibitor Nt-CIF from tobacco: a highly thermostable four-helix bundle with an unusual N-terminal extension
J.Mol.Biol., 335, 2004
|
|
1RJ1
 
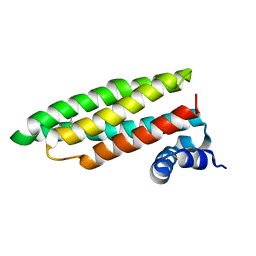 | | Crystal Structure of a Cell Wall Invertase Inhibitor from Tobacco | | Descriptor: | invertase inhibitor | | Authors: | Hothorn, M, D'Angelo, I, Marquez, J.A, Greiner, S, Scheffzek, K. | | Deposit date: | 2003-11-18 | | Release date: | 2004-02-03 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (1.87 Å) | | Cite: | The invertase inhibitor Nt-CIF from tobacco: a highly thermostable four-helix bundle with an unusual N-terminal extension
J.Mol.Biol., 335, 2004
|
|
2XQR
 
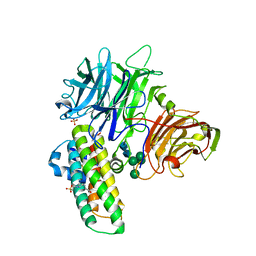 | | Crystal structure of plant cell wall invertase in complex with a specific protein inhibitor | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, ... | | Authors: | Hothorn, M, Van den Ende, W, Lammens, W, Rybin, V, Scheffzek, K. | | Deposit date: | 2010-09-07 | | Release date: | 2010-10-06 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.58 Å) | | Cite: | Structural Insights Into the Ph-Controlled Targeting of Plant Cell-Wall Invertase by a Specific Inhibitor Protein.
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
8QT5
 
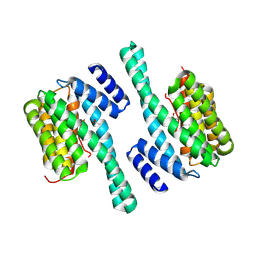 | |
8QTC
 
 | |
8QTT
 
 | |
8QTF
 
 | |
3RJ0
 
 | | Plant steroid receptor BRI1 ectodomain in complex with brassinolide | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Brassinolide, Protein BRASSINOSTEROID INSENSITIVE 1, ... | | Authors: | Hothorn, M. | | Deposit date: | 2011-04-14 | | Release date: | 2011-06-22 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.541 Å) | | Cite: | Structural basis of steroid hormone perception by the receptor kinase BRI1.
Nature, 474, 2011
|
|
3RIZ
 
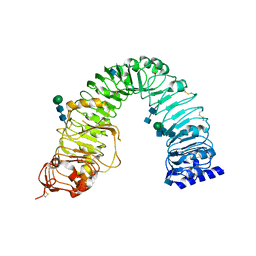 | | Crystal structure of the plant steroid receptor BRI1 ectodomain | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Protein BRASSINOSTEROID INSENSITIVE 1, ... | | Authors: | Hothorn, M. | | Deposit date: | 2011-04-14 | | Release date: | 2011-06-22 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.523 Å) | | Cite: | Structural basis of steroid hormone perception by the receptor kinase BRI1.
Nature, 474, 2011
|
|
6FIF
 
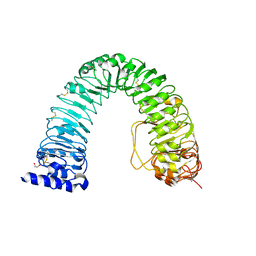 | | Crystal structure of the BRI1 Gly644-Asp (bri1-6) mutant from Arabidopsis thaliana. | | Descriptor: | 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Hothorn, M, Santiago, J, Hohmann, U. | | Deposit date: | 2018-01-18 | | Release date: | 2018-01-31 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.54 Å) | | Cite: | Mechanistic basis for the activation of plant membrane receptor kinases by SERK-family coreceptors.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
6G3W
 
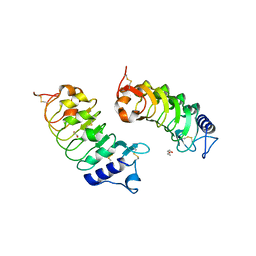 | |
6FG8
 
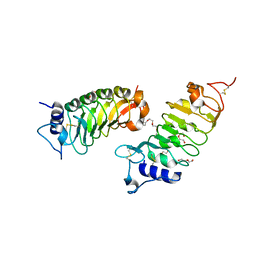 | | Crystal structure of the BIR3 - SERK1 complex from Arabidopsis thaliana. | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, DI(HYDROXYETHYL)ETHER, PENTAETHYLENE GLYCOL, ... | | Authors: | Hothorn, M, Hohmann, U. | | Deposit date: | 2018-01-10 | | Release date: | 2018-01-24 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.25 Å) | | Cite: | The SERK3 elongated allele defines a role for BIR ectodomains in brassinosteroid signalling.
Nat Plants, 4, 2018
|
|
6FG7
 
 | | Crystal structure of the BIR2 ectodomain from Arabidopsis thaliana. | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, Inactive LRR receptor-like serine/threonine-protein kinase BIR2, ... | | Authors: | Hothorn, M, Hohmann, U. | | Deposit date: | 2018-01-10 | | Release date: | 2018-01-24 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The SERK3 elongated allele defines a role for BIR ectodomains in brassinosteroid signalling.
Nat Plants, 4, 2018
|
|
6TO9
 
 | |
6TO5
 
 | |
6TOC
 
 | |
3IID
 
 | | Crystal structure of the macro domain of human histone macroH2A1.1 in complex with ADP-ribose (form A) | | Descriptor: | ADENOSINE-5-DIPHOSPHORIBOSE, Core histone macro-H2A.1, Isoform 1, ... | | Authors: | Hothorn, M, Bortfeld, M, Ladurner, A.G, Scheffzek, K. | | Deposit date: | 2009-07-31 | | Release date: | 2009-08-18 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | A macrodomain-containing histone rearranges chromatin upon sensing PARP1 activation.
Nat.Struct.Mol.Biol., 16, 2009
|
|
3G3O
 
 | |
3G3U
 
 | |
3IIF
 
 | | Crystal structure of the macro domain of human histone macroH2A1.1 in complex with ADP-ribose (form B) | | Descriptor: | ADENOSINE-5-DIPHOSPHORIBOSE, Core histone macro-H2A.1, Isoform 1 | | Authors: | Hothorn, M, Bortfeld, M, Ladurner, A.G, Scheffzek, K. | | Deposit date: | 2009-07-31 | | Release date: | 2009-08-18 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | A macrodomain-containing histone rearranges chromatin upon sensing PARP1 activation.
Nat.Struct.Mol.Biol., 16, 2009
|
|
3G3R
 
 | |
