1EST
 
 | |
1DBS
 
 | |
1UZ2
 
 | | The Cys121Ser Mutant of Beta-Lactoglobulin | | Descriptor: | BETA-LACTOGLOBULIN | | Authors: | McNae, I, Jayat, D, Haertle, T, Holt, C, Sawyer, L. | | Deposit date: | 2004-03-03 | | Release date: | 2004-05-20 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | A recombinant C121S mutant of bovine beta-lactoglobulin is more susceptible to peptic digestion and to denaturation by reducing agents and heating.
Biochemistry, 43, 2004
|
|
1OGW
 
 | | Synthetic Ubiquitin with fluoro-Leu at 50 and 67 | | Descriptor: | UBIQUITIN | | Authors: | Alexeev, D, Ramage, R, Young, D.W, Sawyer, L. | | Deposit date: | 2003-05-13 | | Release date: | 2003-05-30 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.32 Å) | | Cite: | Synthesis, Structural and Biological Studies of Ubiquitin Mutants Containing (2S, 4S)-5-Fluoroleucine Residues Strategically Placed in the Hydrophobic Core
Chembiochem, 4, 2003
|
|
4CK4
 | | Ovine beta-Lactoglobulin at Atomic Resolution | | Descriptor: | ACETATE ION, AMMONIUM ION, BETA_LACTOGLOBULIN-1/B, ... | | Authors: | Kontopidis, G, Nordle, A, Sawyer, L. | | Deposit date: | 2013-12-27 | | Release date: | 2014-10-08 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.12 Å) | | Cite: | Ovine Beta-Lactoglobulin at Atomic Resolution
Acta Crystallogr.,Sect.F, 70, 2014
|
|
1L9W
 
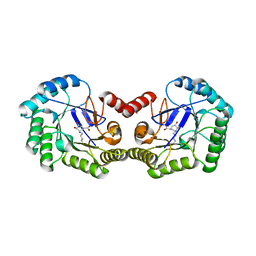 | | CRYSTAL STRUCTURE OF 3-DEHYDROQUINASE FROM SALMONELLA TYPHI COMPLEXED WITH REACTION PRODUCT | | Descriptor: | 3-AMINO-4,5-DIHYDROXY-CYCLOHEX-1-ENECARBOXYLATE, 3-dehydroquinate dehydratase aroD | | Authors: | Lee, W.H, Perles, L.A, Nagem, R.A.P, Shrive, A.K, Hawkins, A, Sawyer, L, Polikarpov, I. | | Deposit date: | 2002-03-26 | | Release date: | 2003-03-26 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Comparison of different crystal forms of 3-dehydroquinase from Salmonella typhi and its implication for the enzyme activity.
Acta Crystallogr.,Sect.D, 58, 2002
|
|
1QFE
 
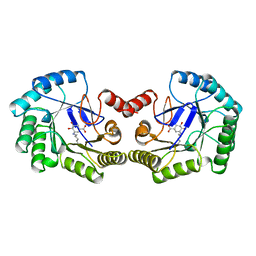 | | THE STRUCTURE OF TYPE I 3-DEHYDROQUINATE DEHYDRATASE FROM SALMONELLA TYPHI | | Descriptor: | 3-AMINO-4,5-DIHYDROXY-CYCLOHEX-1-ENECARBOXYLATE, PROTEIN (3-DEHYDROQUINATE DEHYDRATASE) | | Authors: | Shrive, A.K, Polikarpov, I, Sawyer, L, Coggins, J.R, Hawkins, A.R. | | Deposit date: | 1999-04-05 | | Release date: | 2000-04-05 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The two types of 3-dehydroquinase have distinct structures but catalyze the same overall reaction.
Nat.Struct.Biol., 6, 1999
|
|
1DV9
 
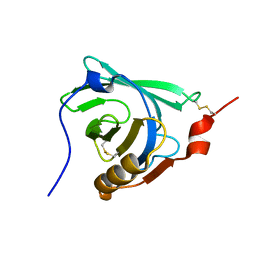 | | STRUCTURAL CHANGES ACCOMPANYING PH-INDUCED DISSOCIATION OF THE B-LACTOGLOBULIN DIMER | | Descriptor: | BETA-LACTOGLOBULIN | | Authors: | Uhrinova, S, Smith, M.H, Jameson, G.B, Uhrin, D, Sawyer, L, Barlow, P.N. | | Deposit date: | 2000-01-20 | | Release date: | 2000-02-09 | | Last modified: | 2022-02-16 | | Method: | SOLUTION NMR | | Cite: | Structural changes accompanying pH-induced dissociation of the beta-lactoglobulin dimer.
Biochemistry, 39, 2000
|
|
1GXA
 
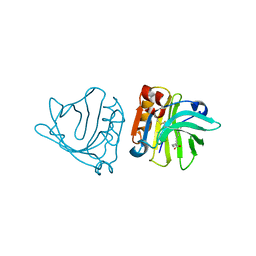 | |
1QG5
 
 | |
1DJE
 
 | | CRYSTAL STRUCTURE OF THE PLP-BOUND FORM OF 8-AMINO-7-OXONANOATE SYNTHASE | | Descriptor: | 8-AMINO-7-OXONANOATE SYNTHASE, PYRIDOXAL-5'-PHOSPHATE, SULFATE ION | | Authors: | Webster, S.P, Alexeev, D, Campopiano, D.J, Watt, R.M, Alexeeva, M, Sawyer, L, Baxter, R.L. | | Deposit date: | 1999-12-02 | | Release date: | 2000-12-04 | | Last modified: | 2018-01-24 | | Method: | X-RAY DIFFRACTION (1.71 Å) | | Cite: | Mechanism of 8-amino-7-oxononanoate synthase: spectroscopic, kinetic, and crystallographic studies.
Biochemistry, 39, 2000
|
|
1DJ9
 
 | | CRYSTAL STRUCTURE OF 8-AMINO-7-OXONANOATE SYNTHASE (OR 7-KETO-8AMINIPELARGONATE OR KAPA SYNTHASE) COMPLEXED WITH PLP AND THE PRODUCT 8(S)-AMINO-7-OXONANONOATE (OR KAPA). THE ENZYME OF BIOTIN BIOSYNTHETIC PATHWAY. | | Descriptor: | 8-AMINO-7-OXONONANOATE SYNTHASE, MAGNESIUM ION, N-[7-KETO-8-AMINOPELARGONIC ACID]-[3-HYDROXY-2-METHYL-5-PHOSPHONOOXYMETHYL-PYRIDIN-4-YL-METHANE], ... | | Authors: | Webster, S.P, Alexeev, D, Campopiano, D.J, Watt, R.M, Alexeeva, M, Sawyer, L, Baxter, R.L. | | Deposit date: | 1999-12-02 | | Release date: | 2000-12-06 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Mechanism of 8-amino-7-oxononanoate synthase: spectroscopic, kinetic, and crystallographic studies.
Biochemistry, 39, 2000
|
|
1B0O
 
 | |
1GX9
 
 | |
1GQN
 
 | | Native 3-dehydroquinase from Salmonella typhi | | Descriptor: | 3-DEHYDROQUINATE DEHYDRATASE | | Authors: | Lee, W.-H, Perles, L.A, Nagem, R.A.P, Polikarpov, I, Sawyer, L. | | Deposit date: | 2001-11-26 | | Release date: | 2003-11-20 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.78 Å) | | Cite: | Comparison of Different Crystal Forms of 3-Dehydroquinase from Salmonella Typhi and its Implications for Enzyme Activity
Acta Crystallogr.,Sect.D, 58, 2002
|
|
1GX8
 
 | |
1BS0
 
 | | PLP-DEPENDENT ACYL-COA SYNTHASE | | Descriptor: | PROTEIN (8-AMINO-7-OXONANOATE SYNTHASE), SULFATE ION | | Authors: | Alexeev, D, Alexeeva, M, Baxter, R.L, Campopiano, D.J, Webster, S.P, Sawyer, L. | | Deposit date: | 1998-08-31 | | Release date: | 1999-08-27 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | The crystal structure of 8-amino-7-oxononanoate synthase: a bacterial PLP-dependent, acyl-CoA-condensing enzyme.
J.Mol.Biol., 284, 1998
|
|
1B8E
 
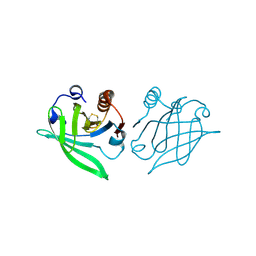 | |
1DTY
 
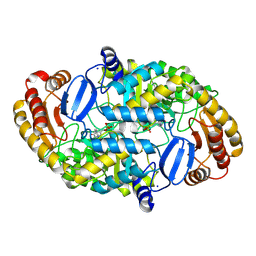 | | CRYSTAL STRUCTURE OF ADENOSYLMETHIONINE-8-AMINO-7-OXONANOATE AMINOTRANSFERASE WITH PYRIDOXAL PHOSPHATE COFACTOR. | | Descriptor: | ADENOSYLMETHIONINE-8-AMINO-7-OXONONANOATE AMINOTRANSFERASE, PYRIDOXAL-5'-PHOSPHATE, SODIUM ION | | Authors: | Alexeev, D, Sawyer, L, Baxter, R.L, Alexeeva, M.V, Campopiano, D. | | Deposit date: | 2000-01-13 | | Release date: | 2000-02-18 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.14 Å) | | Cite: | Crystal structure of adenosylmethionine-8-amino-7-oxonanoate aminotransferase with pyridoxal phosphate cofactor
to be published
|
|
1BEB
 
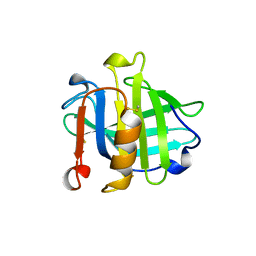 | |
3EST
 
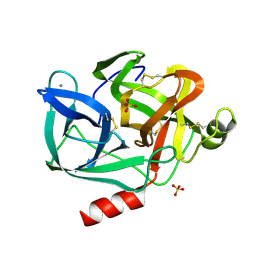 | | STRUCTURE OF NATIVE PORCINE PANCREATIC ELASTASE AT 1.65 ANGSTROMS RESOLUTION | | Descriptor: | CALCIUM ION, PORCINE PANCREATIC ELASTASE, SULFATE ION | | Authors: | Meyer, E.F, Cole, G, Radhakrishnan, R, Epp, O. | | Deposit date: | 1987-09-17 | | Release date: | 1988-01-16 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Structure of native porcine pancreatic elastase at 1.65 A resolutions.
Acta Crystallogr.,Sect.B, 44, 1988
|
|
6GHH
 | |
6GFS
 
 | |
6GF9
 
 | |
6GE7
 
 | |
