7A38
 
 | |
7A2X
 
 | |
7A30
 
 | |
7A34
 
 | |
7A2S
 
 | |
7A2K
 
 | |
7A3B
 
 | |
7A2L
 
 | |
7A2U
 
 | |
7A2O
 
 | |
7A3D
 
 | |
7A2J
 
 | |
7A2V
 
 | |
7A37
 | |
7A2Q
 
 | |
7A33
 | |
7A2W
 
 | |
7A3A
 
 | |
7A2P
 
 | |
7A2N
 
 | |
7A2M
 
 | |
7A2R
 
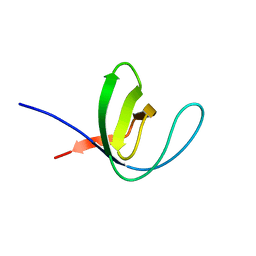 | |
3FJ5
 
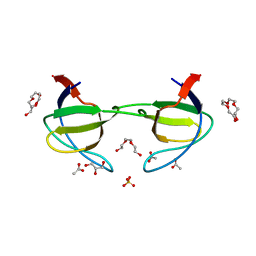 | | Crystal structure of the c-src-SH3 domain | | Descriptor: | ACETATE ION, GLYCEROL, Proto-oncogene tyrosine-protein kinase Src, ... | | Authors: | Camara-Artigas, A. | | Deposit date: | 2008-12-14 | | Release date: | 2009-03-03 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Intertwined dimeric structure for the SH3 domain of the c-Src tyrosine kinase induced by polyethylene glycol binding
Febs Lett., 583, 2009
|
|
2OLP
 
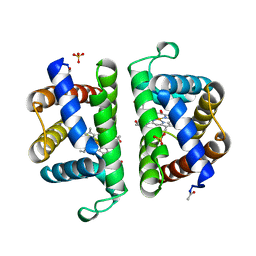 | | Structure and ligand selection of hemoglobin II from Lucina pectinata | | Descriptor: | Hemoglobin II, OXYGEN MOLECULE, PROTOPORPHYRIN IX CONTAINING FE, ... | | Authors: | Gavira, J.A, Camara-Artigas, A, de Jesus, W, Lopez-Garriga, J, Garcia-Ruiz, J.M. | | Deposit date: | 2007-01-19 | | Release date: | 2007-12-18 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.932 Å) | | Cite: | Structure and Ligand Selection of Hemoglobin II from Lucina pectinata
J.Biol.Chem., 283, 2008
|
|
3M0R
 
 | |
