7TU4
 
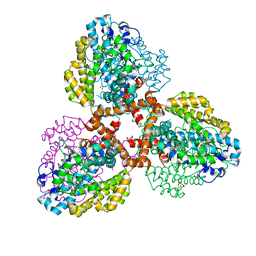 | | Structure of the L. blandensis dGTPase del55-58 mutant bound to Mn | | Descriptor: | 1,2-ETHANEDIOL, MANGANESE (II) ION, SULFATE ION, ... | | Authors: | Sikkema, A.P, Klemm, B.P, Horng, J.C, Hall, T.M.T. | | Deposit date: | 2022-02-02 | | Release date: | 2022-06-01 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.26 Å) | | Cite: | High-resolution structures of the SAMHD1 dGTPase homolog from Leeuwenhoekiella blandensis reveal a novel mechanism of allosteric activation by dATP.
J.Biol.Chem., 298, 2022
|
|
7LWZ
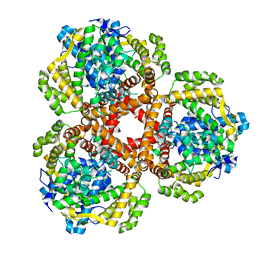 
 | | Apo Structure of Vibrio cholerae dGTPase protein VC1979 | | Descriptor: | Deoxyguanosinetriphosphate triphosphohydrolase-like protein 1, NICKEL (II) ION | | Authors: | Sikkema, A.P, Horng, J, Klemm, B.P, Schaaper, R.M, Hall, T.M.T. | | Deposit date: | 2021-03-02 | | Release date: | 2021-03-10 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.32 Å) | | Cite: | Structure of Vibrio cholerae dGTPase protein VC1979
To Be Published
|
|
3Q0L
 
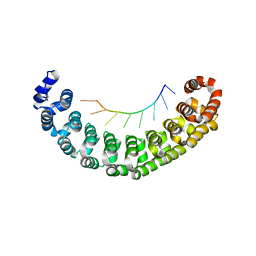 | |
3Q0M
 
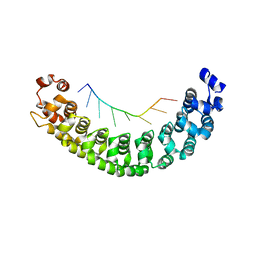 | |
3Q0S
 
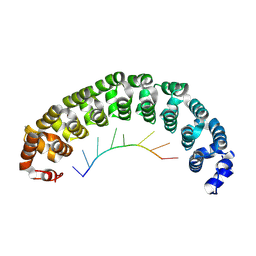 | |
3Q0Q
 
 | |
3Q0P
 
 | |
3Q0R
 
 | |
3Q0O
 
 | |
3Q0N
 
 | |
3QG9
 
 | | crystal structure of FBF-2/gld-1 FBEa A7U mutant complex | | Descriptor: | 1,2-ETHANEDIOL, 5'-R(*UP*GP*UP*GP*CP*CP*UP*UP*A)-3', Fem-3 mRNA-binding factor 2 | | Authors: | Koh, Y.Y, Wang, Y, Qiu, C, Opperman, L, Gross, L, Hall, T.M.T, Wickens, M. | | Deposit date: | 2011-01-24 | | Release date: | 2011-03-23 | | Last modified: | 2018-01-24 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Stacking interactions in PUF-RNA complexes.
Rna, 17, 2011
|
|
3QGC
 
 | | Crystal structure of FBF-2 R288Y mutant in complex with gld-1 FBEa A7U mutant | | Descriptor: | 5'-R(*UP*GP*UP*GP*CP*CP*UP*UP*A)-3', Fem-3 mRNA-binding factor 2 | | Authors: | Wang, Y, Qiu, C, Koh, Y.Y, Opperman, L, Gross, L, Hall, T.M.T, Wickens, M. | | Deposit date: | 2011-01-24 | | Release date: | 2011-03-23 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Stacking interactions in PUF-RNA complexes.
Rna, 17, 2011
|
|
