2L18
 
 | |
2L19
 
 | | An arsenate reductase in the intermediate state | | Descriptor: | Arsenate reductase | | Authors: | Yu, C, Xia, B, Jin, C. | | Deposit date: | 2010-07-26 | | Release date: | 2011-04-13 | | Last modified: | 2011-07-13 | | Method: | SOLUTION NMR | | Cite: | (1)H, (13)C and (15)N resonance assignments of the arsenate reductase from Synechocystis sp. strain PCC 6803
Biomol.Nmr Assign., 5, 2011
|
|
1ZGG
 
 | |
2B8F
 
 | |
2B5X
 
 | |
2B5Y
 
 | |
2B8G
 
 | |
2OZX
 
 | |
2OZW
 
 | |
6AF4
 
 | |
6AF3
 
 | |
2K2V
 
 | |
2K4K
 
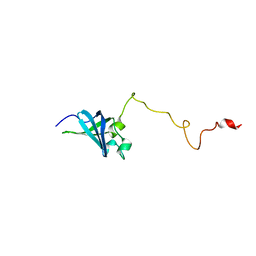 | | Solution structure of GSP13 from Bacillus subtilis | | Descriptor: | General stress protein 13 | | Authors: | Yu, W, Yu, B, Hu, J, Jin, C, Xia, B. | | Deposit date: | 2008-06-13 | | Release date: | 2009-05-12 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | Solution structure of GSP13 from Bacillus subtilis exhibits an S1 domain related to cold shock proteins.
J.Biomol.Nmr, 43, 2009
|
|
2KDV
 
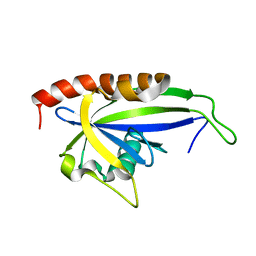 | |
2KNQ
 
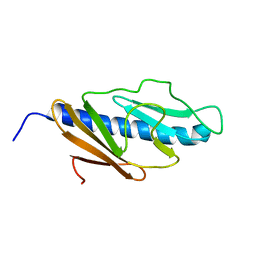 | | Solution structure of E.Coli GspH | | Descriptor: | General secretion pathway protein H | | Authors: | Zhang, Y, Jin, C. | | Deposit date: | 2009-08-31 | | Release date: | 2010-09-08 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution structure of GspH
To be Published
|
|
2KDW
 
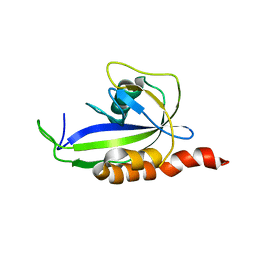 | |
2KCW
 
 | |
2KML
 
 | |
2KW1
 
 | | Solution structure of CTD | | Descriptor: | Fas apoptotic inhibitory molecule 1 | | Authors: | Ma, S, Zheng, X, Jin, C. | | Deposit date: | 2010-03-30 | | Release date: | 2011-04-13 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution structure of CTD
To be Published
|
|
2L16
 
 | |
2MN7
 
 | |
2MN6
 
 | |
2MI2
 
 | |
2MK6
 
 | |
2MRI
 
 | |
