8FI3
 
 | |
8FI4
 
 | |
8FT7
 
 | |
8FUY
 
 | |
8G0V
 
 | |
8G0R
 
 | |
8G0T
 
 | |
8G0U
 
 | |
8G0S
 
 | |
4DC5
 
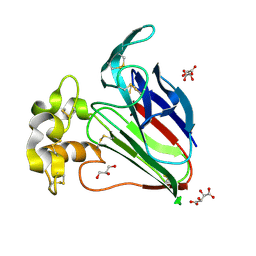 | | Crystal Structure of Thaumatin Unexposed to Excessive SONICC Imaging Laser Dose. | | Descriptor: | GLYCEROL, L(+)-TARTARIC ACID, Thaumatin I | | Authors: | Mulichak, A.M, Becker, M, Kissick, D.J, Keefe, L.J, Fischetti, R.F, Simpson, G.J. | | Deposit date: | 2012-01-17 | | Release date: | 2013-01-23 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.48 Å) | | Cite: | Towards protein-crystal centering using second-harmonic generation (SHG) microscopy.
Acta Crystallogr.,Sect.D, 69, 2013
|
|
4DC8
 
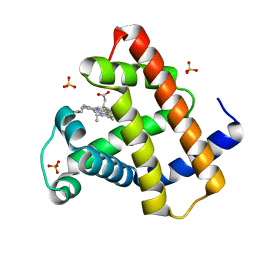 | | Crystal Structure of Myoglobin Unexposed to Excessive SONICC Imaging Laser Dose. | | Descriptor: | Myoglobin, PROTOPORPHYRIN IX CONTAINING FE, SULFATE ION | | Authors: | Becker, M, Mulichak, A.M, Kissick, D.J, Fischetti, R.F, Keefe, L.J, Simpson, D.J. | | Deposit date: | 2012-01-17 | | Release date: | 2013-01-23 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Towards protein-crystal centering using second-harmonic generation (SHG) microscopy.
Acta Crystallogr.,Sect.D, 69, 2013
|
|
4DC6
 
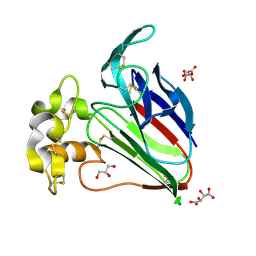 | | Crystal Structure of Thaumatin Exposed to Excessive SONICC Imaging Laser Dose. | | Descriptor: | GLYCEROL, L(+)-TARTARIC ACID, Thaumatin I | | Authors: | Mulichak, A.M, Becker, M, Kissick, D.J, Keefe, L.J, Fischetti, R.F, Simpson, G.J. | | Deposit date: | 2012-01-17 | | Release date: | 2013-01-23 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.48 Å) | | Cite: | Towards protein-crystal centering using second-harmonic generation (SHG) microscopy.
Acta Crystallogr.,Sect.D, 69, 2013
|
|
4DC7
 
 | | Crystal Structure of Myoglobin Exposed to Excessive SONICC Imaging Laser Dose. | | Descriptor: | Myoglobin, PROTOPORPHYRIN IX CONTAINING FE, SULFATE ION | | Authors: | Becker, M, Mulichak, A.M, Kissick, D.J, Fischetti, R.F, Keefe, L.J, Simpson, G.J. | | Deposit date: | 2012-01-17 | | Release date: | 2013-01-23 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Towards protein-crystal centering using second-harmonic generation (SHG) microscopy.
Acta Crystallogr.,Sect.D, 69, 2013
|
|
7TMB
 
 | |
7TMF
 | |
7TM4
 
 | |
7TM8
 
 | |
7TM7
 | |
7TMV
 | |
7TM6
 
 | |
7TMG
 | |
7TM9
 
 | |
7TM5
 
 | |
7TMD
 
 | |
3KOH
 
 | | Cytochrome P450 2E1 with omega-imidazolyl octanoic acid | | Descriptor: | 8-(1H-imidazol-1-yl)octanoic acid, Cytochrome P450 2E1, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Scott, E.E, Porubsky, P.R. | | Deposit date: | 2009-11-13 | | Release date: | 2010-05-12 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Human cytochrome P450 2E1 structures with fatty acid analogs reveal a previously unobserved binding mode.
J.Biol.Chem., 285, 2010
|
|
