1TCJ
 
 | |
1TCG
 
 | |
1TCK
 
 | |
5BJ4
 
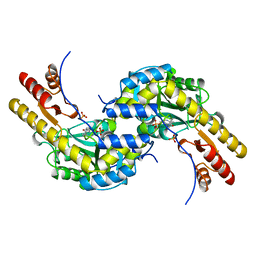 | | THERMUS THERMOPHILUS ASPARTATE AMINOTRANSFERASE TETRA MUTANT 2 | | Descriptor: | PHOSPHATE ION, PROTEIN (ASPARTATE AMINOTRANSFERASE), PYRIDOXAL-5'-PHOSPHATE | | Authors: | Ura, H, Nakai, T, Kawaguchi, S.I, Miyahara, I, Hirotsu, K, Kuramitsu, S. | | Deposit date: | 1999-01-11 | | Release date: | 2003-09-02 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Substrate recognition mechanism of thermophilic dual-substrate enzyme
J.BIOCHEM.(TOKYO), 130, 2001
|
|
3WQB
 
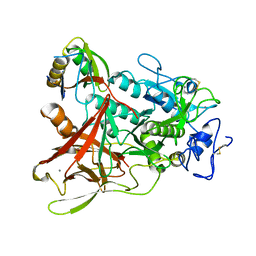 | | Crystal structure of aeromonas sobria serine protease (ASP) and the chaperone (ORF2) complex | | Descriptor: | CALCIUM ION, Extracellular serine protease, Open reading frame 2 | | Authors: | Kobayashi, H, Yoshida, T, Miyakawa, T, Kato, R, Tashiro, M, Yamanaka, H, Tanokura, M, Tsuge, H. | | Deposit date: | 2014-01-24 | | Release date: | 2015-03-25 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.41 Å) | | Cite: | Structural Basis for Action of the External Chaperone for a Propeptide-deficient Serine Protease from Aeromonas sobria.
J.Biol.Chem., 290, 2015
|
|
